Official websites use .gov
A .gov website belongs to an official government organization in the United States.
Secure .gov websites use HTTPS
A lock ( ) or https:// means you've safely connected to the .gov website. Share sensitive information only on official, secure websites.

Public Health Surveillance in Electronic Health Records: Lessons From PCORnet
ORIGINAL RESEARCH — Volume 21 — July 11, 2024
Nidhi Ghildayal, PhD 1 ; Kshema Nagavedu, MPH 1 ; Jennifer L. Wiltz, MD, MPH 2 ; Soowoo Back, MPH 1 ; Tegan K. Boehmer, PhD 3 ; Christine Draper 1 ; Adi V. Gundlapalli, MD, PhD 3 ; Casie Horgan, MPH 1 ; Keith A. Marsolo, PhD 4 ; Nik R. Mazumder, MD, MPH 5 ; Juliane Reynolds, MPH 1 ; Matthew Ritchey, DPT 3 ; Sharon Saydah, PhD 6 ; Yacob G. Tedla 7 ; Thomas W. Carton, PhD 8 ; Jason P. Block, MD, MPH 1 ( View author affiliations )
Suggested citation for this article: Ghildayal N, Nagavedu K, Wiltz JL, Back S, Boehmer TK, Draper C, et al. Public Health Surveillance in Electronic Health Records: Lessons From PCORnet. Prev Chronic Dis 2024;21:230417. DOI: http://dx.doi.org/10.5888/pcd21.230417 .
PEER REVIEWED
Introduction
Acknowledgments, author information.
What is already known on this topic?
Existing survey-based surveillance programs provide important information on the epidemiology of chronic and infectious diseases. Electronic health record (EHR) data can be used to supplement surveillance efforts.
What is added by this report?
In this study, we describe the attributes and challenges of using EHR data for disease surveillance. We describe surveillance case studies and future directions for enhancing opportunities to use EHR data for public health surveillance.
What are the implications for public health practice?
EHR data have an important role for public health surveillance both for chronic and infectious diseases, providing comprehensive information available soon after data collection. Strategic funding and financing models need to be developed, and federal, state, and local support could help establish EHRs as an important sustainable mechanism for surveillance.
PCORnet, the National Patient-Centered Clinical Research Network, is a large research network of health systems that map clinical data to a standardized data model. In 2018, we expanded existing infrastructure to facilitate use for public health surveillance. We describe benefits and challenges of using PCORnet for surveillance and describe case studies.
In 2018, infrastructure enhancements included addition of a table to store patients’ residential zip codes and expansion of a modular program to generate population health statistics across conditions. Chronic disease surveillance case studies conducted in 2019 assessed atrial fibrillation (AF) and cirrhosis. In April 2020, PCORnet established an infrastructure to support COVID-19 surveillance with institutions frequently updating their electronic health record data.
By August 2023, 53 PCORnet sites (84%) had a 5-digit zip code available on at least 95% of their patient populations. Among 148,223 newly diagnosed AF patients eligible for oral anticoagulant (OAC) therapy, 43.3% were on any OAC (17.8% warfarin, 28.5% any novel oral anticoagulant) within a year of the AF diagnosis. Among 60,268 patients with cirrhosis (2015–2019), common documented etiologies included unknown (48%), hepatitis C infection (23%), and alcohol use (22%). During October 2022 through December 2023, across 34 institutions, the proportion of COVID-19 patients who were cared for in the inpatient setting was 9.1% among 887,051 adults aged 20 years or older and 6.0% among 139,148 children younger than 20 years.
Conclusions
PCORnet provides important data that may augment traditional public health surveillance programs across diverse conditions. PCORnet affords longitudinal population health assessments among large catchments of the population with clinical, treatment, and geographic information, with capabilities to deliver rapid information needed during public health emergencies.
Electronic health records (EHRs) contain extensive longitudinal health information about patients and populations (1). Over the last decade, prompted by federal meaningful use guidelines and incentives, EHRs have become ubiquitous in health care settings (2). Because of their wide availability, EHRs are a viable option for disease surveillance and have some advantages over traditional survey-based surveillance methods, such as the National Health and Nutrition Examination Survey and the Behavioral Risk Factor Surveillance System ( Table 1 ) (3,4).
Some of the most important attributes of EHRs for surveillance include timeliness of data and availability for large populations. EHR data are collected daily through routine clinical care delivery and can be made available quickly if resources are available for processing and data curation. In contrast, large national surveillance programs typically use surveys or field data collection, followed by data processing that can lead to extensive lag times between data collection and availability. The scope of EHR data available also can provide important granular information about subgroups. For example, although retrieving metro area and small area modeling estimates via national surveillance surveys is possible, these data are often restricted for privacy reasons and, in some cases, are imputed rather than directly measured (4–6). Furthermore, the sample size of surveys limits the availability of data on rare conditions or less common subgroups of individuals, such as among racial and ethnic minority groups (4,6). Because of the availability of data on vast populations that allow for numbers large enough to stratify by even uncommon subgroups, EHRs can provide data for specific geographic regions and populations (3,6,7).
Another area of potential benefit of EHRs for surveillance is the availability of longitudinal objective, measured data, such as vital signs and laboratory values. These data allow for more accurate definitions (ie, phenotypes) of disease, such as using a combination of medication prescriptions, laboratory values, and vital signs to define chronic disease (4). Measured data can also enable an objective determination of disease severity and disease control over time, such as defining whether patients are meeting guideline control targets for diabetes or hypertension by using glycosylated hemoglobin or measured blood pressure values. These data can provide information both cross-sectionally and longitudinally in cohorts that receive care over time. National surveillance surveys typically rely on self-reported information or single vital sign or laboratory values to define disease prevalence and incidence. Lastly, EHRs can offer longitudinal information with short latency, allowing for capture of information on changing health status, in contrast to that collected through the lengthy process of repeated survey administration (7).
EHR data present some challenges, including with data quality and representativeness. Missing data also are common for myriad reasons. The fragmented health care system in the US precludes comprehensive data integration across care settings, and patients often receive care in multiple institutions with different data systems (8,9). Even when information from other health care institutions can be viewed within the health care system that serves as the medical home for a patient (eg, Care Everywhere in Epic), that information may not be captured in clinical data warehouses that can be used for surveillance. Clinical notes written in free text may not be easily translated to structured data fields, resulting in missing information on symptoms and exposures (10). Furthermore, clinical data available in EHRs do not typically include information on social determinants, quality-of-life measures, and other health behavior information that could be more readily collected through national surveys. Some social determinants data can be integrated when available geographic information can be linked to community-level data from the US Census and other data resources (11,12). Another drawback of EHR surveillance is that data may not be representative, and clinical practice patterns may differ between sites, leading to heterogeneity in data available due solely to the differential ascertainment of diagnoses, for example. Exploration of the epidemiology of disease by geography also is heavily dependent on the number of institutions per geographic area providing data. EHR data have information on patients who are seeking care, likely biasing inferences toward certain demographic groups receiving more medical care and patients who have chronic conditions, have health insurance coverage, or live in urban areas (12). However, unlike claims data that are typically limited to commercial insurance, Medicaid, or Medicare separately, EHR data are typically agnostic to payer source and have information on patients with a diverse array of insurance sources, including those who are uninsured (13) ( Table 1 ).
In this article, we discuss the use of EHR data for public health surveillance in a large national research network and present case studies of its use for chronic disease and its later adaptation for COVID-19 surveillance during a public health emergency.
PCORnet as a data source for public health surveillance
PCORnet, the National Patient-Centered Clinical Research Network, is a research infrastructure program that was established to support use of health care data for comparative effectiveness research (14). This network-of-networks includes more than 60 health care systems embedded in 8 Clinical Research Networks (https://pcornet.org/network/), with a regulatory infrastructure that prioritizes data sharing while protecting patient privacy. Data from millions of patients from different source EHR systems are harmonized locally into a standard data set, called a Common Data Model (CDM). This CDM is updated over time to incorporate new and evolving data elements and is nearly identical across all participating institutions, allowing for centralized querying and interoperability of data across sites (15). The data elements include comprehensive clinical information, including prescriptions, diagnoses, procedures, vital measures, laboratory values, and geographic information, among other data elements, from all care settings relevant for a specific health care system (eg, ambulatory, emergency department, inpatient).
Quarterly data quality reviews allow for a comprehensive assessment of conformance, completeness, plausibility, and persistence, with feedback provided on issues discovered. Data sharing across the network is accommodated by a Master Data Sharing Agreement, with further regulatory processes outlined to accommodate varied circumstances required for research and operations (14,16). PCORnet also has a “front door” mechanism for investigators to request data queries or study collaborators (17).
PCORnet has a distributed query infrastructure, and users can submit a query and obtain a coordinated response that combines data across participating health systems (16). PCORnet also is an engaged network in which investigators, informatics specialists, clinicians, patients, and other partners from sites can provide context and information regarding the data available from that site.
Reusable SAS-based tools that have been developed for PCORnet are available for querying data, with regular updates for CDM changes and to enhance functionality for new data needs. These tools are modular descriptive programs that can be quickly adapted to create and characterize cohorts with aggregate data, using tables and variables defined in the PCORnet CDM. While PCORnet has protocols allowing for the transfer of patient-level data to requestors, the availability of a reusable process for obtaining aggregate data from partners allows for assessments that can often be completed quickly. While less flexible than centralized, pooled data available for analysis, aggregate data are typically sufficient for surveillance.
PCORnet has several capabilities that foster successful public health surveillance. As a national EHR surveillance program with multiple contributing entities, PCORnet contains data on more than 30 million patients annually (16). PCORnet has broad geographic representation with most sites providing data from both inpatient and outpatient settings (https://pcornet.org/data/). The network provides access to patients with longitudinal follow-up, often over many years; populations large enough to allow for examination of subgroups, such as by race and ethnicity, geography, and multimorbidity; and opportunities to capture adequate numbers of patients with rare diseases to make important inferences about prevalence. Data captured on race are considerably more complete than those found in some other commonly used clinical data sets (18). For example, among all patients with encounters in 34 PCORnet sites during October 2022 through December 2023, race and ethnicity information was missing for 9.5% of those younger than 20 years and 8.7% of those aged 20 years or older ( Table 2 ). Race and ethnicity missingness was lower for patients with diagnostic codes for COVID-19, positive laboratory tests for SARS-CoV-2, or recent prescriptions for COVID-19 medications: 7.6% for patients younger than 20 years and 5.1% for patients aged 20 years or older (data not shown).
In 2018, PCORnet began to expand capabilities of the network to conduct EHR-based surveillance, specifically focused on chronic disease. The program was initially used for pilot projects that built capacity for geographic data capture. In March 2020, the network began exploring whether its resources, including the newly established capabilities for chronic disease surveillance, could be adapted for COVID-19 surveillance. This shift required some changes, especially to provide more timely data. PCORnet expanded its infrastructure to include the ability to frequently, up to twice monthly, refresh data. With regularly refreshed data and modular programs, data can now be available for public health professionals and researchers in a matter of weeks. Simple analyses that only require basic counts and frequencies can be provided even more quickly.
Expanded data and tools for surveillance and case studies in PCORnet
Starting in 2019 with CDM version 5.0, PCORnet incorporated a new, optional CDM table containing patient-level geographic information. This table allowed sites to include patient information on 9- and 5-digit zip code, city, state, and the start and end date for that address information. To accommodate surveillance queries in PCORnet, we developed a geographic assessment module to query this address data (16). The module allows for the characterization of a cohort based on the most recent address stratified by zip code, city, state, or Census region. Queries also can pull patient-level data with zip-code or mapped US Census Bureau’s data elements. The geographic module was piloted at several PCORnet health systems for chronic disease surveillance case studies, including atrial fibrillation (AF) and liver cirrhosis.
Starting in April 2020, select PCORnet institutions collaborated on a response to the COVID-19 pandemic that would allow for more frequent querying of data. Institutions developed a CDM that contained data for a subset of their total patient population, including only patients who had a diagnostic code for a respiratory virus or infection or a viral laboratory test for SARS-CoV-2. The inclusion criteria for this subset CDM were later expanded to include COVID-19 therapeutics and vaccines. Filtering the broader population using these criteria allowed for quicker refreshes of data, facilitating reports on data with a latency of a few weeks, in contrast to the regular quarterly updates. This process also was a more practical approach for sites, given that frequent refreshes of their complete patient population data would take extensive effort and data storage. Sites initially updated their filtered CDM biweekly and then later monthly or on request.
The PCORnet team leading this surveillance effort also changed the modular statistical programs to allow for characterization of cohorts using results of qualitative viral testing information, available mortality information (ie, typically deaths reported to the health care system or in-hospital deaths) and records of vaccinations given in the health care system or populated in EHRs from state registry linkages, when available. The statistical programs also were updated to allow for distributed advanced analytics, including the use of multiple regression models that execute behind institutions’ firewalls and return only summary model output; these results can be combined across sites using meta-analytic techniques (19). Since October 2020, PCORnet has participated in a cooperative agreement funded by the Centers for Disease Control and Prevention (CDC) to provide COVID-19 information from up to 43 PCORnet institutions on a biweekly basis.
The geographic query module was released for use in PCORnet in July 2019. Geographic data returned from queries were well distributed but contained many sparsely populated zip codes. Zip code data typically were not available retrospectively; many sites only began capturing the geographic information prospectively at the time of its CDM release. County information was added to the CDM during the release of CDM 6.1 in April 2023.
By August 2023, 62 of 63 (98%) PCORnet institutions had populated geographic information. Among sites, 59 (94%) had at least some information on 5-digit zip code, with 53 (84%) having 5-digit zip available on at least 95% of their patient population. For 9-digit zip codes, 42 (67%) sites had some information on patients, with 10 (16%) having this information populated for at least 75% of their population. The pilot projects on AF and liver cirrhosis tested the implementation of this geographic data table and use of the geographic query module.
Oral anticoagulant use
Oral anticoagulant (OAC) therapy is proven to reduce the risk of stroke and is the standard treatment for stroke risk reduction in patients with AF (20,21). Some local studies have found that about half of patients with AF at risk of stroke do not get OAC prescriptions (22–25). However, little information exists on the rate of prescriptions of OACs across US states. We used data from 4 PCORnet Clinical Research Networks (CAPriCORN, STAR, REACHnet, and ADVANCE) and investigated the OAC prescription rate in 22 states. Patients newly diagnosed with AF between January 2014 and December 2019, with a CHADSVASC score of 2 or more, no history of stroke, and known zip code were included in our analysis. The CHADSVASC score includes information on risk factors for stroke among patients with AF and is used to calculate a predicted probability of stroke; a score of 2 or more is considered high risk for stroke (26). Among 148,223 newly diagnosed AF patients eligible for an OAC, 43.3% were on any OAC, 17.8% received any warfarin, and 28.5% received any novel oral anticoagulant (NOAC) in the year following AF diagnosis. OAC prescription rates varied greatly across states, ranging from 28.4% in Virginia to 54.0% in Indiana.
OAC prescriptions continue to be low in patients with AF and vary across health systems and geographic regions. These results are consistent with findings from previous studies (22–25). Our findings provided comprehensive information on OAC use across regions but were not nationally representative. The study only examined health systems that were part of the CRNs involved in the study: 6 from CAPriCORN, 2 from REACHnet, and 1 each from STAR and ADVANCE.
Cirrhosis, irreversible damage to the liver, is a leading cause of illness and death in the US (27). Despite its importance as a major medical condition, one of the most important challenges for determining population prevalence and geographic distribution is the lack of a unified repository of patients with cirrhosis. PCORnet provided an opportunity to explore the epidemiology of cirrhosis using diagnostic codes in EHRs. In this pilot study, we included any patient aged 18 years or older with a qualifying International Classification of Diseases (ICD) code for cirrhosis (ie, ICD-9 or ICD-10) who received care at a participating center during the calendar years 2015–2018. The study included 9 health systems from 3 Clinical Research Networks, with strong overlap with the AF pilot: STAR, CAPriCORN, and REACHnet. Patient zip code was assessed as zip code of residence both within 90 days of cohort inclusion and within any prior period before inclusion.
Overall, we identified 60,268 patients with ICD codes for cirrhosis. Patients were 58% (n = 34,908) male, 57% (n = 34,458) White race, and 81% (n = 48,646) non-Hispanic ethnicity, with a mean age of 58 years. The most common etiologies for cirrhosis were hepatitis C (n = 13,882; 23%) and alcohol (n = 13,187; 22%); however, nearly half of patients (n = 29,177, 48%) did not have a clear etiology of liver disease documented in the EHR. When geographic data were restricted to a period that was within 90 days of diagnosis of cirrhosis, residential zip code was highly missing (86% missing); missingness was much lower for records of any zip code documented in the EHR before study inclusion (33% missing). This study was conducted relatively soon after the geographic information was first provided in the CDM. Because most sites populated their geographic information prospectively, missingness will improve over time, allowing for PCORnet to be effectively used for important public health surveillance of cirrhosis by geography.
COVID-19 surveillance
PCORnet was able to quickly transition to infectious disease surveillance and began reporting COVID-19 national data in April 2020. Since October 1, 2020, working with CDC, 43 PCORnet institutions have been engaged in a broad surveillance effort in which queries are conducted up to twice monthly on varied topics, with aggregate data provided to CDC in support of pandemic response. The surveillance effort has led to over 50 data queries. In a recent query, focused on the period of October 1, 2022, to December 31, 2023, 34 of the participating PCORnet institutions recorded 887,051 patients aged 20 years or older and 139,148 patients younger than 20 years who tested positive for SARS-COV-2, received a COVID-19 therapeutic, or had an ICD-10 code for COVID-19, with geographic information available by state. Among these patients, 80,712 (9%) of the patients aged 20 years or older and 8,322 (6%) of the patients younger than 20 years were cared for in the inpatient setting. We have mapped zip code data available in this population to designated geographic variables, including US Census Bureau variables for rurality and urbanicity and area deprivation index. These variables were well populated with missing data for 3%–4% of the population. Most patients lived in urban settings (88%–89%). For area deprivation index scores, 44% of patients aged 20 years or older and 51% of patients younger than 20 years were in the top 2 quartiles (ie, higher area deprivation).
This COVID-19 surveillance program has generated important information on the prevalence of post-acute sequelae of SARS-CoV-2 infection (28), disparities in uptake of COVID-19 therapeutics (18,29), cardiac complications after COVID-19 mRNA vaccines and SARS-CoV-2 infection (30), and association of uncontrolled diabetes and hypertension and severe COVID-19 (19). Information also was captured on trends in chronic and infectious disease incidence and preventive care services before and during the pandemic and the incidence of and therapeutics for mpox to support CDC’s response. The infrastructure developed for this CDC-funded project also was leveraged for other large-scale research programs, such as providing preliminary data for the National Institutes of Health RECOVER Initiative (31).
With the availability of patient-level geographic information, large populations, and comprehensive longitudinal clinical data, PCORnet and similar networks can fill in gaps for existing national surveillance infrastructure. Pilot surveillance projects provided valuable lessons for use of PCORnet infrastructure that was leveraged for the national COVID-19 public health response.
Streamlined regulatory processes are critical to accommodate efficient surveillance work. For example, we pursued individual institutional review board (IRB) approvals for each chronic disease surveillance pilot project; all IRB approvals required more than 6 months to complete. Lead sites faced difficulties in coordinating single IRBs (eg, through SMART IRB) across participating sites and ascertaining whether sites should be obtaining IRB determination for limited or nonlimited data sets. These processes were streamlined during the COVID-19 pandemic. The collaborative PCORnet CDC COVID-19 project was exempt from IRB review because it constituted public health surveillance required or authorized by a public health authority, as specified under the Common Rule. Our experience demonstrates that clear network guidance on approvals necessary for varied types of data exchange could help streamline surveillance projects. Fewer requirements should be needed for projects using only aggregate data, even with the inclusion of some geographic information (32). This guidance will be most important for surveillance projects that are not directed by a public health authority and, thus, not exempt from IRB review (33).
Implementation of the pilot projects also revealed both issues and benefits that arose from using various organizing units for geographic data. The city variable was not useful due to varied spellings. Zip code was well-populated at the 5-digit level, providing expanded geographic capabilities that go beyond other data sources, such as insurance claims. Counties can be useful in some cases, such as for states that use geographic divisions other than standard ones (eg, Utah Small Area Codes); county was added as a new geographic unit for the PCORnet CDM in 2023. Ultimately, census tracts or block groups are likely most useful because these geographic units are typically more uniform than larger zip code areas. Regulatory processes could help ease the way for networks to use this information more readily. Direct linkage to US Census Bureau and other community-level data might also preclude the need to share actual geographic identifiers.
Implications for practice
EHR-based networks have important potential for surveillance of key priority areas that align with health and public health missions. When rapidly refreshed data with short latency are required, PCORnet has shown that it can be used for COVID-19 surveillance and other infectious disease outbreaks or epidemics, with data that are available with often very short latency from the time of collection. These rapidly available data in PCORnet allowed for timely reporting of infection trends, including information on patient demographics, comorbidities, and treatments used or prescribed. Timely data can also be important for chronic disease surveillance but may not require updates as frequently as for emerging infectious diseases.
EHR data could be improved to address some of the challenges for its use in surveillance. Developing and deploying population statistical weighting schemes for data in EHRs, which have data only on patients seeking care from designated health care institutions, could help alleviate issues related to generalizability of populations (34,35). Strategic funding and financing models should be developed, and federal, state, and local support could help establish EHRs as an important sustainable mechanism for surveillance (36). The continued success of using PCORnet for large-scale surveillance also can expand its engagement of partners to ensure that data can be used most efficiently to support population health priorities (36).
Dr Block and Dr Carton were co-senior authors and contributed equally to this article. This study was funded in part by a grant from the Patient-Centered Outcomes Research Institute (PCORI 283-3709) and Cooperative Agreement number 6-NU38OT000316, funded by CDC. The authors declare no potential conflicts of interest with respect to the research, authorship, or publication of this article. No copyrighted materials, surveys, instruments, or tools were used in this research.
Corresponding Author: Jason P. Block, MD, MPH, Department of Population Medicine, Harvard Medical School, Harvard Pilgrim Health Care Institute, 401 Park Dr, Ste 401, Boston, MA 02215 ( [email protected] ).
Author Affiliations: 1 Department of Population Medicine, Harvard Medical School, Harvard Pilgrim Health Care Institute, Boston, Massachusetts. 2 National Center for Chronic Disease Prevention and Health Promotion, Centers for Disease Control and Prevention, Atlanta, Georgia. 3 Office of Public Health Data, Surveillance, and Technology, Centers for Disease Control and Prevention, Atlanta, Georgia. 4 Department of Population Health Sciences, Duke Clinical Research Institute, Duke University School of Medicine, Durham, North Carolina. 5 Department of Internal Medicine, University of Michigan Health, Ann Arbor, Michigan. 6 Coronavirus and Other Respiratory Viruses Division, Centers for Disease Control and Prevention, Atlanta, Georgia. 7 Department of Medicine, Vanderbilt University Medical Center, Nashville, Tennessee. 8 Louisiana Public Health Institute, New Orleans, Louisiana.
- Gunter TD, Terry NP. The emergence of national electronic health record architectures in the United States and Australia: models, costs, and questions. J Med Internet Res . 2005;7(1):e3. PubMed doi:10.2196/jmir.7.1.e3
- Agency for Healthcare Research and Quality. Module 17, electronic health records and meaningful use. https://www.ahrq.gov/ncepcr/tools/pf-handbook/mod17.html
- Birkhead GS, Klompas M, Shah NR. Uses of electronic health records for public health surveillance to advance public health. Annu Rev Public Health . 2015;36(1):345–359. PubMed doi:10.1146/annurev-publhealth-031914-122747
- Klompas M, Cocoros NM, Menchaca JT, Erani D, Hafer E, Herrick B, et al. . State and local chronic disease surveillance using electronic health record systems. Am J Public Health . 2017;107(9):1406–1412. PubMed doi:10.2105/AJPH.2017.303874
- Centers for Disease Control and Prevention. PLACES: local data for better health, methodology. Accessed March 28, 2024. https://www.cdc.gov/places/methodology/
- Casey JA, Schwartz BS, Stewart WF, Adler NE. Using electronic health records for population health research: a review of methods and applications. Annu Rev Public Health . 2016;37(1):61–81. PubMed doi:10.1146/annurev-publhealth-032315-021353
- McVeigh KH, Newton-Dame R, Chan PY, Thorpe LE, Schreibstein L, Tatem KS, et al. . Can electronic health records be used for population health surveillance? Validating population health metrics against established survey data. EGEMS (Wash DC) . 2016;4(1):1267. PubMed doi:10.13063/2327-9214.1267
- Eggleston EM, Weitzman ER. Innovative uses of electronic health records and social media for public health surveillance. Curr Diab Rep . 2014;14(3):468. PubMed doi:10.1007/s11892-013-0468-7
- Lenert L, Sundwall DN. Public health surveillance and meaningful use regulations: a crisis of opportunity. Am J Public Health . 2012;102(3):e1–e7. PubMed doi:10.2105/AJPH.2011.300542
- Willis SJ, Cocoros NM, Randall LM, Ochoa AM, Haney G, Hsu KK, et al. . Electronic health record use in public health infectious disease surveillance, USA, 2018–2019. Curr Infect Dis Rep . 2019;21(10):32. PubMed doi:10.1007/s11908-019-0694-5
- Elliott AF, Davidson A, Lum F, Chiang MF, Saaddine JB, Zhang X, et al. . Use of electronic health records and administrative data for public health surveillance of eye health and vision-related conditions in the United States. Am J Ophthalmol . 2012;154(6 Suppl):S63–S70. PubMed doi:10.1016/j.ajo.2011.10.002
- Birkhead GS. Successes and continued challenges of electronic health records for chronic disease surveillance. Am J Public Health . 2017;107(9):1365–1367. PubMed doi:10.2105/AJPH.2017.303938
- Huguet N, Angier H, Rdesinski R, Hoopes M, Marino M, Holderness H, et al. . Cervical and colorectal cancer screening prevalence before and after Affordable Care Act Medicaid expansion. Prev Med . 2019;124:91–97. PubMed doi:10.1016/j.ypmed.2019.05.003
- Forrest CB, McTigue KM, Hernandez AF, Cohen LW, Cruz H, Haynes K, et al. . PCORnet® 2020: current state, accomplishments, and future directions. J Clin Epidemiol . 2021;129:60–67. PubMed doi:10.1016/j.jclinepi.2020.09.036
- National Patient-Centered Clinical Research Network. PCORnet data, common data model (CDM) specification, version 6.0. Accessed March 28, 2024. https://pcornet.org/wp-content/uploads/2020/12/PCORnet-Common-Data-Model-v60-2020_10_221.pdf
- National Patient-Centered Clinical Research Network. Data. Accessed March 28, 2024. https://pcornet.org/data/
- National Patient-Centered Clinical Research Network. Front door. Accessed March 28, 2024. https://pcornet.org/front-door/
- Boehmer TK, Koumans EH, Skillen EL, Kappelman MD, Carton TW, Patel A, et al. . Racial and ethnic disparities in outpatient treatment of COVID-19 — United States, January–July 2022. MMWR Morb Mortal Wkly Rep . 2022;71(43):1359–1365. PubMed doi:10.15585/mmwr.mm7143a2
- Jackson SL, Woodruff RC, Nagavedu K, Fearrington J, Rolka DB, Twentyman E, et al. ; PCORnet Collaboration Authors. Association between hypertension and diabetes control and COVID-19 severity: National Patient-Centered Clinical Research Network, United States, March 2020 to February 2022. J Am Heart Assoc . 2023;12(21):e030240. PubMed doi:10.1161/JAHA.122.030240
- Connolly SJ, Ezekowitz MD, Yusuf S, Eikelboom J, Oldgren J, Parekh A, et al. ; RE-LY Steering Committee and Investigators. Dabigatran versus warfarin in patients with atrial fibrillation. N Engl J Med . 2009;361(12):1139–1151. PubMed doi:10.1056/NEJMoa0905561
- Patel MR, Mahaffey KW, Garg J, Pan G, Singer DE, Hacke W, et al. ; ROCKET AF Investigators. Rivaroxaban versus warfarin in nonvalvular atrial fibrillation. N Engl J Med . 2011;365(10):883–891. PubMed doi:10.1056/NEJMoa1009638
- Dentali F, Riva N, Crowther M, Turpie AG, Lip GY, Ageno W. Efficacy and safety of the novel oral anticoagulants in atrial fibrillation: a systematic review and meta-analysis of the literature. Circulation . 2012;126(20):2381–2391. PubMed doi:10.1161/CIRCULATIONAHA.112.115410
- Marzec LN, Wang J, Shah ND, Chan PS, Ting HH, Gosch KL, et al. . Influence of direct oral anticoagulants on rates of oral anticoagulation for atrial fibrillation. J Am Coll Cardiol . 2017;69(20):2475–2484. PubMed doi:10.1016/j.jacc.2017.03.540
- Essien UR, Holmes DN, Jackson LR II, Fonarow GC, Mahaffey KW, Reiffel JA, et al. . Association of race/ethnicity with oral anticoagulant use in patients with atrial fibrillation: findings from the Outcomes Registry for Better Informed Treatment of Atrial Fibrillation II. JAMA Cardiol . 2018;3(12):1174–1182. PubMed doi:10.1001/jamacardio.2018.3945
- Birman-Deych E, Radford MJ, Nilasena DS, Gage BF. Use and effectiveness of warfarin in Medicare beneficiaries with atrial fibrillation. Stroke . 2006;37(4):1070–1074. PubMed doi:10.1161/01.STR.0000208294.46968.a4
- Camm AJ, Lip GY, De Caterina R, Savelieva I, Atar D, Hohnloser SH, et al. ; ESC Committee for Practice Guidelines (CPG). 2012 Focused update of the ESC Guidelines for the Management of Atrial Fibrillation: an update of the 2010 ESC Guidelines for the Management of Atrial Fibrillation. Developed with the special contribution of the European Heart Rhythm Association. Eur Heart J . 2012;33(21):2719–2747. PubMed doi:10.1093/eurheartj/ehs253
- Centers for Disease Control and Prevention, National Center for Health Statistics. Chronic liver disease and cirrhosis. Accessed March 28, 2024. https://www.cdc.gov/nchs/fastats/liver-disease.htm
- Hernandez-Romieu AC, Carton TW, Saydah S, Azziz-Baumgartner E, Boehmer TK, Garret NY, et al. . Prevalence of select new symptoms and conditions among persons aged younger than 20 years and 20 years or older at 31 to 150 days after testing positive or negative for SARS-CoV-2. JAMA Netw Open . 2022;5(2):e2147053. PubMed doi:10.1001/jamanetworkopen.2021.47053
- Wiltz JL, Feehan AK, Molinari NM, Ladva CN, Truman BI, Hall J, et al. . Racial and ethnic disparities in receipt of medications for treatment of COVID-19 — United States, March 2020–August 2021. MMWR Morb Mortal Wkly Rep . 2022;71(3):96–102. PubMed doi:10.15585/mmwr.mm7103e1
- Block JP, Boehmer TK, Forrest CB, Carton TW, Lee GM, Ajani UA, et al. . Cardiac complications after SARS-CoV-2 infection and mRNA COVID-19 vaccination — PCORnet, United States, January 2021–January 2022. MMWR Morb Mortal Wkly Rep . 2022;71(14):517–523. PubMed doi:10.15585/mmwr.mm7114e1
- RECOVER: Researching COVID to Enhance Recovery. Building capacity for collaboration. March 28, 2024. https://recovercovid.org/infrastructure
- US Department of Health and Human Services. Standards for privacy of individually identifiable health information; final rule. Accessed March 28, 2024. https://www.hhs.gov/sites/default/files/ocr/privacy/hipaa/administrative/privacyrule/privrulepd.pdf
- US Department of Health and Human Services. Activities deemed not to be research: public health surveillance 2018 requirements. Accessed March 28, 2024. https://www.hhs.gov/ohrp/regulations-and-policy/requests-for-comments/draft-guidance-activities-deemed-not-be-research-public-health-surveillance/index.html
- Hohman KH, Martinez AK, Klompas M, Kraus EM, Li W, Carton TW, et al. Leveraging electronic health record data for timely chronic disease surveillance: the Multi-State EHR-Based Network for Disease Surveillance. J Public Health Manag Pract . 2023;29(2):162–73.
- Nasuti L, Andrews B, Li W, Wiltz J, Hohman KH, Patanian M. Using latent class analysis to inform the design of an EHR-based national chronic disease surveillance model. Chronic Illn . 2023;19(3):675–680. PubMed doi:10.1177/17423953221099043
- Patient-Centered Outcomes Research Institute. PCORnet: progress, challenges, and opportunities ahead. Accessed March 28, 2024. https://www.pcori.org/blog/pcornet-progress-challenges-and-opportunities-ahead
| Surveillance system attributes | Traditional national surveillance surveys | EHRs | ||
|---|---|---|---|---|
| Strengths | Weaknesses | Strengths | Weaknesses | |
| NA | Can take years between data collection and availability | Available soon after collected | NA | |
| In-depth availability of patient-reported data on behaviors; extensive collection of social determinants of health data | Limited sample sizes, especially for less common sociodemographic groups | Data on millions of patients provides ability to estimate disease prevalence for rare diseases, less common subgroups (Native Hawaiian/Pacific Islander, American Indian/Alaska Native), and small area geographic units and population-based cohorts | Limited availability of patient-reported data; social determinants data availability increasing but limited to insurance type and linked Census data for many EHRs | |
| Objectively measured health outcomes (vitals, laboratory values) according to study protocol | Cross-sectional or panel designs limit longitudinal follow-up | Longitudinal follow-up on patients allows tracking changes over time; data available on disease control over time | Many data are unstructured (eg, patient notes) and less available for use; structured data standardization is variable; identification of diseases often depends on use of nonspecific diagnostic codes; prescription data typically available but pharmacy dispensing may not be | |
| Nationally representative by design; typically covers entire US population with probability-based sampling strategies | Certain populations can be under-represented (eg, people without a landline telephone, the institutionalized population); characteristics of respondents may differ from nonrespondents in measured or unmeasured ways | Some research networks have data available on people in all US states and territories; patients with multiple types of insurance (commercial and government insurance) are typically available | Representative of care-seeking population, which may limit broad surveillance questions at the population level; representativeness of urban versus rural populations dependent on institutions contributing data | |
| Data collected according to study protocol; robust data completeness and curation | Telephone surveys used in some programs reliant on self-report; all surveys subject to nonresponse | Objective measures of some disease (eg, diabetes, obesity) and robust computable phenotypes of others | Missing data are common; data not collected according to a standardized protocol | |
| Infrastructure established by federal agencies to collect data; sampling and weighting strategies well validated and centrally applied by data collectors; some flexibility on adding new questions and data elements | Requires substantial resources and staff to facilitate | Data collected for routine clinical activities and only additional resources for collection required for new data elements | Data processing requires substantial resources, especially to address data quality issues that can arise; adding new data elements challenging | |
Abbreviation: NA, not applicable. a Examples: National Health and Nutrition Examination Survey (NHANES, www.cdc.gov/nchs/nhanes ), Behavioral Risk Factor Surveillance System (BRFSS, www.cdc.gov/brfss ). b Example: National Patient-Centered Clinical Research Network (PCORnet).
| Race and ethnicity | Children, adolescents, young adults (aged <20 y) | Adults (aged ≥20 y) |
|---|---|---|
| N (%) | ||
| NH American Indian/Alaska Native | 32,351 (0.4) | 100,070 (0.5) |
| NH Asian | 276,545 (3.4) | 686,624 (3.2) |
| NH Black or African American | 1,266,244 (15.7) | 2,973,069 (13.8) |
| Hispanic | 1,743,201 (21.6) | 3,246,099 (15.1) |
| NH Multiple race | 115,798 (1.4) | 54,400 (0.3) |
| NH Native Hawaiian/Other Pacific Islander | 23,348 (0.3) | 40,290 (0.2) |
| NH Other race | 233,508 (2.9) | 436,754 (2.0) |
| NH White | 3,746,223 (46.5) | 12,430,453 (57.8) |
| Missing | 768,425 (9.5) | 1,869,061 (8.7) |
Abbreviations: NH, Non-Hispanic; PCORnet, National Patient-Centered Clinical Research Network. a Includes any patients with a designated ethnicity as Hispanic, regardless of race. All racial groups had ethnicity categories of non-Hispanic or missing/other Hispanic ethnicity.
The opinions expressed by authors contributing to this journal do not necessarily reflect the opinions of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors’ affiliated institutions.
Exit Notification / Disclaimer Policy
- The Centers for Disease Control and Prevention (CDC) cannot attest to the accuracy of a non-federal website.
- Linking to a non-federal website does not constitute an endorsement by CDC or any of its employees of the sponsors or the information and products presented on the website.
- You will be subject to the destination website's privacy policy when you follow the link.
- CDC is not responsible for Section 508 compliance (accessibility) on other federal or private website.
Thank you for visiting nature.com. You are using a browser version with limited support for CSS. To obtain the best experience, we recommend you use a more up to date browser (or turn off compatibility mode in Internet Explorer). In the meantime, to ensure continued support, we are displaying the site without styles and JavaScript.
- View all journals
- Explore content
- About the journal
- Publish with us
- Sign up for alerts
- Review Article
- Published: 04 July 2024
Harnessing EHR data for health research
- Alice S. Tang ORCID: orcid.org/0000-0003-4745-0714 1 ,
- Sarah R. Woldemariam 1 ,
- Silvia Miramontes 1 ,
- Beau Norgeot ORCID: orcid.org/0000-0003-2629-701X 2 ,
- Tomiko T. Oskotsky ORCID: orcid.org/0000-0001-7393-5120 1 &
- Marina Sirota ORCID: orcid.org/0000-0002-7246-6083 1 , 3
Nature Medicine ( 2024 ) Cite this article
660 Accesses
13 Altmetric
Metrics details
- Computational biology and bioinformatics
- Machine learning
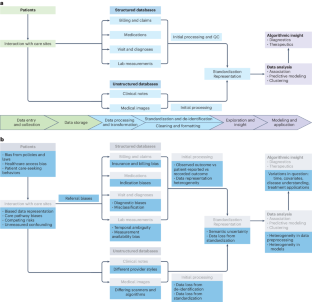
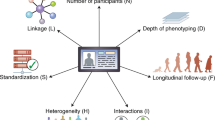
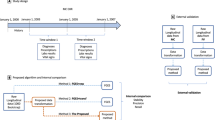
With the increasing availability of rich, longitudinal, real-world clinical data recorded in electronic health records (EHRs) for millions of patients, there is a growing interest in leveraging these records to improve the understanding of human health and disease and translate these insights into clinical applications. However, there is also a need to consider the limitations of these data due to various biases and to understand the impact of missing information. Recognizing and addressing these limitations can inform the design and interpretation of EHR-based informatics studies that avoid confusing or incorrect conclusions, particularly when applied to population or precision medicine. Here we discuss key considerations in the design, implementation and interpretation of EHR-based informatics studies, drawing from examples in the literature across hypothesis generation, hypothesis testing and machine learning applications. We outline the growing opportunities for EHR-based informatics studies, including association studies and predictive modeling, enabled by evolving AI capabilities—while addressing limitations and potential pitfalls to avoid.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
24,99 € / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
195,33 € per year
only 16,28 € per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
Axes of a revolution: challenges and promises of big data in healthcare
A novel method for causal structure discovery from EHR data and its application to type-2 diabetes mellitus
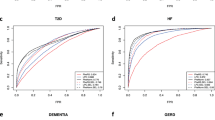

Quantitative disease risk scores from EHR with applications to clinical risk stratification and genetic studies
Gillum, R. F. From papyrus to the electronic tablet: a brief history of the clinical medical record with lessons for the digital age. Am. J. Med. 126 , 853–857 (2013).
Article PubMed Google Scholar
US Food and Drug Administration. Real-World Evidence. FDA https://www.fda.gov/science-research/science-and-research-special-topics/real-world-evidence/ (5 February 2023).
Office of the National Coordinator for Health Information Technology. National Trends in Hospital and Physician Adoption of Electronic Health Records. HealthIT.gov https://www.healthit.gov/data/quickstats/national-trends-hospital-and-physician-adoption-electronic-health-records/ (2021).
Liu, F. & Panagiotakos, D. Real-world data: a brief review of the methods, applications, challenges and opportunities. BMC Med. Res. Methodol. 22 , 287 (2022).
Article PubMed PubMed Central Google Scholar
Cowie, M. R. et al. Electronic health records to facilitate clinical research. Clin. Res. Cardiol. 106 , 1–9 (2017).
Kierkegaard, P. Electronic health record: wiring Europe’s healthcare. Comput. Law Secur. Rev. 27 , 503–515 (2011).
Article Google Scholar
Wen, H. -C., Chang, W. -P., Hsu, M. -H., Ho, C. -H. & Chu, C. -M. An assessment of the interoperability of electronic health record exchanges among hospitals and clinics in Taiwan. JMIR Med. Inform. 7 , e12630 (2019).
Johnson, A. E. W. et al. MIMIC-IV, a freely accessible electronic health record dataset. Sci. Data 10 , 1 (2023).
Article CAS PubMed PubMed Central Google Scholar
All of Us Research Program Investigators. The ‘All of Us’ Research Program. N. Engl. J. Med . 381 , 668–676 (2019).
Sudlow, C. et al. UK Biobank: an open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS Med. 12 , e1001779 (2015).
Sinha, P., Sunder, G., Bendale, P., Mantri, M. & Dande, A. Electronic Health Record: Standards, Coding Systems, Frameworks, and Infrastructures (Wiley, 2012); https://doi.org/10.1002/9781118479612
Overhage, J. M., Ryan, P. B., Reich, C. G., Hartzema, A. G. & Stang, P. E. Validation of a common data model for active safety surveillance research. J. Am. Med. Inform. Assoc. 19 , 54–60 (2012).
Murugadoss, K. et al. Building a best-in-class automated de-identification tool for electronic health records through ensemble learning. Patterns 2 , 100255 (2021).
Yogarajan, V., Pfahringer, B. & Mayo, M. A review of automatic end-to-end de-identification: is high accuracy the only metric? Appl. Artif. Intell. 34 , 251–269 (2020).
Mandl, K. D. & Perakslis, E. D. HIPAA and the leak of ‘deidentified’ EHR data. N. Engl. J. Med. 384 , 2171–2173 (2021).
Norgeot, B. et al. Protected Health Information filter (Philter): accurately and securely de-identifying free-text clinical notes. NPJ Digit. Med. 3 , 57 (2020).
Steurer, M. A. et al. Cohort study of respiratory hospital admissions, air quality and sociodemographic factors in preterm infants born in California. Paediatr. Perinat. Epidemiol. 34 , 130–138 (2020).
Costello, J. M., Steurer, M. A., Baer, R. J., Witte, J. S. & Jelliffe‐Pawlowski, L. L. Residential particulate matter, proximity to major roads, traffic density and traffic volume as risk factors for preterm birth in California. Paediatr. Perinat. Epidemiol. 36 , 70–79 (2022).
Yan, C. et al. Differences in health professionals’ engagement with electronic health records based on inpatient race and ethnicity. JAMA Netw. Open 6 , e2336383 (2023).
Lotfata, A., Moosazadeh, M., Helbich, M. & Hoseini, B. Socioeconomic and environmental determinants of asthma prevalence: a cross-sectional study at the U.S. county level using geographically weighted random forests. Int. J. Health Geogr. 22 , 18 (2023).
Li, L. et al. Identification of type 2 diabetes subgroups through topological analysis of patient similarity. Sci. Transl. Med. 7 , 311ra174 (2015).
De Freitas, J. K. et al. Phe2vec: automated disease phenotyping based on unsupervised embeddings from electronic health records. Patterns 2 , 100337 (2021).
Tang, A. S. et al. Deep phenotyping of Alzheimer’s disease leveraging electronic medical records identifies sex-specific clinical associations. Nat. Commun. 13 , 675 (2022).
Su, C. et al. Clinical subphenotypes in COVID-19: derivation, validation, prediction, temporal patterns, and interaction with social determinants of health. NPJ Digit. Med. 4 , 110 (2021).
Glicksberg, B. S. et al. PatientExploreR: an extensible application for dynamic visualization of patient clinical history from electronic health records in the OMOP common data model. Bioinformatics 35 , 4515–4518 (2019).
Huang, Z., Dong, W., Bath, P., Ji, L. & Duan, H. On mining latent treatment patterns from electronic medical records. Data Min. Knowl. Discov. 29 , 914–949 (2015).
Zaballa, O., Pérez, A., Gómez Inhiesto, E., Acaiturri Ayesta, T. & Lozano, J. A. Identifying common treatments from electronic health records with missing information. An application to breast cancer. PLoS ONE 15 , e0244004 (2020).
Lou, S. S., Liu, H., Harford, D., Lu, C. & Kannampallil, T. Characterizing the macrostructure of electronic health record work using raw audit logs: an unsupervised action embeddings approach. J. Am. Med. Inform. Assoc. 30 , 539–544 (2023).
Glicksberg, B. S. et al. Comparative analyses of population-scale phenomic data in electronic medical records reveal race-specific disease networks. Bioinformatics 32 , i101–i110 (2016).
Obermeyer, Z., Powers, B., Vogeli, C. & Mullainathan, S. Dissecting racial bias in an algorithm used to manage the health of populations. Science 366 , 447–453 (2019).
Article CAS PubMed Google Scholar
Smith, M. A. et al. Insights into measuring health disparities using electronic health records from a statewide network of health systems: a case study. J. Clin. Transl. Sci. 7 , e54 (2023).
Swerdel, J. N., Hripcsak, G. & Ryan, P. B. PheValuator: development and evaluation of a phenotype algorithm evaluator. J. Biomed. Inform. 97 , 103258 (2019).
Denny, J. C. et al. PheWAS: demonstrating the feasibility of a phenome-wide scan to discover gene-disease associations. Bioinformatics 26 , 1205–1210 (2010).
Chen, C., Ding, S. & Wang, J. Digital health for aging populations. Nat. Med. 29 , 1623–1630 (2023).
Woldemariam, S. R., Tang, A. S., Oskotsky, T. T., Yaffe, K. & Sirota, M. Similarities and differences in Alzheimer’s dementia comorbidities in racialized populations identified from electronic medical records. Commun. Med. 3 , 50 (2023).
Austin, P. C. An introduction to propensity score methods for reducing the effects of confounding in observational studies. Multivariate Behav. Res. 46 , 399–424 (2011).
Karlin, L. et al. Use of the propensity score matching method to reduce recruitment bias in observational studies: application to the estimation of survival benefit of non-myeloablative allogeneic transplantation in patients with multiple myeloma relapsing after a first autologous transplantation. Blood 112 , 1133 (2008).
Ho, D., Imai, K., King, G. & Stuart, E. A. MatchIt: nonparametric preprocessing for parametric causal inference. J. Stat. Softw. 42 , 8 (2011).
Zhang, Z., Kim, H. J., Lonjon, G. & Zhu, Y. Balance diagnostics after propensity score matching. Ann. Transl. Med. 7 , 16 (2019).
Landi, I. et al. Deep representation learning of electronic health records to unlock patient stratification at scale. NPJ Digit. Med. 3 , 96 (2020).
Bai, W. et al. A population-based phenome-wide association study of cardiac and aortic structure and function. Nat. Med . https://doi.org/10.1038/s41591-020-1009-y (2020).
Engels, E. A. et al. Comprehensive evaluation of medical conditions associated with risk of non-Hodgkin lymphoma using medicare claims (‘MedWAS’). Cancer Epidemiol. Biomark. Prev. 25 , 1105–1113 (2016).
Article CAS Google Scholar
Bastarache, L., Denny, J. C. & Roden, D. M. Phenome-wide association studies. J. Am. Med. Assoc. 327 , 75–76 (2022).
Yazdany, J. et al. Rheumatology informatics system for effectiveness: a national informatics‐enabled registry for quality improvement. Arthritis Care Res. 68 , 1866–1873 (2016).
Nelson, C. A., Bove, R., Butte, A. J. & Baranzini, S. E. Embedding electronic health records onto a knowledge network recognizes prodromal features of multiple sclerosis and predicts diagnosis. J. Am. Med. Inform. Assoc. 29 , 424–434 (2022).
Tang, A. S. et al. Leveraging electronic health records and knowledge networks for Alzheimer’s disease prediction and sex-specific biological insights. Nat. Aging 4 , 379–395 (2024).
Mullainathan, S. & Obermeyer, Z. Diagnosing physician error: a machine learning approach to low-value health care. Q. J. Econ. 137 , 679–727 (2022).
Makin, T. R. & Orban De Xivry, J. -J. Ten common statistical mistakes to watch out for when writing or reviewing a manuscript. eLife 8 , e48175 (2019).
Carrigan, G. et al. External comparator groups derived from real-world data used in support of regulatory decision making: use cases and challenges. Curr. Epidemiol. Rep. 9 , 326–337 (2022).
Hersh, W. R. et al. Caveats for the use of operational electronic health record data in comparative effectiveness research. Med. Care 51 , S30–S37 (2013).
Rudrapatna, V. A. & Butte, A. J. Opportunities and challenges in using real-world data for health care. J. Clin. Invest. 130 , 565–574 (2020).
Belthangady, C. et al. Causal deep learning reveals the comparative effectiveness of antihyperglycemic treatments in poorly controlled diabetes. Nat. Commun. 13 , 6921 (2022).
Roger, J. et al. Leveraging electronic health records to identify risk factors for recurrent pregnancy loss across two medical centers: a case–control study. Preprint at Res. Sq. https://doi.org/10.21203/rs.3.rs-2631220/v2 (2023).
Gervasi, S. S. et al. The potential for bias in machine learning and opportunities for health insurers to address it: article examines the potential for bias in machine learning and opportunities for health insurers to address it. Health Aff. 41 , 212–218 (2022).
Sai, S. et al. Generative AI for transformative healthcare: a comprehensive study of emerging models, applications, case studies, and limitations. IEEE Access 12 , 31078–31106 (2024).
Wang, M. et al. A systematic review of automatic text summarization for biomedical literature and EHRs. J. Am. Med. Inform. Assoc. 28 , 2287–2297 (2021).
Katsoulakis, E. et al. Digital twins for health: a scoping review. NPJ Digit. Med. 7 , 77 (2024).
Thirunavukarasu, A. J. et al. Large language models in medicine. Nat. Med. 29 , 1930–1940 (2023).
Meskó, B. & Topol, E. J. The imperative for regulatory oversight of large language models (or generative AI) in healthcare. NPJ Digit. Med. 6 , 120 (2023).
Hastings, J. Preventing harm from non-conscious bias in medical generative AI. Lancet Digit. Health 6 , e2–e3 (2024).
Lett, E., Asabor, E., Beltrán, S., Cannon, A. M. & Arah, O. A. Conceptualizing, contextualizing, and operationalizing race in quantitative health sciences research. Ann. Fam. Med. 20 , 157–163 (2022).
Belonwu, S. A. et al. Sex-stratified single-cell RNA-seq analysis identifies sex-specific and cell type-specific transcriptional responses in Alzheimer’s disease across two brain regions. Mol. Neurobiol. https://doi.org/10.1007/s12035-021-02591-8 (2021).
Krumholz, A. Driving and epilepsy: a review and reappraisal. J. Am. Med. Assoc. 265 , 622–626 (1991).
Xu, J. et al. Data-driven discovery of probable Alzheimer’s disease and related dementia subphenotypes using electronic health records. Learn. Health Syst. 4 , e10246 (2020).
Vyas, D. A., Eisenstein, L. G. & Jones, D. S. Hidden in plain sight—reconsidering the use of race correction in clinical algorithms. N. Engl. J. Med. 383 , 874–882 (2020).
Dagdelen, J. et al. Structured information extraction from scientific text with large language models. Nat. Commun. 15 , 1418 (2024).
Hu, Y. et al. Improving large language models for clinical named entity recognition via prompt engineering. J. Am. Med. Inform. Assoc. 27 , ocad259 (2024).
Microsoft. microsoft/FHIR-Converter (2024).
Torfi, A., Fox, E. A. & Reddy, C. K. Differentially private synthetic medical data generation using convolutional GANs. Inf. Sci. 586 , 485–500 (2022).
Yoon, J., Jordon, J. & van der Schaar, M. GAIN: missing data imputation using generative adversarial nets. Preprint at https://arxiv.org/abs/1806.02920v1 (2018).
Shi, J., Wang, D., Tesei, G. & Norgeot, B. Generating high-fidelity privacy-conscious synthetic patient data for causal effect estimation with multiple treatments. Front. Artif. Intell. 5 , 918813 (2022).
Stuart, E. A. Matching methods for causal inference: a review and a look forward. Stat. Sci. 25 , 1–21 (2010).
Murali, L., Gopakumar, G., Viswanathan, D. M. & Nedungadi, P. Towards electronic health record-based medical knowledge graph construction, completion, and applications: a literature study. J. Biomed. Inform. 143 , 104403 (2023).
Li, Y. et al. BEHRT: transformer for electronic health records. Sci. Rep. 10 , 7155 (2020).
Guo, L. L. et al. EHR foundation models improve robustness in the presence of temporal distribution shift. Sci. Rep. 13 , 3767 (2023).
Zhu, R. et al. Clinical pharmacology applications of real‐world data and real‐world evidence in drug development and approval—an industry perspective. Clin. Pharmacol. Ther. 114 , 751–767 (2023).
Voss, E. A. et al. Accuracy of an automated knowledge base for identifying drug adverse reactions. J. Biomed. Inform. 66 , 72–81 (2017).
Taubes, A. et al. Experimental and real-world evidence supporting the computational repurposing of bumetanide for APOE4-related Alzheimer’s disease. Nat. Aging 1 , 932–947 (2021).
Gold, R. et al. Using electronic health record-based clinical decision support to provide social risk-informed care in community health centers: protocol for the design and assessment of a clinical decision support tool. JMIR Res. Protoc. 10 , e31733 (2021).
Varga, A. N. et al. Dealing with confounding in observational studies: a scoping review of methods evaluated in simulation studies with single‐point exposure. Stat. Med. 42 , 487–516 (2023).
Carrigan, G. et al. Using electronic health records to derive control arms for early phase single‐arm lung cancer trials: proof‐of‐concept in randomized controlled trials. Clin. Pharmacol. Ther. 107 , 369–377 (2020).
Infante-Rivard, C. & Cusson, A. Reflection on modern methods: selection bias—a review of recent developments. Int. J. Epidemiol. 47 , 1714–1722 (2018).
Degtiar, I. & Rose, S. A review of generalizability and transportability. Annu. Rev. Stat. Appl. 10 , 501–524 (2023).
Badhwar, A. et al. A multiomics approach to heterogeneity in Alzheimer’s disease: focused review and roadmap. Brain 143 , 1315–1331 (2020).
Stuart, E. A. & Rubin, D. B. Matching with multiple control groups with adjustment for group differences. J. Educ. Behav. Stat. 33 , 279–306 (2008).
Hernan, M. A. & Robins, J. M. Causal Inference: What If (Taylor and Francis, 2024).
Hernan, M. A. Causal knowledge as a prerequisite for confounding evaluation: an application to birth defects epidemiology. Am. J. Epidemiol. 155 , 176–184 (2002).
Dang, L. E. et al. A causal roadmap for generating high-quality real-world evidence. J. Clin. Transl. Sci. 7 , e212 (2023).
Hernán, M. A. & Robins, J. M. Using big data to emulate a target trial when a randomized trial is not available. Am. J. Epidemiol. 183 , 758–764 (2016).
Oskotsky, T. et al. Mortality risk among patients with COVID-19 prescribed selective serotonin reuptake inhibitor antidepressants. JAMA Netw. Open 4 , e2133090 (2021).
Sperry, M. M. et al. Target-agnostic drug prediction integrated with medical record analysis uncovers differential associations of statins with increased survival in COVID-19 patients. PLoS Comput. Biol. 19 , e1011050 (2023).
Amit, G. et al. Antidepressant use during pregnancy and the risk of preterm birth – a cohort study. NPJ Womens Health 2 , 5 (2024); https://doi.org/10.1038/s44294-024-00008-0
Download references
Author information
Authors and affiliations.
Bakar Computational Health Sciences Institute, University of California, San Francisco, San Francisco, CA, USA
Alice S. Tang, Sarah R. Woldemariam, Silvia Miramontes, Tomiko T. Oskotsky & Marina Sirota
Qualified Health, Palo Alto, CA, USA
Beau Norgeot
Department of Pediatrics, University of California, San Francisco, San Francisco, CA, USA
Marina Sirota
You can also search for this author in PubMed Google Scholar
Corresponding author
Correspondence to Marina Sirota .
Ethics declarations
Competing interests.
B.N. is an employee at Qualified Health. The other authors declare no competing interests.
Peer review
Peer review information.
Nature Medicine thanks Wenbo Wu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Karen O’Leary, in collaboration with the Nature Medicine team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
Reprints and permissions
About this article
Cite this article.
Tang, A.S., Woldemariam, S.R., Miramontes, S. et al. Harnessing EHR data for health research. Nat Med (2024). https://doi.org/10.1038/s41591-024-03074-8
Download citation
Received : 03 January 2024
Accepted : 17 May 2024
Published : 04 July 2024
DOI : https://doi.org/10.1038/s41591-024-03074-8
Share this article
Anyone you share the following link with will be able to read this content:
Sorry, a shareable link is not currently available for this article.
Provided by the Springer Nature SharedIt content-sharing initiative
Quick links
- Explore articles by subject
- Guide to authors
- Editorial policies
Sign up for the Nature Briefing newsletter — what matters in science, free to your inbox daily.
- SUGGESTED TOPICS
- The Magazine
- Newsletters
- Managing Yourself
- Managing Teams
- Work-life Balance
- The Big Idea
- Data & Visuals
- Reading Lists
- Case Selections
- HBR Learning
- Topic Feeds
- Account Settings
- Email Preferences
It’s Time for a New Kind of Electronic Health Record
- John Glaser

We need to shift from reactive to preventative care.
Well before the Covid-19 pandemic struck, electronic health records were the bane of physicians’ existences. In all too many cases, EHRs seemed to create a huge amount of extra work and generate too few benefits. The pandemic has made the deficiencies of these systems glaringly apparent. This article discusses how EHRs should be transformed so they become an indispensable tool in keeping individual patients and patient populations healthy.
In these difficult times, we’ve made a number of our coronavirus articles free for all readers. To get all of HBR’s content delivered to your inbox, sign up for the Daily Alert newsletter.
The Covid-19 pandemic presents the U.S. health care system with a mind-boggling array of challenges. One of the most urgent is coping with a simultaneous glut and dearth of information . Between tracking outbreaks, staying abreast of the latest information on effective treatments and vaccine development, keeping tabs on how each patient is doing, and recognizing and documenting a seemingly endless stream of weird new symptoms, the entire medical community is being chronically overwhelmed.
- John Glaser is an executive in residence at Harvard Medical School. He previously served as the CIO of Partners Healthcare (now Mass General Brigham), a senior vice president at Cerner, and the CEO of Siemens Health Services. He is co-chair of the HL7 Advisory Council and a board member of the National Committee for Quality Assurance.
Partner Center

Certification of Health IT
Health information technology advisory committee (hitac), health equity, hti-1 final rule, hti-2 proposed rule, information blocking, interoperability, patient access to health records, clinical quality and safety, health it and health information exchange basics, health it in health care settings, health it resources, laws, regulation, and policy, onc funding opportunities, onc hitech programs, privacy, security, and hipaa, scientific initiatives, standards & technology, usability and provider burden.

Specialists Achieve Meaningful Use with Support from Kentucky’s Regional Extension Center and the Department for Medicaid Services
Vendors and communities working together: a catalyst for interoperability and exchange, successful electronic information exchange through direct pilot implementation with cerner and the lewis and clark information exchange (lacie), medallies and the direct project support secure exchange of clinical information in ehr systems, care coordination improved through health information exchange, viewing patients as partners: patient portal implementation and adoption, urban health plan in new york uses its ehr meaningfully to improve care coordination, solo family practitioner demonstrates care coordination with referring physicians, small practice monitors clinical quality through ehr system templates, rural health clinic exchanges information with hospitals and physicians for improved coordination of care, reducing vaccine preventable disease through immunization registries, quality improvement in a primary care practice.
Open Survey
EDITORIAL article
Editorial: ethical considerations in electronic data in healthcare.

- 1 Department of Computer Engineering, Faculty of Engineering, The Hashemite University, Zarqa, Jordan
- 2 College of Technological Innovation, Zayed University, Abu Dhabi, United Arab Emirates
Editorial on the Research Topic Ethical considerations in electronic data in healthcare
1 Introduction
Electronic data has revolutionized the healthcare sector in the digital age, promising enhanced patient care, streamlined operations, and groundbreaking medical research. However, this transformation has complex ethical challenges that need careful consideration. The surge in electronic health records (EHRs), big data analytics, and telemedicine raises significant questions about privacy, consent, data ownership, and equity. Integrating these technologies into our healthcare systems is crucial to navigating these ethical dilemmas thoughtfully.
This editorial explores the ethical considerations surrounding electronic data in healthcare, drawing insights from a series of articles that explore various facets of this multifaceted issue. These contributions collectively provide a comprehensive view of the challenges and propose pathways for ethically sound practices in managing electronic healthcare data.
2 Privacy and confidentiality: safeguarding patient information
One of the foremost ethical concerns is the protection of patient privacy in an era where data breaches and cyber-attacks are increasingly common. Carmichael et al. 's article, “ Personal Data Store Ecosystems in Health and Social Care ,” underscores the need for robust security measures to prevent unauthorized access to sensitive patient information. She highlights the tension between the accessibility of data for medical purposes and the imperative to protect patient confidentiality.
3 Informed consent: respecting patient autonomy
Informed consent is a cornerstone of ethical healthcare practices, but its application becomes complex with electronic data. Benevento et al. explore this Research Topic in their article, “ Measuring the willingness to share personal Health information: a systematic review .”
4 Data ownership and control: who owns the data?
The question of data ownership is another critical ethical issue. In the article, “ Brave (in a) New World: An Ethical Perspective on Chatbots for Medical Advice ,” Erren et al. examine the legal and ethical implications of data ownership in the healthcare sector. They discuss the competing interests of patients, healthcare providers, and third-party companies, and advocate for policies that prioritize patient rights.
5 Equity and access: bridging the digital divide
The digital divide presents a significant barrier to equitable healthcare. Adepoju et al. address this in their piece, “ Access to Technology, Internet Usage, and Online Health information-seeking behaviors in a racially diverse, lower-income population .” They highlight how disparities in digital access can exacerbate existing health inequalities, with marginalized communities often being the most disadvantaged. The authors advocate for policies and initiatives that promote digital literacy and provide equitable access to technology, ensuring that the benefits of electronic data in healthcare are shared broadly across all segments of society.
6 Ethical use of big data: balancing innovation and privacy
The utilization of big data in healthcare offers immense innovation potential, but it also poses significant ethical challenges. Pu et al. 's article, “ A Medical Big Data Access Control Model Based on Smart Contracts and Risk in the Blockchain Environment ,” investigates the ethical considerations of using large datasets for medical research and decision-making. He discusses the balance between the benefits of big data, such as improved patient outcomes and medical advancements, and the risks, including privacy violations and data misuse. Pu et al. emphasizes the need for ethical frameworks that guide the responsible use of big data while fostering innovation.
7 Conclusion
As we navigate the digital transformation of healthcare, it is imperative to address the ethical challenges associated with electronic data. Protecting patient privacy, ensuring informed consent, safeguarding against digital threats, promoting equity and access, and maintaining transparency and accountability are all critical components of ethical practice in this new landscape. The insights from the articles in this series highlight the complexities and propose thoughtful approaches to managing these Research Topic. The ethical considerations in healthcare data demand our attention and action. Together, these articles offer a roadmap for healthcare providers, policymakers, and technology developers to build a more ethical and inclusive healthcare system, where the promise of electronic data can be fully realized without compromising ethical standards.
8 Summary of contributing articles
1. “ Barriers and facilitators related to healthcare practitioner use of real-time prescription monitoring tools in Australia ” by Hoppe et al. :
- Using an online survey, investigate the barriers and facilitators related to healthcare practitioners' use of real-time prescription monitoring (RTPM) tools in Australia.
- Further research is needed to gain an understanding of healthcare practitioners' use of RTPM tools and how to minimize barriers and optimize use for the essential delivery of quality healthcare.
2. “ Measuring the willingness to share personal health information: a systematic review ” by Benevento et al. :
- Analyze the determinants and describe the measurement of the willingness to disclose personal health information.
- Systematic review of articles assessing willingness to share personal health information as a primary or secondary outcome.
3. “ Brave (in a) new world: an ethical perspective on chatbots for medical advice ” by Erren et al. :
- Emphasizes the significant ethical challenges associated with the use of AI chatbots in medical contexts, such as privacy and confidentiality.
- Discusses the necessity of regulating AI, particularly in the medical field, to avoid potential harms, and raises critical questions about who controls AI, how personal data is protected, and who is liable for the advice provided by AI.
4. “ Access to technology, internet usage, and online health information-seeking behaviors in a racially diverse, lower-income population ” by Adepoju et al. :
- Examines access to technology, internet usage, and online health information-seeking behaviors, in a racially diverse, lower-income population using a survey.
- Identifies the gap between technology adoption and effective use for health purposes, highlighting a critical area for improving public health efforts to leverage digital resources.
- Revealed that higher income, higher education levels, and female gender were significantly associated with increased online health information-seeking behaviors.
5. “ Personal data store ecosystems in health and social care ” by Carmichael et al. :
- Highlights the potential of personal data storage to transform health and social care through enhanced individual data control and usage.
- Points out the significant challenges that need to be addressed for their successful adoption, such as Technical and Operational Hurdles, User Engagement, and Data Governance.
6. “ A Medical Big Data Access Control Model based on Smart Contracts and Risk in the Blockchain Environment ” by Pu et al. :
- Proposes a smart contract and risk-based access control model (SCR-BAC) integrated with traditional risk-based access control and deploys risk-based access control policies in the form of smart contracts into the blockchain, thereby ensuring the protection of medical data.
- Demonstrates that the access control model effectively curbs the access behavior of malicious doctors to a certain extent and imposes a limiting effect on the internal abuse and privacy leakage of medical big data.
7. “ Large language models in physical therapy: time to adapt and adept ” by Naqvi et al. :
- Examines how large language models (LLMs) driven by deep ML can offer human-like performance but face challenges in accuracy due to vast data in Physical Therapy (PT) and rehabilitation practice.
- Urges PTs to engage in learning and shaping AI models by highlighting the need for ethical use and human supervision to address potential biases.
Through a comprehensive understanding and proactive management of these ethical issues, we can ensure that the digital revolution in healthcare is both transformative and just, benefiting all patients and society.
Author contributions
DM: Conceptualization, Investigation, Methodology, Project administration, Resources, Supervision, Writing – original draft, Writing – review & editing. MA-K: Conceptualization, Investigation, Methodology, Validation, Writing – review & editing.
The author(s) declare that no financial support was received for the research, authorship, and/or publication of this article.
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Publisher's note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Keywords: patient privacy, informed consent, data security, digital health equity, ethical data use
Citation: Mustafa D and Al-Kfairy M (2024) Editorial: Ethical considerations in electronic data in healthcare. Front. Public Health 12:1454323. doi: 10.3389/fpubh.2024.1454323
Received: 24 June 2024; Accepted: 04 July 2024; Published: 15 July 2024.
Edited and reviewed by: Himel Mondal , All India Institute of Medical Sciences, Deoghar (AIIMS Deoghar), India
Copyright © 2024 Mustafa and Al-Kfairy. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY) . The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Dheya Mustafa, dheya@hu.edu.jo
Disclaimer: All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Electronic Health Records: A Case Study of an Implementation
- Conference paper
- Cite this conference paper

- Guillaume Cusseau 17 ,
- Jon Grinsell 17 ,
- Christopher Wenzel 17 &
- Fan Zhao 17
Part of the book series: Lecture Notes in Computer Science ((LNISA,volume 8005))
Included in the following conference series:
- International Conference on Human-Computer Interaction
5438 Accesses
Since healthcare institutions have to manage efficiently many terabytes of data on their patients, they need tools that allow them to have an easy access to their data and that enable them to share their data with every specialist involved in the treatment of a patient. That’s why they increasingly adopt EMR and EHR systems. As they are quite recent systems, healthcare institutions usually lack of experience to implement these systems. The purpose of this paper is to do a case study on the implementation of an EHR system in a local healthcare institution, and then to analyze this case study to give directions so as to avoid some arising issues.
Download to read the full chapter text
Chapter PDF
Similar content being viewed by others.

A Study of Electronic Health Record to Unfold Its Significance for Medical Reforms

A Technological Framework for EHR Interoperability: Experiences from Italy

Views on Electronic Health Record
- Implementation
Bureau of Labor Statistics, Occupational Outlook Handbook, 2010-2011 Edition, http://www.bls.gov/oco/ocos074.htm#emply
Burns, F.: Information for Health: An information strategy for the modern NHS 1998-2005. A national strategy for local implementation (2006)
Google Scholar
Tatum, M.: What are Electronic Health Records? Wisegeek.com, n.d. Web, http://www.wisegeek.com/what-are-electronic-health-records.htm (November 21, 2011)
Walker, J., Bieber, E., Richards, F.: Implementing an Electronic Health Record System. Springer, London (2005)
Book Google Scholar
Carter, J.: Electronic Health Records: A Guide for Clinicians and Administrators, 2nd edn. American College of Physicians, U.S.A. (2008)
Hoerbst, A., Ammenwerth, E.: A Structural Model for Quality Requirements regarding Electronic Health Records – State of the art and first concepts
Blacharski, D.: What is ERP (Enterprise Resource Planning)? Wisegeek.com. Web (September 22, 2011), http://www.wisegeek.com/what-is-enterprise-resource-planning.htm (November 21, 2011)
Chester, K.: Benefits of Enterprise Resource Planning (ERP) Systems. Ezinearticles.com. Web (February 04, 2011), http://ezinearticles.com/?Benefits-Of-Enterprise-Resource-Planning-ERP-Systems&id=5855906 (November 21, 2011)
Ge, X., Paige, R.F., McDermid, J.A.: Domain analysis on an Electronic Health Records System
Hristidis, V., Clarke, P.J., Prabakar, N., Deng, Y., White, J.A., Burke, R.P.: A Flexible Approach for Electronic Medical Records Exchange
Torrey, T.: The Benefits of Electronic Medical Records (EMRs). About.com. Web (April 11, 2011), http://patients.about.com/od/electronicpatientrecords/a/EMRbenefits.htm (November 21, 2011)
Gurley, L.: Advantages and Disadvantages of the Electronic Medical Record. Aameda.org. Web (2004), http://www.aameda.org/MemberServices/Exec/Articles/spg04/Gurley%20article.pdf (November 21, 2011)
Chimezie, O.: Electronic Health Record. Articlebase.com. Web (April 11, 2011), http://www.articlesbase.com/health-articles/electronic-health-record-4581106.html (November 21, 2011)
Artio, C.: Advantages of Electronic Health Record System. Ezinearticles.com. Web (August 06, 2009), http://ezinearticles.com/?Advantages-of-Electronic-Health-Record-System&id=2720601 (November 21, 2011)
Pounders, A.: What Are Electronic Health Records? Ezinearticles.com. Web (October 01, 2011), http://ezinearticles.com/?What-Are-Electronic-Health-Records?&id=6598849 (November 21, 2011)
Oz, E.: Management Information Systems, 5th edn. Thomson Learning, Canada (2009)
Stahl, B.: Information Systems: Critical Perspectives, 6th edn. Routledge, Oxon (2008)
Kuziemsky, C.E., Williams, J.B.: Towards Electronic Health Record Support for Collaborative Processes
Young, K.: Informatics for Healthcare Professionals. F.A. Davis, Philadelphia (2000)
Frankk, D.: Enterprise Resource Planning - An Introduction. Ezinearticles.com. Web (August 01, 2011), http://ezinearticles.com/?Enterprise-Resource-Planning---An-Introduction&id=6464120 (November 21, 2011)
Sartipi, K., Yarmand, M.H., Down, D.G.: Mined-knowledge and Decision Support Services in Electronic Health
Tang, C., Carpendale, S.: Evaluating the Deployment of a Mobile Technology in a Hospital Ward
Ebadollahi, S., Tanenblatt, M.A., Coden, A.R., Chang, S.-F., Syeda-Mahmood, T., Amir, A.: Concept-Based Electronic Health Records: Opportunities and Challenges
Download references
Author information
Authors and affiliations.
Florida Gulf Coast University, Florida, USA
Guillaume Cusseau, Jon Grinsell, Christopher Wenzel & Fan Zhao
You can also search for this author in PubMed Google Scholar
Editor information
Editors and affiliations.
The Open University of Japan, 2-11 Wakaba, Mihama-ku, 261-8586, Chiba-shi, Japan
Masaaki Kurosu
Rights and permissions
Reprints and permissions
Copyright information
© 2013 Springer-Verlag Berlin Heidelberg
About this paper
Cite this paper.
Cusseau, G., Grinsell, J., Wenzel, C., Zhao, F. (2013). Electronic Health Records: A Case Study of an Implementation. In: Kurosu, M. (eds) Human-Computer Interaction. Applications and Services. HCI 2013. Lecture Notes in Computer Science, vol 8005. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-39262-7_6
Download citation
DOI : https://doi.org/10.1007/978-3-642-39262-7_6
Publisher Name : Springer, Berlin, Heidelberg
Print ISBN : 978-3-642-39261-0
Online ISBN : 978-3-642-39262-7
eBook Packages : Computer Science Computer Science (R0)
Share this paper
Anyone you share the following link with will be able to read this content:
Sorry, a shareable link is not currently available for this article.
Provided by the Springer Nature SharedIt content-sharing initiative
- Publish with us
Policies and ethics
- Find a journal
- Track your research
An official website of the United States government
The .gov means it’s official. Federal government websites often end in .gov or .mil. Before sharing sensitive information, make sure you’re on a federal government site.
The site is secure. The https:// ensures that you are connecting to the official website and that any information you provide is encrypted and transmitted securely.
- Publications
- Account settings
- My Bibliography
- Collections
- Citation manager
Save citation to file
Email citation, add to collections.
- Create a new collection
- Add to an existing collection
Add to My Bibliography
Your saved search, create a file for external citation management software, your rss feed.
- Search in PubMed
- Search in NLM Catalog
- Add to Search
Implementing an Open Source Electronic Health Record System in Kenyan Health Care Facilities: Case Study
Affiliations.
- 1 KEMRI/Wellcome Trust Research Programme, Nairobi, Kenya.
- 2 e-Health and Systems Development Unit, Ministry of Health, Nairobi, Kenya.
- 3 Vimak Company Limited, Nairobi, Kenya.
- 4 Brown Center for Biomedical Informatics, Brown University, Providence, RI, United States.
- 5 Nuffield Department of of Primary Care Health Sciences, University of Oxford, Oxford, United Kingdom.
- 6 Nuffield Department of Medicine, University of Oxford, Oxford, United Kingdom.
- 7 Centre for Tropical Medicine and Global Health, Nuffield Department of Medicine, University of Oxford, Oxford, United Kingdom.
- PMID: 29669709
- PMCID: PMC5932328
- DOI: 10.2196/medinform.8403
Background: The Kenyan government, working with international partners and local organizations, has developed an eHealth strategy, specified standards, and guidelines for electronic health record adoption in public hospitals and implemented two major health information technology projects: District Health Information Software Version 2, for collating national health care indicators and a rollout of the KenyaEMR and International Quality Care Health Management Information Systems, for managing 600 HIV clinics across the country. Following these projects, a modified version of the Open Medical Record System electronic health record was specified and developed to fulfill the clinical and administrative requirements of health care facilities operated by devolved counties in Kenya and to automate the process of collating health care indicators and entering them into the District Health Information Software Version 2 system.
Objective: We aimed to present a descriptive case study of the implementation of an open source electronic health record system in public health care facilities in Kenya.
Methods: We conducted a landscape review of existing literature concerning eHealth policies and electronic health record development in Kenya. Following initial discussions with the Ministry of Health, the World Health Organization, and implementing partners, we conducted a series of visits to implementing sites to conduct semistructured individual interviews and group discussions with stakeholders to produce a historical case study of the implementation.
Results: This case study describes how consultants based in Kenya, working with developers in India and project stakeholders, implemented the new system into several public hospitals in a county in rural Kenya. The implementation process included upgrading the hospital information technology infrastructure, training users, and attempting to garner administrative and clinical buy-in for adoption of the system. The initial deployment was ultimately scaled back due to a complex mix of sociotechnical and administrative issues. Learning from these early challenges, the system is now being redesigned and prepared for deployment in 6 new counties across Kenya.
Conclusions: Implementing electronic health record systems is a challenging process in high-income settings. In low-income settings, such as Kenya, open source software may offer some respite from the high costs of software licensing, but the familiar challenges of clinical and administration buy-in, the need to adequately train users, and the need for the provision of ongoing technical support are common across the North-South divide. Strategies such as creating local support teams, using local development resources, ensuring end user buy-in, and rolling out in smaller facilities before larger hospitals are being incorporated into the project. These are positive developments to help maintain momentum as the project continues. Further integration with existing open source communities could help ongoing development and implementations of the project. We hope this case study will provide some lessons and guidance for other challenging implementations of electronic health record systems as they continue across Africa.
Keywords: Kenya; electronic health records; medical records; open source; software.
©Naomi Muinga, Steve Magare, Jonathan Monda, Onesmus Kamau, Stuart Houston, Hamish Fraser, John Powell, Mike English, Chris Paton. Originally published in JMIR Medical Informatics (http://medinform.jmir.org), 18.04.2018.
PubMed Disclaimer
Conflict of interest statement
Conflicts of Interest: HF is a cofounder of the OpenMRS EHR project and unpaid member of the OpenMRS leadership team.
Proposed electronic health record at…
Proposed electronic health record at facility level (source: Kenyan Ministry of Health). LMIS:…
Similar articles
- Digital health Systems in Kenyan Public Hospitals: a mixed-methods survey. Muinga N, Magare S, Monda J, English M, Fraser H, Powell J, Paton C. Muinga N, et al. BMC Med Inform Decis Mak. 2020 Jan 6;20(1):2. doi: 10.1186/s12911-019-1005-7. BMC Med Inform Decis Mak. 2020. PMID: 31906932 Free PMC article.
- Site readiness assessment preceding the implementation of a HIV care and treatment electronic medical record system in Kenya. Muthee V, Bochner AF, Kang'a S, Owiso G, Akhwale W, Wanyee S, Puttkammer N. Muthee V, et al. Int J Med Inform. 2018 Jan;109:23-29. doi: 10.1016/j.ijmedinf.2017.10.019. Epub 2017 Oct 27. Int J Med Inform. 2018. PMID: 29195702
- Open-Source Electronic Health Record Systems for Low-Resource Settings: Systematic Review. Syzdykova A, Malta A, Zolfo M, Diro E, Oliveira JL. Syzdykova A, et al. JMIR Med Inform. 2017 Nov 13;5(4):e44. doi: 10.2196/medinform.8131. JMIR Med Inform. 2017. PMID: 29133283 Free PMC article. Review.
- People, Process and Technology: Strategies for Assuring Sustainable Implementation of EMRs at Public-Sector Health Facilities in Kenya. Kang'a SG, Muthee VM, Liku N, Too D, Puttkammer N. Kang'a SG, et al. AMIA Annu Symp Proc. 2017 Feb 10;2016:677-685. eCollection 2016. AMIA Annu Symp Proc. 2017. PMID: 28269864 Free PMC article.
- A national standards-based assessment on functionality of electronic medical records systems used in Kenyan public-Sector health facilities. Kang'a S, Puttkammer N, Wanyee S, Kimanga D, Madrano J, Muthee V, Odawo P, Sharma A, Oluoch T, Robinson K, Kwach J, Lober WB. Kang'a S, et al. Int J Med Inform. 2017 Jan;97:68-75. doi: 10.1016/j.ijmedinf.2016.09.013. Epub 2016 Sep 23. Int J Med Inform. 2017. PMID: 27919397 Review.
- Implementing Clinical Information Systems in Sub-Saharan Africa: Report and Lessons Learned From the MatLook Project in Cameroon. Bediang G. Bediang G. JMIR Med Inform. 2023 Oct 18;11:e48256. doi: 10.2196/48256. JMIR Med Inform. 2023. PMID: 37851502 Free PMC article.
- Determinants of Implementation of a Critical Care Registry in Asia: Lessons From a Qualitative Study. Tolppa T, Pari V, Pell C, Aryal D, Hashmi M, Shamal Ghalib M, Jawad I, Tripathy S, Tirupakuzhi Vijayaraghavan BK, Beane A, Dondorp AM, Haniffa R; Collaboration of Research Implementation & Training in Critical Care in Asia Investigators. Tolppa T, et al. J Med Internet Res. 2023 Mar 6;25:e41028. doi: 10.2196/41028. J Med Internet Res. 2023. PMID: 36877557 Free PMC article.
- Adapting Longstanding Public Health Collaborations between Government of Kenya and CDC Kenya in Response to the COVID-19 Pandemic, 2020-2021. Herman-Roloff A, Aman R, Samandari T, Kasera K, Emukule GO, Amoth P, Chen TH, Kisivuli J, Weyenga H, Hunsperger E, Onyango C, Juma B, Munyua P, Wako D, Akelo V, Kimanga D, Ndegwa L, Mohamed AA, Okello P, Kariuki S, De Cock KM, Bulterys M; CDC-Kenya COVID-19 Response Team. Herman-Roloff A, et al. Emerg Infect Dis. 2022 Dec;28(13):S159-S167. doi: 10.3201/eid2813.211550. Emerg Infect Dis. 2022. PMID: 36502403 Free PMC article. Review.
- Implementing Electronic Health Records in Primary Care Using the Theory of Change: Nigerian Case Study. Adedeji T, Fraser H, Scott P. Adedeji T, et al. JMIR Med Inform. 2022 Aug 11;10(8):e33491. doi: 10.2196/33491. JMIR Med Inform. 2022. PMID: 35969461 Free PMC article.
- Open-Source Software for Public Health: Opportunities and Approaches. Wanger J, Cullen T, Dunbar EL, Flowers JL. Wanger J, et al. Sex Transm Dis. 2023 Aug 1;50(8S Suppl 1):S31-S33. doi: 10.1097/OLQ.0000000000001689. Epub 2022 Aug 10. Sex Transm Dis. 2023. PMID: 35948283 Free PMC article. No abstract available.
- Fraser HS, Biondich P, Moodley D, Choi S, Mamlin BW, Szolovits P. Implementing electronic medical record systems in developing countries. Inform Prim Care. 2005;13(2):83–95. - PubMed
- Fraser HS, Blaya J. Implementing medical information systems in developing countries, what works and what doesn't. AMIA Annu Symp Proc. 2010;2010:232–6. http://europepmc.org/abstract/MED/21346975 - PMC - PubMed
- Bates DW, Cohen M, Leape LL, Overhage JM, Shabot MM, Sheridan T. Reducing the frequency of errors in medicine using information technology. J Am Med Inform Assoc. 2001;8(4):299–308. http://jamia.oxfordjournals.org/cgi/pmidlookup?view=long&pmid=11418536 - PMC - PubMed
- English M, Irimu G, Agweyu A, Gathara D, Oliwa J, Ayieko P, Were F, Paton C, Tunis S, Forrest CB. Building learning health systems to accelerate research and improve outcomes of clinical care in low- and middle-income countries. PLoS Med. 2016 Apr;13(4):e1001991. doi: 10.1371/journal.pmed.1001991. http://dx.plos.org/10.1371/journal.pmed.1001991 PMEDICINE-D-15-01349 - DOI - DOI - PMC - PubMed
- Friedman CP, Wong AK, Blumenthal D. Achieving a nationwide learning health system. Sci Transl Med. 2010 Nov 10;2(57):57cm29. doi: 10.1126/scitranslmed.3001456.2/57/57cm29 - DOI - PubMed
Related information
Grants and funding.
- WT_/Wellcome Trust/United Kingdom
- MR/N005600/1/MRC_/Medical Research Council/United Kingdom
- PDA/02/06/096/DH_/Department of Health/United Kingdom
- U54 GM115677/GM/NIGMS NIH HHS/United States
LinkOut - more resources
Full text sources.
- Europe PubMed Central
- JMIR Publications
- PubMed Central
Other Literature Sources
- scite Smart Citations
- Citation Manager
NCBI Literature Resources
MeSH PMC Bookshelf Disclaimer
The PubMed wordmark and PubMed logo are registered trademarks of the U.S. Department of Health and Human Services (HHS). Unauthorized use of these marks is strictly prohibited.
- Introduction
- Conclusions
- Article Information
CDW indicates clinical data warehouse.
Data Sharing Statement
See More About
Sign up for emails based on your interests, select your interests.
Customize your JAMA Network experience by selecting one or more topics from the list below.
- Academic Medicine
- Acid Base, Electrolytes, Fluids
- Allergy and Clinical Immunology
- American Indian or Alaska Natives
- Anesthesiology
- Anticoagulation
- Art and Images in Psychiatry
- Artificial Intelligence
- Assisted Reproduction
- Bleeding and Transfusion
- Caring for the Critically Ill Patient
- Challenges in Clinical Electrocardiography
- Climate and Health
- Climate Change
- Clinical Challenge
- Clinical Decision Support
- Clinical Implications of Basic Neuroscience
- Clinical Pharmacy and Pharmacology
- Complementary and Alternative Medicine
- Consensus Statements
- Coronavirus (COVID-19)
- Critical Care Medicine
- Cultural Competency
- Dental Medicine
- Dermatology
- Diabetes and Endocrinology
- Diagnostic Test Interpretation
- Drug Development
- Electronic Health Records
- Emergency Medicine
- End of Life, Hospice, Palliative Care
- Environmental Health
- Equity, Diversity, and Inclusion
- Facial Plastic Surgery
- Gastroenterology and Hepatology
- Genetics and Genomics
- Genomics and Precision Health
- Global Health
- Guide to Statistics and Methods
- Hair Disorders
- Health Care Delivery Models
- Health Care Economics, Insurance, Payment
- Health Care Quality
- Health Care Reform
- Health Care Safety
- Health Care Workforce
- Health Disparities
- Health Inequities
- Health Policy
- Health Systems Science
- History of Medicine
- Hypertension
- Images in Neurology
- Implementation Science
- Infectious Diseases
- Innovations in Health Care Delivery
- JAMA Infographic
- Law and Medicine
- Leading Change
- Less is More
- LGBTQIA Medicine
- Lifestyle Behaviors
- Medical Coding
- Medical Devices and Equipment
- Medical Education
- Medical Education and Training
- Medical Journals and Publishing
- Mobile Health and Telemedicine
- Narrative Medicine
- Neuroscience and Psychiatry
- Notable Notes
- Nutrition, Obesity, Exercise
- Obstetrics and Gynecology
- Occupational Health
- Ophthalmology
- Orthopedics
- Otolaryngology
- Pain Medicine
- Palliative Care
- Pathology and Laboratory Medicine
- Patient Care
- Patient Information
- Performance Improvement
- Performance Measures
- Perioperative Care and Consultation
- Pharmacoeconomics
- Pharmacoepidemiology
- Pharmacogenetics
- Pharmacy and Clinical Pharmacology
- Physical Medicine and Rehabilitation
- Physical Therapy
- Physician Leadership
- Population Health
- Primary Care
- Professional Well-being
- Professionalism
- Psychiatry and Behavioral Health
- Public Health
- Pulmonary Medicine
- Regulatory Agencies
- Reproductive Health
- Research, Methods, Statistics
- Resuscitation
- Rheumatology
- Risk Management
- Scientific Discovery and the Future of Medicine
- Shared Decision Making and Communication
- Sleep Medicine
- Sports Medicine
- Stem Cell Transplantation
- Substance Use and Addiction Medicine
- Surgical Innovation
- Surgical Pearls
- Teachable Moment
- Technology and Finance
- The Art of JAMA
- The Arts and Medicine
- The Rational Clinical Examination
- Tobacco and e-Cigarettes
- Translational Medicine
- Trauma and Injury
- Treatment Adherence
- Ultrasonography
- Users' Guide to the Medical Literature
- Vaccination
- Venous Thromboembolism
- Veterans Health
- Women's Health
- Workflow and Process
- Wound Care, Infection, Healing
Get the latest research based on your areas of interest.
Others also liked.
- Download PDF
- X Facebook More LinkedIn
Mermin-Bunnell K , Zhu Y , Hornback A, et al. Use of Natural Language Processing of Patient-Initiated Electronic Health Record Messages to Identify Patients With COVID-19 Infection. JAMA Netw Open. 2023;6(7):e2322299. doi:10.1001/jamanetworkopen.2023.22299
Manage citations:
© 2024
- Permissions
Use of Natural Language Processing of Patient-Initiated Electronic Health Record Messages to Identify Patients With COVID-19 Infection
- 1 Currently a medical student at Emory University School of Medicine, Atlanta, Georgia
- 2 School of Electrical and Computer Engineering, Georgia Institute of Technology, Atlanta
- 3 School of Computational Science and Engineering, Georgia Institute of Technology, Atlanta
- 4 Division of Infectious Diseases, Emory University School of Medicine, Atlanta, Georgia
- 5 Division of General Internal Medicine, Emory University School of Medicine, Atlanta, Georgia
- 6 Department of Biomedical Informatics, Emory University School of Medicine, Atlanta, Georgia
- 7 Emory University School of Medicine, Atlanta, Georgia
- 8 Atlanta Veterans Affairs Healthcare System, Decatur, Georgia
- 9 Department of Biomedical Engineering, Georgia Institute of Technology and Emory University, Atlanta, Georgia
Question Can a natural language processing (NLP) model accurately classify patient-initiated electronic health record (EHR) messages and triage positive COVID-19 cases?
Findings In this cohort study of 10 172 patients, 3048 messages reported COVID-19–positive test results, and the mean (SD) message response time for patients who received treatment (364.10 [784.47] minutes) was faster than for those who did not (490.38 [1132.14] minutes). This novel NLP model classified patient messages with 94% accuracy and a sensitivity of 85% for messages that mentioned confirmed COVID-19 infection, discussed COVID-19 without mentioning a positive test result, or were unrelated to COVID-19.
Meaning These findings suggest that NLP-EHR integration can effectively triage patients reporting positive at-home COVID-19 test results via the EHR, reducing the time to first message response and increasing the likelihood of receiving an antiviral prescription within the 5-day treatment window.
Importance Natural language processing (NLP) has the potential to enable faster treatment access by reducing clinician response time and improving electronic health record (EHR) efficiency.
Objective To develop an NLP model that can accurately classify patient-initiated EHR messages and triage COVID-19 cases to reduce clinician response time and improve access to antiviral treatment.
Design, Setting, and Participants This retrospective cohort study assessed development of a novel NLP framework to classify patient-initiated EHR messages and subsequently evaluate the model’s accuracy. Included patients sent messages via the EHR patient portal from 5 Atlanta, Georgia, hospitals between March 30 and September 1, 2022. Assessment of the model’s accuracy consisted of manual review of message contents to confirm the classification label by a team of physicians, nurses, and medical students, followed by retrospective propensity score–matched clinical outcomes analysis.
Exposure Prescription of antiviral treatment for COVID-19.
Main Outcomes and Measures The 2 primary outcomes were (1) physician-validated evaluation of the NLP model’s message classification accuracy and (2) analysis of the model’s potential clinical effect via increased patient access to treatment. The model classified messages into COVID-19–other (pertaining to COVID-19 but not reporting a positive test), COVID-19-positive (reporting a positive at-home COVID-19 test result), and non–COVID-19 (not pertaining to COVID-19).
Results Among 10 172 patients whose messages were included in analyses, the mean (SD) age was 58 (17) years; 6509 patients (64.0%) were women and 3663 (36.0%) were men. In terms of race and ethnicity, 2544 patients (25.0%) were African American or Black, 20 (0.2%) were American Indian or Alaska Native, 1508 (14.8%) were Asian, 28 (0.3%) were Native Hawaiian or other Pacific Islander, 5980 (58.8%) were White, 91 (0.9%) were more than 1 race or ethnicity, and 1 (0.01%) chose not to answer. The NLP model had high accuracy and sensitivity, with a macro F1 score of 94% and sensitivity of 85% for COVID-19–other, 96% for COVID-19–positive, and 100% for non–COVID-19 messages. Among the 3048 patient-generated messages reporting positive SARS-CoV-2 test results, 2982 (97.8%) were not documented in structured EHR data. Mean (SD) message response time for COVID-19–positive patients who received treatment (364.10 [784.47] minutes) was faster than for those who did not (490.38 [1132.14] minutes; P = .03). Likelihood of antiviral prescription was inversely correlated with message response time (odds ratio, 0.99 [95% CI, 0.98-1.00]; P = .003).
Conclusions and Relevance In this cohort study of 2982 COVID-19–positive patients, a novel NLP model classified patient-initiated EHR messages reporting positive COVID-19 test results with high sensitivity. Furthermore, when responses to patient messages occurred faster, patients were more likely to receive antiviral medical prescription within the 5-day treatment window. Although additional analysis on the effect on clinical outcomes is needed, these findings represent a possible use case for integration of NLP algorithms into clinical care.
The emergence and rapid spread of SARS-CoV-2 and subsequent variants has posed a unique threat to health care capacity worldwide. The COVID-19 pandemic has increased reliance on telemedicine and electronic health record (EHR) communications, as patient-initiated EHR message rates increased over 200%. 1 Electronic messaging has the potential to improve clinician-patient communication, but high message volumes can impair efficiency and impose burdens on clinicians already experiencing burnout, ultimately resulting in patient morbidity from delayed responses. 2
At-home rapid SARS-CoV-2 testing offers near-immediate results, increases testing access, and facilitates implementation of appropriate isolation measures without exposing clinicians and other patients. 3 Many patients report positive SARS-CoV-2 test results to clinicians via patient-initiated EHR messages. When message responses are delayed due to increased EHR message burden, treatment windows may be missed. 4 - 6 Oral antiviral therapies may decrease hospitalization, long-term sequelae, and death but have only been studied and authorized for use within 5 days of symptom onset. 3 , 7 There are few examples of natural language processing (NLP) analysis of patient messages for use in clinical care. 8 , 9
We developed an artificial intelligence solution to EHR message burden that rapidly identifies patient-authored messages reporting positive SARS-CoV-2 test results with the aim of facilitating timely administration of oral antiviral treatment. In a retrospective cohort analysis, we assessed whether there was an association between the time from when a patient sent a message reporting a positive test result to when their message was first opened by a member of their clinical team and whether the patient received antiviral treatment.
In this cohort study, data from 5 Atlanta, Georgia, hospitals was approved by Emory University’s Institutional Review Board. The requirement for informed consent was waived based on negligible risk to patients and impracticality in obtaining consent from tens of thousands of patients.
We obtained internally completed SARS-CoV-2 polymerase chain reaction and rapid antigen test results and recorded external test results from Emory’s clinical data warehouse on a flow sheet via an honest broker informatician. A total of 187 217 messages sent by adult patients to their health care teams via a patient portal between March 30 and September 1, 2022, were analyzed. Race and ethnicity data were extracted from the EHR to characterize the study population and reduce uncontrolled confounding in model development and analysis. To train the model, a random sample of 14 537 messages were categorized by study clinicians as (1) mentioning confirmed COVID-19 infection (COVID-19 positive), (2) discussing COVID-19 without mention of a positive test result (COVID-19 other), or (3) unrelated to COVID-19 (non–COVID-19). We used transfer learning with a Bidirectional Encoding Representations of Transformers (BERT) NLP model 10 to classify messages. The model, eCOV, was trained and validated on 14 537 clinician-labeled messages with a train to validation to testing ratio of 6:2:2; 5-fold cross-validation was applied to evaluate the performance across all samples in the testing set. 11 All data were obtained and reported in concordance with the Strengthening the Reporting of Observational Studies in Epidemiology ( STROBE ) reporting guideline.
Several BERT models were deployed and tested, including the base BERT model, 10 Bio ClinicalBERT, 12 and distilBERT. 13 The Bio ClinicalBERT model was initialized from the BioBERT model and pretrained on MIMIC-III (Medical Information Mart for Intensive Care III) clinical notes; the model shows performance superior to that of general-purpose embeddings on several clinical NLP tasks, such as the MedNLI (natural language inference) task annotated by physicians, nurses, and medical students and named entity recognition tasks. Meanwhile, distilBERT, a lightweight transformer model distilled from the base BERT model, preserved 95% performance in language understanding tasks with 40% fewer parameters and 60% faster computation. Using all 3 models yielded very similar classification performance. The language in messages from patients is similar to general-purpose text and very different from clinical notes; thus, the Bio ClinicalBERT model fails to show better classification performance than distilBERT for EHR text written by patients. Overall, the distilBERT model was found to be optimal for this specific application due to faster and more efficient computation.
To test the robustness of the distilBERT model, we conducted experiments to evaluate its classification performance with limited message samples. We divided the data set into a training and validation set of 10 000 samples and a holdout testing set of 4537 samples. We randomly extracted 1000 to 10 000 samples from the training and validation sets, with a 6:2 ratio of training to validation. We trained and evaluated each model on the holdout testing set. The plot shows that more training and validation samples led to better results, but even with just 1000 samples, the model achieved competitive performance. This finding is important because it suggests that clinicians can label a small number of messages and still achieve reasonable results when using the same NLP approach in a new hospital or for a new clinical task.
After clinical care for the episode in question had ceased, SQL (structured query language) queries were used to extract EHR data. Matching to chronic conditions, demographics, and medications, for example, was accomplished via matching of unique identifiers and was date matched to ensure medications and conditions were active during the clinical period of interest.
Once trained, the eCOV model’s 3-label classifier was prospectively deployed on 2907 new messages to evaluate accuracy. The first message reporting positive results for each patient was included; subsequent messages describing positive results within the study time frame were ignored.
A subset of messages representing patients self-reporting positive results was adjudicated by clinicians to determine whether the message was sent within the time frame of effectiveness for oral antivirals. 14 The 2 primary outcomes were (1) physician-validated evaluation of the NLP model’s message classification accuracy and (2) analysis of the model’s potential clinical outcomes via increased patient access to treatment. Candidates without documented prescriptions for nirmatrelvir and ritonavir or for molnupiravir were identified as untreated, and those who had a prescription for nirmatrelvir and ritonavir or for molnupiravir within 5 days of the message creation date were identified as treated. Propensity score matching was performed to create similar classes to control for confounders. Logistic regression was performed on the COVID-19–positive group with the binary outcome of receiving or not receiving antiviral treatments in the 5-day window with covariates age, sex, White race, Charlson Comorbidity Index (CCI) score, and body mass index (BMI), similar to prior analyses of monoclonal antibody recipients. 15 White race was a binary variable in this analysis because it was the most common race among the patient population.
Clinician-assigned labels were compared with model-assigned labels to calculate the class-specific and weighted to unweighted mean of sensitivity (recall), specificity, precision, and F1 score. True-positive (TP), false-negative (FN), true-negative (TN), and false-positive (FP) labels were calculated for each class using a one-vs-rest approach. Class-specific sensitivity (recall) was calculated as TP/(TP + FN) and measured the proportion of patients with COVID-19 who were labeled as COVID-19 positive by the model. Class-specific specificity was calculated as TN/(TN + FP) and measured the proportion of patients without COVID-19 who were labeled as COVID-19 negative by the model. Class-specific precision was calculated as TP/(TP + FP) and measured the indications denoting, among all samples with positive test results, how many truly belonged to the target class. Class-specific F1 score was calculated as 2TP/(2TP + FP + FN). The F1 score is the arithmetic mean of precision and recall and is a balanced summarization metric of NLP model performance. 16 , 17 The unweighted mean (macro mean) of each performance metric is the arithmetic mean of the above performance metrics across all classes, ignoring the number of samples of each class. The weighted mean of each performance metric includes the number of samples (support) in each class in calculation.
Time from message creation to first staff interaction, defined as the time when a clinical staff member first viewed the message, was calculated using clinical data warehouse timestamps. A paired t test was used to compare time to first message interaction between untreated and treated patients. The Figure depicts a flowchart of study methods, including NLP model training and evaluation as well as cohort study analyses. Two-sided α = .05 indicated statistical significance.
Of the 10 172 patients whose messages were included in analyses, the mean (SD) age was 58 (17) years; 6509 (64.0%) were women and 3663 (36.0%) were men. In terms of race and ethnicity, 2544 patients (25.0%) were African American or Black, 20 (0.2%) were American Indian or Alaska Native, 1508 (14.8%) were Asian, 28 (0.3%) were Native Hawaiian or other Pacific Islander, 5980 (58.8%) were White, 91 (0.9%) were more than 1 race or ethnicity, and 1 (0.01%) chose not to answer. Table 1 gives complete demographic information. Overall, 3048 of 187 217 incoming patient-generated messages (1.6%) were related to self-reported positive SARS-CoV-2 test results and, of those, 2982 (97.8%) were not otherwise documented in structured EHR data. An evaluation of 50 patients’ external pharmacy records in the EHR showed concordance between internal and external antiviral prescriptions in 45 (90.0%) of the 50 patients examined. The 3048 patients who sent messages reporting COVID-19 diagnoses that were not recorded in structured EHR data were evaluated in retrospective cohort analysis. The eCOV model’s macro F1 score was 94%, with individual class F1 scores of 87% for COVID-19–other, 95% for COVID-19–positive, and 100% for non–COVID-19 messages. Sensitivities were 85% for COVID-19–other, 96% for COVID-19–positive, and 100% for non–COVID-19 messages ( Table 2 ).
The treated and untreated groups had significant differences in age, BMI, and CCI ( Table 3 ). Mean (SD) response time was significantly faster for the treated group (364.10 [784.47] minutes) than the untreated group (490.38 [1132.14] minutes; P = .03). Logistic regression showed an association between shorter response time and likelihood of receiving an antiviral prescription (odds ratio, 0.99 [95% CI, 0.98-1.00]; P = .003) when controlling for age, BMI, CCI, male sex, and White race ( Table 4 ).
Electronic health record messaging offers convenient patient self-reporting of COVID-19 test results to clinicians, but high message volumes, demands on clinician time, and lack of documentation into structured EHR data elements are barriers to timely treatment. The eCOV model is the first clinically deployable NLP model to triage incoming, self-reported positive COVID-19 cases in real time. The eCOV model identified acute COVID-19 cases from patient-initiated EHR messages with 94% sensitivity.
For large health care systems receiving thousands of messages daily, the only viable option for triaging clinically urgent messages is a technological one. Whether a physician acts on a message the day it is sent or multiple days later might determine whether oral antiviral treatment can be appropriately administered and whether benefits of reduced risk of hospitalization or mortality can be realized. 18 This remains critically important as concurrent COVID-19, influenza, and respiratory syncytial virus epidemics threaten hospital capacity. This method also allows for more accurate reporting of positivity, as only 66 cases identified in our cohort (2.2%) were otherwise documented in the structured data elements more easily accessed via traditional EHR database queries.
The availability and convenience of at-home SARS-CoV-2 tests makes them an attractive option for patients. Patients with positive test results for COVID-19 at home and who experience severe symptoms can then send an EHR message to their primary care physician reporting the positive test results and inquiring about treatment options. Some patients can then be adequately treated remotely by their primary care physician, reducing hospitalization rates, infection risk for health care workers, and burden on the medical care system. However, this requires initiation of treatment in a timely fashion, and the large burden of EHR tasks, including patient-initiated messages, makes it difficult for physicians to rapidly respond to all patient messages, many of which are not urgent. Patients whose messages have slower response times are less likely to receive an antiviral prescription within the 5-day treatment initiation window. The eCOV model accurately and instantaneously identifies and triages patient-initiated messages reporting positive COVID-19 test results. By classifying patient messages accurately and improving the speed of treatment access, NLP, when integrated into the EHR, has the potential to improve clinical outcomes while simultaneously reducing health care system burden. Additional analyses on outcomes following clinical integration are needed to quantify true clinical impact.
We found an association between longer response time and absence of antiviral prescription. Although propensity score matching was performed to create similar classes, factors such as age and preexisting medical conditions may contribute to the decision to prescribe antivirals independent of response time. Additionally, some patients may have received treatments other than oral antivirals that were not captured, such as monoclonal antibody therapies or intravenous remdesivir, during the treatment window. There are a multitude of factors that contribute to treatment decisions—including patient preference, vaccination status, insurance status, and cost of medications—that were not captured in this analysis. Further research is required to determine a causal relationship between response time and treatment. Additional limitations of this study include absence of visual validation of test results and inability to systematically verify prescriptions from other facilities or treatment adherence. However, underreporting rates likely significantly outweigh false-positive reporting. 3 , 19 Misclassification based on external prescription of antivirals not identified by our database is possible but likely small based on evaluation of patients’ external pharmacy records in the EHR, which showed concordance between internal and external prescriptions in 45 (90.0%) of 50 patients examined. Additionally, symptom onset could not be systematically evaluated, so a subset of patients identified as antiviral candidates may have been out of the treatment window by the time a message was sent.
The findings of this cohort study suggest that nearly 98% of patients reporting their positive at-home COVID-19 test results were not otherwise documented as SARS-CoV-2 positive in the EHR. These represent opportunities for patients whose messages are at risk of being missed among high volumes of EHR communications to receive oral antivirals within the 5-day treatment window. The eCOV model uses NLP to classify messages with high accuracy and can be deployed as an automated triage tool to facilitate timely identification of treatment candidates. Although additional analysis of the effect on clinical outcomes is needed, these findings represent a possible use case for integration of NLP algorithms into clinical care.
Accepted for Publication: May 19, 2023.
Published: July 7, 2023. doi:10.1001/jamanetworkopen.2023.22299
Open Access: This is an open access article distributed under the terms of the CC-BY License . © 2023 Mermin-Bunnell K et al. JAMA Network Open .
Corresponding Author: Blake Anderson, MD, Division of General Internal Medicine, Emory University School of Medicine, 2200 Peachtree Rd NW, Atlanta, GA 30309 ( [email protected] ).
Author Contributions: Dr Anderson had full access to all of the data in the study and takes responsibility for the integrity of the data and the accuracy of the data analysis.
Concept and design: Mermin-Bunnell, Zhu, Damhorst, Johnson, Anderson.
Acquisition, analysis, or interpretation of data: Zhu, Hornback, Walker, Robichaux, Mathew, Jaquemet, Peters, Wang, Anderson.
Drafting of the manuscript: Mermin-Bunnell, Zhu, Mathew, Jaquemet, Peters, Anderson.
Critical revision of the manuscript for important intellectual content: Mermin-Bunnell, Zhu, Hornback, Damhorst, Walker, Robichaux, Jaquemet, Johnson, Wang, Anderson.
Statistical analysis: Zhu, Hornback, Robichaux, Mathew, Anderson.
Administrative, technical, or material support: Mermin-Bunnell, Zhu, Mathew, Jaquemet, Peters, Johnson, Wang, Anderson.
Supervision: Mermin-Bunnell, Walker, Johnson, Wang, Anderson.
Conflict of Interest Disclosures: Mr Zhu reported receiving grant funding from the Georgia Institute of Technology during the conduct of the study. Prof Johnson reported receiving grant funding from Dexcom, consulting for Easai Co Ltd, and personal fees for authorship of the nocturia card from UpToDate unrelated to the current project. Dr Wang reported receiving a collaborative grant from Emory University, faculty fellow awards from the Georgia Institute of Technology, a National Science Foundation travel grant funding to support students attending a biomedical informatics conference, and grant funding from the Imlay Foundation and Enduring Hearts Foundation outside the submitted work and serving as an advisor to Switchboard startup. Dr Anderson reported receiving a collaborative grant from Emory University and intellectual property at stake for himself and his institution around this use of transformer architecture natural language processing, which will likely include payment in the future. No other disclosures were reported.
Funding/Support: This research was supported by COVID-19 CURE Award 00103333 from the O. Wayne Rollins Foundation and the William Randolph Hearst Foundation.
Role of the Funder/Sponsor: The sponsors had no role in the design and conduct of the study; collection, management, analysis, and interpretation of the data; preparation, review, or approval of the manuscript; and decision to submit the manuscript for publication.
Data Sharing Statement: See the Supplement .
- Register for email alerts with links to free full-text articles
- Access PDFs of free articles
- Manage your interests
- Save searches and receive search alerts
- Research article
- Open access
- Published: 31 December 2012
We are bitter, but we are better off: case study of the implementation of an electronic health record system into a mental health hospital in England
- Amirhossein Takian 1 , 3 ,
- Aziz Sheikh 2 &
- Nicholas Barber 3
BMC Health Services Research volume 12 , Article number: 484 ( 2012 ) Cite this article
14k Accesses
54 Citations
3 Altmetric
Metrics details
In contrast to the acute hospital sector, there have been relatively few implementations of integrated electronic health record (EHR) systems into specialist mental health settings. The National Programme for Information Technology (NPfIT) in England was the most expensive IT-based transformation of public services ever undertaken, which aimed amongst other things, to implement integrated EHR systems into mental health hospitals. This paper describes the arrival, the process of implementation, stakeholders’ experiences and the local consequences of the implementation of an EHR system into a mental health hospital.
Longitudinal, real-time, case study-based evaluation of the implementation and adoption of an EHR software (RiO) into an English mental health hospital known here as Beta. We conducted 48 in-depth interviews with a wide range of internal and external stakeholders, undertook 26 hours of on-site observations, and obtained 65 sets of relevant documents from various types relating to Beta. Analysis was both inductive and deductive, the latter being informed by the ‘sociotechnical changing’ theoretical framework.
Many interviewees perceived the implementation of the EHR system as challenging and cumbersome. During the early stages of the implementation, some clinicians felt that using the software was time-consuming leading to the conclusion that the EHR was not fit for purpose. Most interviewees considered the chain of deployment of the EHR–which was imposed by NPfIT–as bureaucratic and obstructive, which restricted customization and as a result limited adoption and use. The low IT literacy among users at Beta was a further barrier to the implementation of the EHR. This along with inadequate training in using the EHR software led to resistance to the significant cultural and work environment changes initiated by EHR. Despite the many challenges, Beta achieved some early positive results. These included: the ability to check progress notes and monitor staff activities; improving quality of care as a result of real-time, more accurate and shared patient records across the hospital; and potentially improving the safety of care through increasing the legibility of the clinical record.
Conclusions
Notwithstanding what was seen as a turbulent, painful and troublesome implementation of the EHR system, Beta achieved some early clinical and managerial benefits from implementing EHRs. The ‘sociotechnical changing’ framework helped us go beyond the dichotomy of success versus failure, when conducting the evaluation and interpreting findings. Given the scope for continued development, there are good reasons, we argue, to scale up the intake of EHR systems by mental health care settings. Software customization and appropriate support are essential to work EHR out in such organizations.
Peer Review reports
Provision of mental health services often involves professionals located in disparate locations. It has been suggested that the use of integrated electronic health record (EHR) systems (a digital longitudinal record of a citizen’s health and healthcare interventions that can be accessed by healthcare providers from across a defined range of healthcare settings) will help to improve the quality of care for mental health patients [ 1 , 2 ] through, for example, preventing loss of records, increasing accessibility of the records, improving medication management, reducing medical errors and costs [ 3 – 5 ], and empowering patients through greater engagement in their care provision [ 6 ].
Despite these anticipated benefits, few implementations of integrated EHR systems have taken place across mental health settings [ 7 , 8 ]. Possible reasons for this lack of progress include: sensitivities in relation to the potential for stigma and discrimination associated with the unambiguous recording of diagnoses in medical records [ 9 ]; healthcare professionals’ reluctance to use EHRs [ 10 ]; concerns about lowering productivity and inaccurate clinical notes [ 11 ]; data security and confidentiality due to very sensitive and specific nature of mental health [ 12 , 13 ]; concerns about the quality of patient-provider relationship [ 14 , 15 ]; and considerably lower frequency of self-determination in decisions about mental health, compared to acute settings [ 16 ]. As a result, little has been published on the evaluation of the implementation of EHRs in the context of mental health [ 8 ].
Launched in 2002 and officially dismantled in 2011 [ 17 , 18 ], the National Programme for Information Technology (NPfIT) included the first sustained national attempt to introduce centrally-procured EHR systems across the National Health Service’s (NHS) hospitals [ 19 , 20 ], including mental health settings [ 21 – 23 ]. We conducted the first national evaluation of implementation and adoption of EHR systems in NHS ‘early adopter’ hospitals and have reported on this in detail elsewhere [ 24 , 25 ]. Here, we report on a case study of the implementation of an EHR (RiO) into a mental health setting delivered though the NPfIT and analyzed using our adapted ‘sociotechnical changing framework’ (see below for more elaboration). We investigated the arrival and the implementation process, stakeholders’ experiences and perceptions, and local consequences of adopting this nationally procured ‘off the shelf’ EHR software that, in various forms, had been used for up to 15 years in a few mental health and community centres in the UK.
Ethical considerations
Our research was classified as a service evaluation (ref. 08/H0703/112). We obtained informed consent from the participating hospital and individuals and guaranteed their anonymity.
Design, sampling and data collection
This was a prospective, longitudinal [ 26 ], sociotechnical [ 27 ], and real-time case study-based evaluation [ 28 ]. Beta was selected from a purposive sample [ 29 ] of 12 diverse, NHS ‘early adopter’ hospitals studied over a 30-month period from September 2008 until February 2011. This paper focuses in-depth on one of these sites (Beta), in which the lead researcher (AT) collected a broad range of qualitative data (see Table 1 for characteristics of Beta and the dataset). We conceptualized Beta as an independent case study to reflect the importance of local contingencies [ 30 ]. This allowed the specific character of the implementation and adoption of the EHR software to be revealed, whilst attempting to make general inferences transferable to other contexts [ 31 , 32 ].
Within Beta, we purposefully (and at times opportunistically) conducted semi-structured interviews with a diverse range of stakeholders with broad range of perspectives, from inside and outside of the hospitals (see Table 1 for more details). We developed generic interview guides that were then tailored for specific participants [See Additional files 1 , 2 and 3 ]. The majority of interviews were audio-recorded and transcribed verbatim. Interviews were complemented by the researcher’s field notes, as well as observational and documentary data of various types with regard to planning, implementing, and using EHR systems at Beta. The opportunity to triangulate between these data sources enhanced our understanding of the evolving process of implementation.
Data analysis
Data analysis was an iterative process. We followed a two-step approach: initially, at Beta case study level using a combination of deductive and inductive approaches [ 33 , 34 ], and then a meta-synthesis that drew upon the analytical themes from other case studies, which were predominantly in acute hospitals. We used an adapted sociotechnical framework for data organiziation and classification of findings [ 27 ], and ‘sociotechnical changing perspective’ [ 35 ] for data analysis and interpretation, which we discuss below. Further, we presented primary findings from each case study through two complementary fora: regular analysis workshops with the wider evaluation team and formative feedback sessions with hospital representatives. This helped with validating the case study findings and furthermore integrating findings with our broader evaluation, enabling us to draw out some transferable findings.
Our theoretical perspective: ‘sociotechnical changing’
Most EHR evaluations have drawn upon a broadly positivist ontology with a view to making causal inferences about the effectiveness of EHRs [ 30 ]. In the context of our evaluation, it was however not possible to ‘control’ for contextual factors using standard experimental or quazi-experimental designs. Moreover, it was also important for us to be cognizant of the fact that both the social and technical dimensions of EHR had the potential to shape each other over time in the complex and evolving environment of healthcare settings [ 36 ]. A number of theoretical frameworks have been developed and deployed to study this reciprocal relationship between EHR and the organization including: the role of leadership and envisioning the implementation of EHR as change management [ 37 ]; engaging with various groups of stakeholders [ 38 ], taking wider social context into consideration [ 39 ]; integration of EHR with workflows and care pathways [ 40 ]; organizational culture and behaviour [ 41 ]; and the ever-evolving contextual flux [ 42 ]. Drawing on these theoretical prepositions, Aarts et al. (2004) [ 43 ] has highlighted three dimensions to understanding the implementation of EHR: the interrelation of the organizational environment and the technology; a constantly changing mileu of the organization and environement: “emergent change”; and the interaction between system’s functioning, the organization’s needs and working patterns: the sociotechnical approach [ 44 ].
Our approach [ 30 ], pursued over the course of a 30 month longitudinal evaluation of national EHR systems in the English hospitals [ 21 , 24 ], underscored the emerging nature of change and its characteristics when evaluating EHRs. We refer to such change, as it happens as ‘changing’ (present participle) [ 35 ]. This implies that the EHR software, the clinical practice, the care giving, the organizational structures, and the carriers of institutional and professional norms were all in a state of flux, moving from somewhere now lost in the past to somewhere in the uncertain future: ‘becoming’ [ 45 ]. We therefore focused on the activity ‘in between’, the period of implementing EHR during which things were changing, rather than some predicted state of achieved change.
In our analysis, we adopted a social construction [ 46 ] and performative view [ 30 ]. We applied this performative view to explore how a diverse set of stakeholders [ 47 ] performed to make sense of new circumstances under EHR and make it work [ 42 , 48 ], enabled and constrained, as they were by their own skills, attitudes and the various technologies and other resources available to them. We sought to explain how stakeholders’ understandings and actions shaped adoption or non-adoption of RiO at Beta [ 49 , 50 ]. This is what we call the notion of “working-out” to signify a dynamic process of change over time that involved the ensemble of people, existing and emerging work practices and tools, individuals and organizational beliefs, assumptions, and expectations [ 48 , 51 , 52 ], which can be understood as both cause and consequence of longer-term processes of changing [ 53 ].
We sought to explore, understand and narrate the stories of EHR “in-the-making” [ 54 , 55 ]. Thus, we were less concerned with assessing the progress or achievements of implementing EHR systems measured against predefined criteria of success or failure, expectations and project milestones [ 43 ]; rather, drawing on the principles of Actor Network Theory (ANT) [ 54 , 56 ], and other studies on impact of perceptions of EHR systems on the implementation [e.g. [ 52 ]], our focus was on exploring what people understood about EHR (perceptions, hopes, fears) and what they actually did in their day to day practices (uses and practice) to ‘make it work’.
We report on four main findings. First, we describe the arrival of EHR at Beta through reflecting on the underlying reasons that led Beta to decide to implement RiO. Second, we provide an overview of the process of implementation through describing the management strategies pursued by Beta in implementing the software. Third, we consider the experiences, perceptions, and attitudes of users with RiO software. Finally, the local consequences, including some early benefits of the EHR software, realized by users at Beta will be described.
The arrival of EHR at Beta
Compared to acute hospitals, many of which traditionally used some sort of computerized systems to manage or deliver patient care, mental health settings in England at the time of this study typically lacked any ‘joined up’ electronic information system. It has been suggested that there is ‘an intrinsic lack of interest in information systems among many staff in mental health’ [ 57 ]. It is therefore perhaps unsurprising that computerized patient administration system (PAS) consisting of basic patient demographics, with little or no clinical functionality, had hitherto been the dominant form of electronic records in mental health settings. The organization of mental health Trusts (the administrative unit in England, which can include one or more hospital or clinic) involves a close working relationship with primary care and social services to manage a range of often complex cases involving several stakeholders. Episodes of care in mental health hospitals typically last longer than in acute settings, on occasions up to several years. Record keeping is also very different from the approach used in the acute sectors as the notes tend in mental health settings to be more narrative in nature. Paper record systems were the standard method of record keeping in mental health settings, these offering the advantages of being self-contained, (manually) transferable between clinical locations and well suited to narrative-based recording of clinical entries [ 58 ]. Consultations also tend to last longer (about an hour) and consequently notes tend to be very long:
" “I suppose our note keeping is very different because it’s therapeutic so we’re writing an hour session where it’s just based on talking to somebody so our notes are a lot longer they have a lot more detail…” (Nurse)."
NPfIT initiated the introduction of integrated EHR software with clinical functionality to mental health hospitals. The financial benefits of the hospital being part of NPfIT, i.e. virtually free software and support for early adopter hospitals (up until 2015), as well as RiO’s ability to connect to the national Spine [ 1 ] led Beta to implement the EHR software procured by NPfIT:
"“ You couldn’t really say how long it [the legacy PAS software] would develop or be in existence with the bigger systems coming in. We had to take a view then what is our strategy through to 2015 on this to get that developed” (Manager)."
Beta perceived the deployment of a modern clinically-oriented EHR as an essential step to maintain Foundation Hospital status (which would provide greater financial independence from the central NHS). EHR was seen as an opportunity to strengthen the information technology (IT) at Beta:
" “We’ve probably invested in the structure around this project [EHR] more than we’ve invested in anything else. We’ve employed a lot of external people to come in and roll this project out and quite a lot of investment in rolling it out. This is pivotal to improving our IT capability” (Doctor)."
Beta perceived the NPfIT-procured EHR as a potential enabler, facilitating integration of its care services with other care settings in their region and nationally. Moreover, as an ‘early adopter’ of NPfIT-supported EHR software, Beta was financially incentivized to implement RiO.
The process of implementing EHR
Implementation of RiO system into Beta proved challenging, the key issues encountered are described below.
Management of the implementation (the process of changing)
Beta followed an incremental approach in implementing RiO. This was crucial in order to connect the main hospital and a number of community centres at Beta that were physically dispersed. RiO version 5.1, which was implemented in Beta, was the first version with connectivity to the Spine (NHS national database and messaging service). On the basis of various services and physical sites within the organization, Beta divided the implementation into three distinct phases. The hospital adopted a ‘big bang’ approach within each deployment phase, in which most services went live and were migrated to RiO simultaneously, over a single night. External stakeholders (interviewees who were not employed by Beta) stated that Beta managed to plan well and put reasonable infrastructure in place for implementing EHR; this had positive effects on the experience of EHR implementation:
" “I’m very impressed here [Beta]. They understand the importance of RiO and implementing it correctly and I was surprised to see the seriousness that they’ve taken the project and therefore, the amount of resources that are allocated to it. That is the reason that I’m confident that RiO will be implemented successfully and the Trust will benefit from it” (NPfIT)."
RiO was not however linked to the local authority databases specified for social care services. This proved to be a major barrier to the integrated provision of mental and social care services in England, and led to data entry duplications, which in turn had adverse effects on users’ attitude towards the EHR:
" “You’d like to sort of knock their heads together and say, yes, it’s wonderful having a [city name] wide solution for mental health. Why didn’t anybody think of integrating social services into it? This is the problem that existed for a long time” (Manager)."
Beta used virtual databases for training purposes, which were criticized for not being rooted in users’ needs or their actual work practices:
" “Training was not useful for what we needed to know to be able to do our jobs. It didn’t tie in our processes and PIs [performance indicators] and things like that” (Manager)."
This led to “ when users actually go back onto their desk they realize, oh, I can’t remember this, and because their drop downs are totally different.” (Manager). Consequently, most interviewees preferred to learn through using the software in practice rather than in traditional classroom environments.
Beta mostly outsourced the implementation team responsible for putting RiO in practice with experienced people, who had deployed RiO in other settings. This was perceived to be a significant advantage:
" “I think key to our success here is that you get an individual in each of the work streams that has RiO knowledge and has gone through deployment and understands the problems” (IT Manager)."
Nevertheless, despite the advantages, there were two main disadvantages to this: high deployment costs; and challenges with employing a team predominantly comprising of temporary staff who were likely to be shed when returning to business as usual, with the potential for considerable loss of experience and expertise.
Users’ experiences with EHR software
In this section, we describe the experiences of EHR users at Beta and the ways that various users ‘worked out’ what EHR was and how to use it in their day to day practices. To begin with, some interviewees wished that the software was designed more around clinicians’ needs: “ It would be interesting to know how many medical types were involved in the setting up of it” (Nurse). RiO was described as “ too clumsy ” (Nurse), “quite an old looking tool” (Doctor), and not easy to navigate:
" “It is disappointing to have a clinical tool that is not as advanced as what I can do when I go and do my Internet shopping for my Tesco weekly shop” (Doctor) . "
RiO was thus seen as being unfit for at least some clinical purposes. Some described RiO as being designed on a simplistic, linear interpretation of the workflows in mental health settings. They thus criticized the software as not reflecting the contextual differences across care settings:
" “The reality is completely different. We see lots of different people. The type of contact we have with people is completely complex and it’s very variable. It varies from hospital to hospital how people deal with their clients. We had to adjust to that” (Care Manager)."
Although RiO was an established software that had existed in the British market for more than a decade, it was seen as lacking some essential assessment and clinical functionalities:
" “In old age psychiatry we use a Single Assessment Process (SAP) which isn’t on RiO at all. We are just going to have to continue using that as a Word document and uploading it. The core assessment is completely unsuitable for our use” (Doctor) . "
The degree of cultural change as a result of using EHR left some users feel uncomfortable:
" “It was quite a shock to not being able to do the things that we used to do on ward. For example, risk assessments used to be just typing little bits. We need to do everything in a different setting in RiO” (Nurse)."
On the one hand, some clinical functionalities of RiO including ICD (International Classification of Diseases) 10 coding for patients’ categorization were seen as “not perfect, because there are a number of diagnosis that aren’t coded. That’s all we’ve got and you have to fit people into it” (Doctor, Beta). This added to consultants’ workload because “ICD coding has to be a consultant that puts it in and can’t even be a junior doctor, so that’s just slightly irritating to me” (Doctor).
On the other hand, there were a few functionalities that users perceived as unnecessary and irritating:
" “You can’t delete out the bits that aren’t relevant, so you would have the whole document which includes things like forensic history and murder, which are perhaps not appropriate to an elderly person with some mild memory problems” (Doctor)."
Many interviewees perceived RiO as being incapable of meeting a number of important expectations of users. For instance, the very sensitive and distinct therapeutic nature of the relationship between patient and carer in mental health settings was not, it was suggested, appropriately considered by the designers of the RiO software. Given the very scattered distribution of community centres affiliated with mental health hospitals, it was perceived that the implementation team underestimated the practicalities of real-time data entry at the point of care:
"“ I need a clinical tool that I can sit here and stick my card in and look at the patient notes from my patient’s that are up the road at a different hospital and enter notes and read what the community worker has done and in their conversation with the family” (Doctor)."
Several practical issues with regard to the day-to-day use of RiO arose. For example, frustrations were reported because of staff being automatically logged-off if the software was not used for 30 minute. In addition, RiO was used on desktops in an office environment where the public had no access, so it was seen as “secure enough” to ensure confidentiality – however, Smartcards (a chip and PIN card that grants access to patient information based on healthcare practitioners’ work and level of involvement in patient care) were sometimes left in computers.
Logging-in and -out of multiple systems (including the legacy systems and the dataset for social care services which ran separately from RiO) was viewed a time consuming, unnecessary and cumbersome procedure, which “defeats the purpose” (Manager).
Depending on their position and responsibilities, staff used the EHR applications differently. For instance, senior psychiatrists used RiO less than their junior colleagues:
" “I probably use it [RiO] less than 10% of the time. It would be my junior doctor that’s inputting the information, not me. They probably use it 80% of their day” (Doctor)."
Other staff groups at Beta who had to enter data complained about increased administrative burden and, as a result, a reduction in time they had available to spend with patients.
The local consequences of EHR systems: early benefits of RiO at Beta
Despite challenges and difficulties, our evaluation revealed several perceptions of positive changes in work practices and patient care as a result of using EHR systems at Beta. These are described below.
Changing work environment
Generally, implementation of EHR systems brought the hospital the opportunity to strengthen its IT infrastructure:
" “RiO pushes IT to the front and just as important as clinical practices. Therefore, the Trust needs to have an on-going budget to be able to maintain their IT equipment and also look at advanced technologies that can go directly into RiO” (IT Manager)."
This enabled, some interviewees envisaged, the hospital to transform a number of its daily work dimensions towards more efficient services:
" “We are at the beginning of a big organizational change. People don’t have to travel backwards and forwards. They could just work outside of the office all the time. Therefore, you see more patients in a day. Your electronic record is up to date. It just goes on from there” (Manager)."
Data sharing was also perceived to be quicker, more transparent and secure, and timely through using RiO:
" “EHR brought in the standardization process in all the practices. That was very key. Because of the data warehouse on EHR, as long as the data is put into EHR, we have the ability to report on every single field” ( IT Manager)."
In addition, RiO was seen as contributing to the standardization of “ the context and structure of the letters that were sent to a patient, a GP or a carer” (IT Manager), which increased consistency of record keeping and enabled data extraction.
Patients’ care was also perceived to have become safer, for example, because of improved access to important information, and “the communication [with clinical colleagues] has improved a lot. There is just the reduction of clinical risk, the fact that information is available to all the clinicians who are involved in someone’s care” (Clinical Lead).
Users’ attitude about the consequences of EHR
The attitudes of clinical staff varied with regards to the consequences of EHR systems on their work practices. For instance:
" “The audio landscape in the office has changed. It’s not phones ringing and people talking. It’s the kind of sound of typing in the office, which is a little bit creepy” (Nurse)."
Within the early stages of implementing RiO, some users expressed their stress and anxiety, stating that they had less interaction with colleagues and spent more time than usual sitting in front of computers.
Interestingly, and in contrast to the experience with Smartcards noted above, most interviewees were confident that EHR provided a more confidential record of patient data than paper:
" “The system will track the person who is illegitimately looking at my record and figure out why they are entering it” (Nurse)."
Increasing quality of care and improving patient safety
As described earlier, although Beta did not have much flexibility for software configuration following the NPfIT chain of deployment, “that’s outweighed by we get a standardized build and overall I think that has been a great benefit to us” (Manager). Clinical users consistently praised the ability to see patients’ notes on RiO quickly, completely, in real-time and live across the whole hospital and affiliated community health centres:
" “I think a great benefit [of RiO] is being able to access records. What we’ll be able to do when we have RiO is access the notes of where they’ve been seen, wherever it was within the organization” (Doctor)."
This was seen as bringing direct benefits to the patients because:
" “Sometimes RiO makes things for our patients easier. Instead of waiting for me to write a referral form and then send in CPA b and then send in risk assessment and then wait for them to meet them up, they can go on RiO and have a look at the CPA. It’s actually speeding things up and it’s more reliable about information which is live” (Nurse)."
EHR systems were perceived as an enabler to keep mental health patients’ information safer than the previously used paper-based system, particularly when patients moved across care settings or got transferred:
" “I think it’s very easy for things to get missed when people are being transferred from one site to another. Another advantage is when you are looking at progress notes, you can filter them. If you are going to find something, you will be able to find it much more easily” (Doctor)."
EHR was also seen as making communications faster and more reliable because “you are not faxing and you are not saying, that fax machine not working. I’ll email it to you” (Nurse). Further, EHR was perceived to assist more careful and systematic monitoring and greater efficiency when utilizing resource across mental health hospitals and affiliated centres:
" “Users must outcome their appointments every two days after it’s been actually conducted. We know exactly if the patient was visited or if it was cancelled. In that sense, we can do it a lot of tracking and a lot of monitoring and better performance indicators” (IT Manager)."
The ability to check progress notes was seen by some as an opportunity to monitor staff activities and, if necessary, take remedial actions:
" “Because I have RiO I can actually go into the record myself and I can see what who has been discharged… I can see that that person from that team actually didn’t record it in their diary, so that’s why the report didn’t pick it up. Then I can flag it” (Manager)."
"Quality of care was also seen as being improved because the EHR “allowed us to look at our practice and make it more transparent. We have got so tight with doing everything correctly. We contact the GP and email the assessment out within three days. We want to make sure it’s done properly” (Care Manager)."
In addition, healthcare providers were less anxious about misunderstandings and mistakes about their planned orders for patients being carried out correctly:
" “When I’m on RiO, I’ll just quickly type in what I expect my nurse to do, at the time, rather than thinking, well, she’s got the notes so hopefully she’ll make the entry confirming what I’ve said ” (Doctor)."
Also, clinicians found the availability of information valuable:
" “I obviously get a lot of phone calls from patients, involved professionals and carers. You maybe don’t remember the exact and you don’t have the notes, immediately to hand. Obviously, now I’m on the phone, I’ll be able to tap in and get the details up [on RiO] and make any changes or suggestions, I can immediately type them in as well” (Doctor)."
This partly happened because users became more aware of the need to write patients’ notes more accurately:
" “I think now, with RiO it makes it all the more visible and people have to be more careful about how they write things” ( Manager)."
The greater visibility of health records led to reducing patients’ risk of poor treatment because of missing data or actions:
" “I have better quality of care now. There is nowhere to hide with EHR. If you didn’t put something down, it’s going to be missing and you can see straightaway” (Care Manger) . "
Given the very text-based environment of mental health, EHR was perceived to have improved patient safety by enhancing readability of patient notes:
" “The main thing really is that we can read people’s writing. That was a big thing before that you couldn’t actually read what people were writing in the NHS across the board” (Nurse)."
This advantage was more visible when staff were on leave and their assigned patients were taken care of by other members of the team:
" “The ward might have not put in the community slant of things on their ward notes. And then information would have gone amiss or they would have been delayed. Now I can just log in and have a look at the patient notes and I can see what the ward has entered ” (Nurse)." "Further, “using [electronic] records for other purposes like research is much easier now” (Doctor, Beta)." "All-in-all, the EHR was seen to have “played a key part here to push the hospital to be more modernized. Be more electronic orientated. I think that’s the most benefit that it brings ” (IT Manager). As a result, “ when we were going to negotiate or bid for contracts with the PCT (Primary Care Trust responsible for purchasing services from Beta, now being restructured), we had more accurate figures on which to base our bid. This very much helps the business function of the Trust” (Senior Manager)."
Our adapted ‘sociotechnical changing’ framework has three main dimensions: the constantly evolving nature of the contexts: i.e. environment, organization, perceptions, and consequences of the implementation of EHR systems; the performative nature of evaluating the implementation and adoption of EHR to explore how it ‘worked out’ and was ‘made to work’; and finally exploring and narrating the implementation of EHRs ‘in the making’, beyond the potentially misleading dichotomy of success or failure. Such an approach enabled us to learn how the EHR was formed, translated and reproduced in various entities at Beta [ 59 ] and the different meanings it embodied for various stakeholders, at different times and locations [ 60 ]. Our study revealed the usefulness of this approach to shed light on empirical aspects of the implementation and adoption and to plan for improving the process.
The decisions to procure EHR, the selection of the specific software (RiO), the process of implementation and the attempts to make RiO work at Beta, all proceeded in a rapidly changing NHS environment. NPfIT was dismantled in 2011. If Beta had decided not to join NPfIT, the organization may have lost the opportunity of being an ‘early adopter’ of NHS-centrally procured EHR systems. At the time of making the decision to proceed in 2008, the financial and non-fiscal incentives to be early adopter, and Beta’s desire to seize the opportunity of integrated EHR to get closer to the Foundation c status to help the organization survive, were constantly and quickly changing over time. For stakeholders at Beta, EHR embodied certain interests of (e.g. senior managers, doctors, IT staff, managers, etc.) that was linked to systems of politics and power relations [ 48 ], which shaped perceptions and actions, as opposed to a discrete and contextual resource deployed in planned processes of change [ 42 , 61 ]. In hindsight, irrespective of shortcomings of the implementation and some negative experiences by the users, Beta’s decision was a right choice for the organization and the quality of care for its patients.
Our performative and social construction view helped explore the implementation of RiO in the making and portray how users from various disciplines shifted their perceptions and attitudes towards the EHR system in use, and became generally positive to make it work for their organization. In this way, change is rarely a fast or direct movement from ‘the old’ to ‘the new’, rather the new is born within the old and co-exists with it, and the old and the older still remain sedimented within the most new [ 35 , 54 ]. In addition to capturing what people said they did versus what they were doing, we managed to reconcile between the state of being (e.g. being a nurse, or doctor, or computer, etc.) and practice of doing (e.g. order entry, putting notes, etc.) [ 35 ]. Our longitudinal evaluation allowed us to understand the implementation process through engaging with actors who experienced changing in their daily interaction with EHR, and who also were being changed by it.
Initially, the users expressed mixed feelings about RiO and perceived it as being somewhat inadequate. They complained that it lacked some key clinical functionality versus loads of useless functions on RiO, and the significant cultural and work environmental changes that EHR brought to mental health settings. For instance, in line with the literature, some clinical users were concerned about adverse effects of EHR on healthcare practitioner-patient relationship [ 14 , 15 , 62 , 63 ]. However, a lot of users’ initial anxiety, negative attitudes, and stress were replaced with hope and satisfaction. This partly happened as a result of attempts to make RiO work and appropriate preparation to adopt EHR, which led to experiencing and recognizing some early benefits. Modest early benefits led users change their behaviour in substantial ways, many of whom, including doctors and nurses mentioned the greater degree to which they paid attention to creating more accurate and meaningful notes on RiO, because they were constantly seen and judged by their colleagues.
Our theoretical perspective helped ensure that we did not reduce the EHR to delivery, implementation and immediate use [ 53 ], but understand it as both cause and consequence of longer-term processes of changing, during which people and EHR came together to perform actions and tasks [ 56 ] as co-constitutive entities [ 43 , 55 ]. Such a social and cultural shift did not happen serendipitously and over a night. Rather, we noticed that the vision of change management [ 64 ], the leadership of the organization who made the decision to join the NPfIT despite the negative climate and the uncertain future of NPfIT, and constant support and help from senior management who invested in appropriate infrastructure, were the main reasons behind changing towards improvement, and reducing degrees of resistance to adopt EHR, and making RiO work at Beta.
The process of the implementation of RiO, as we understood it, involved multiple intricately woven moments of changing including inter alia combinations of the organizational, technical, social, professional and care, which was materialized as it was performed by various stakeholders with different sets of attitudes and perceptions, at different times and locations, across our context of investigation: Beta. This led us to learn insights that could be obtained from approaches that sought to ‘tell the whole story’ not just the ending [ 30 , 65 ]. Our ‘sociotechnical changing’ framework enabled us to manifest changing by capturing stakeholders’ perceptions of EHR as instances of both projection (what is possibly becoming new) and remembrance (what is old and difficult to give up) [ 42 ]. For us, studying implementation and adoption of EHR was inevitably and eternally a process or performance, suspended between what was and what might one day be. The EHR thus comes into being as and when it is performed (not when software is delivered and installed) even to the extent that it ‘vanishes when it is no longer performed’ [ 66 ]. In addition, we observed redistribution of professional responsibility and degrees of job change as users attempted to inscribe their interests into EHR [ 50 ]. Initially, there were complaints about extra burden of administrative job. Some users, senior doctors in particular, for whom administrative job was conducted by junior doctors or nurses traditionally, were reluctant to put notes on RiO. Nurses, in contrast, were generally more compliant as they projected EHR as a chance to take more control over their work (remembrance).
Further, by exploring EHR system (RiO in this case) in-the-making, we focused on real concerns of policy makers and managers – the causal texture within which the implementation happened. Our findings brought to the fore the intricate set of interlocking changes in practice that EHR implies, a more formative view than the image of discrete change, and a detailed stock of knowledge that informed key stakeholders at the time that it was, we believe, most needed. For illustrative purposes, we refer to feedback, recognizing that our ‘sociotechnical changing’ approach resulted in useful outputs that informed strategies and brought improvements to the implementation of RiO at Beta:
" ‘…excellent stuff that truly gave us insights we as a deployment team had not perhaps fully thought about or understood… I think a second phase review of perceptions of the system after it has settled down would be extremely beneficial, warts and all, and help with the formation of our future strategies and approach.’ (Senior manager)."
Although not optimal, people worked hard around issues to make the system more compatible with their organizational needs. As a result, they harvested some modest benefits for both patients and the organization [ 67 ] and valued the system [ 37 ] eventually. From our perspective, non- or partial adoption but also rejection, mis-use, non-use, resistance to EHR and workarounds, all are not simply negative effects, pathologies or signs of failure, but are alternative enactments upon technology, which may pave the trajectory of organizational learning towards future smoother implementation process [ 42 , 68 , 69 ]. In this way, as an intertwined product of technology; work practices; and people who make them work, EHR is made to actively produce a fit system to the needs of organization [ 43 ].
All in all, the ‘sociotechnical changing’ perspective helped us move away from static before and after implementation ‘impacts’ or notions of discrete change. Instead, we focused on nominalism (rather than essentialism), crossing of temporalities (rather than before-after dualisms) and practice (rather than strategic or functional) orientation [ 35 ].
Our findings are in contrast to the claims that EHR may lead to impersonal and inaccurate clinical notes in mental health settings [ 11 ]. Given a great desire of mental health patients to receive a copy of their summary notes (78% of patients reported that it was helpful to receive the letter, and 83% reported that they would like to continue receiving them) [ 70 ], EHR may lead to enhanced patient satisfaction by producing more accurate notes. The evaluation of the implementation of EHR in an NHS community mental health setting showed similar results: high degree of users’ satisfaction and some tangible benefits to clinical staff [ 71 ].
Lessons for implementers
On the basis of challenges encountered during the implementation of EHRs, and early benefits realized at Beta, we consider some policy implications below that may help facilitate the improved implementation of EHR systems into mental health settings.
First, stakeholders need to be identified prior to planning to procure and implement EHR software, and their computer literacy and ability to access the technology needs to be adjusted accordingly [ 72 ]. Engaging with healthcare professionals from early stages of planning and as EHR partners is pivotal to maximize efficacy and improve patient care.
Second, although overlooked by the NPfIT, it is important to understand whether both mental health service providers and users would like to have EHR systems–and for what purposes–before embarking on the large-scale implementation of EHR systems.
Third, EHR needs to be seen as a sociotechnical entity by stakeholders, thereby ensuring a user-centred design of EHR [ 73 , 74 ]. It is important to address concern of users who may present less interest and enthusiasm about EHR.
Fourth, contextualization and taking heterogeneity across mental health settings is crucial to implement EHR initiatives. This might also help identify areas in need of additional support when implementing EHR software.
Fifth, given a huge cultural shift that EHR brings [ 75 ] to heavily text-based notes in mental health, healthcare practitioners must be educated and protected with regards to transparency and observing confidentiality of patient notes.
Last but not least, the safety of EHR systems needs to be ensured prior to and during the implementation [ 76 ], and their efficacy requires to be evaluated using robust, independent, and forethought evaluation programs that employ reflexive and multidisciplinary research team [ 30 ].
Strengths and limitations of this work
Our findings need to be interpreted with caution. We evaluated one ‘early adopter’ mental hospital in England, during a relatively short period of EHR implementation and in the beginning of a long journey to full integration. We did not intend to evaluate the software specifications per se . Rather, we attempted to understand what was ‘going on’ in terms of the implementation and adoption of EHR in the studied settings, namely the process of implementation not outcomes. The in-depth case study approach [ 28 , 30 ] was helpful to ensure an understanding of the contextual aspects of the implementation, however generalizable lessons can only be drawn with great caution. In addition, our adapted ‘sociotechnical changing’ perspective may have narrowed down our focus on the micro level, thus hindering the bigger picture to be portrayed. Nonetheless, we managed to collect data from various stakeholders from outside Beta, and compared our analytic themes with other case studies in our evaluation. This may have expanded our understanding of the phenomenon. We acknowledge that many perceptions and attitudes described here may be altered in times to come, as there is a natural learning and adoption curve in any organizational change initiative. Finally, we did not study patients in our evaluation. Other studies on the impact of EHR use on the quality of the patient-psychiatrist relationship found no change in satisfaction scores among adult psychiatric patients for whom EHR was used during outpatient encounters instead of paper charting [ 7 ].
Nevertheless, despite the above limitations, little has previously been published on EHRs in mental health settings, let alone in the context of national implementation endeavors. This paper may shed light on some practical dimensions of EHR implementation and things to consider when planning implementing integrated EHR systems in mental health settings. As such, we hope that will help in the many future implementation and adoption of EHR systems in mental health settings that are now underway or are planned in countries with similar healthcare systems, and possibly beyond.
There is now a strong policy drive to implement EHRs in mental health settings. Despite substantial initial challenges, the English mental health hospital reported on in this paper achieved some early-perceived benefits from implementing the EHR system. These benefits related to improved legibility and accessibility of patient records, and transparency of care processes. Because of the nature of mental health and the specific conditions of its patients, some of them have difficulty in describing their problems and occasionally their medical history appropriately, shared electronic records proved to be potentially useful for their safety. As mental health settings face greater challenges for providing a quality service at an acceptable cost, wise implementation of suitable EHR applications may boost the chances for the success.
a A collection of national applications, services and directories that support the NHS and the exchange of information across national and local NHS systems. The project began in 2003, and now every NHS organization can access the Spine services ( http://www.connectingforhealth.nhs.uk/systemsandservices/spine ).
b Care programme approach: Anyone experiencing mental health problems is entitled to an assessment of their needs with a mental healthcare professional, and to have a care plan that is regularly reviewed by that professional (NHS Choice 2012).
c Greater autonomy and freedoms for NHS hospitals within a national framework of standards [DH 2005].
Jones IR, Ahmed N, Catty J, McLaren S, Rose D, Wykes T, Burns T: Illness careers and continuity of care in mental health services: a qualitative study of service users and carers. Soc Sci Med. 2009, 69: 632-639. 10.1016/j.socscimed.2009.06.015.
Article PubMed Google Scholar
Burns T, Catty J, White S, Clement S, Ellis G, Jones IR, Lissouba P, McLaren S, Rose D, Wykes T: Continuity of care in mental health: understanding and measuring a complex phenomenon. Psychol Med. 2009, 39: 313-323. 10.1017/S0033291708003747.
Article CAS PubMed Google Scholar
Tsai J, Bond G: A comparison of electronic records to paper records in mental health centers. Int J Qual Health Care. 2008, 20 (2): 136-143.
Hillestad R, Bigelow J, Bower A, Girosi F, Meili R, Scoville R, Taylor R: Can electronic medical record systems transform health care? Potential health benefits, savings, and costs. Health Aff (Millwood). 2005, 24: 1103-1117. 10.1377/hlthaff.24.5.1103.
Article Google Scholar
Wang SJ, Middleton B, Prosser LA, Bardon CG, Spurr CD, Carchidi PJ, Kittler AF, Goldszer RC, Fairchild DG, Sussman AJ, Kuperman GJ, Bates DW: A cost-benefit analysis of electronic medical records in primary care. Am J Med. 2003, 114: 397-403. 10.1016/S0002-9343(03)00057-3.
Department of Health: Health Informatics Review Report. 2008, Stationery Office, London
Google Scholar
Stewart RF, Kroth PJ, Schuyler M, Bailey R: Do electronic health records affect the patient-psychiatrist relationship? A before & after study of psychiatric outpatients. BMC Psychiatry. 2010, 8: 10-13.
Ennis L, Rose D, Callard F, Denis M, Wykes T: Rapid progress or lengthy process? electronic personal health records in mental health. BMC Psychiatry. 2011, 26: 11-117.
Callard F, Wykes T: Mental health and perceptions of biomarker research - possible effects on participation. J Ment Health. 2008, 17: 1-7. 10.1080/09638230801931944.
Essex B, Doig R, Renshaw J: Pilot study of records of shared care for people with mental illnesses. BMJ. 1990, 300: 1442-1446. 10.1136/bmj.300.6737.1442.
Article CAS PubMed PubMed Central Google Scholar
Plovnick RM: The progression of electronic health records and implications for psychiatry. Am J Psychiatry. 2010, 167: 498-500. 10.1176/appi.ajp.2009.09101473.
Weitzman ER, Kaci L, Mandl KD: Acceptability of a personally controlled health record in a community-based setting: implications for policy and design. J Med Internet Res. 2009, 11: e14-10.2196/jmir.1187.
Article PubMed PubMed Central Google Scholar
Rothstein MA: The Hippocratic bargain and health information technology. J Law Med Ethics. 2010, 38 (1): 7-13. 10.1111/j.1748-720X.2010.00460.x.
Koide D, Asonuma M, Naito K, Igawa S, Shimizu S: Evaluation of electronic health records from viewpoint of patients. Stud Health Technol Inform. 2006, 122: 304-308.
PubMed Google Scholar
Garrison GM, Bernard ME, Rasmussen NH: 21st-century health care: the effect of computer use by physicians on patient satisfaction at a family medicine clinic. Fam Med. 2002, 34 (5): 362-368.
Hamann J, Leucht S, Kissling W: Shared decision making in psychiatry. Acta Psychiatr Scand. 2003, 107: 403-409. 10.1034/j.1600-0447.2003.00130.x.
Department of Health: ‘Delivering the NHS Plan: next steps on investment, next steps on reform’. 2002, Department of Health, London
Cabinet Office: ‘Major Projects Authority Programme Assessment Review of the National Programme for IT’. 2011, http://www.cabinetoffice.gov.uk/sites/default/files/resources/mpa-review-nhs-it.pdf . (Last accessed 21.04.2012)
Brennan S: The biggest computer programme in the world ever! How's it going?. J Inf Tech. 2007, 22: 201-211.
Currie WL, Guah MW: Conflicting institutional logics: a national programme for it in the organizational field of healthcare. J Inf Tech. 2007, 22 (3): 235-247. 10.1057/palgrave.jit.2000102.
Cresswell K, Ali M, Avery A, Barber N, Cornford T, Crowe S, Fernando B, Jacklin A, Jani Y, Klecun E, Lichtner V, Marsden K, Morrison Z, Paton J, Petrakaki D, Prescott R, Quinn C, Robertson A, Takian A, Voutsina K, Waring J, Sheikh A: The long and winding road…an independent evaluation of the implementation and adoption of the national health service care records service (NHS CRS) in secondary care in England. 2011, Available from: http://www.haps.bham.ac.uk/publichealth/cfhep/005.shtml (Last accessed: 20/04/2012)
Brennan S: THE NHS IT PROJECT the biggest computer programme in the world.ever!. 2005, Radcliffe
National Audit Office: The National Programme for IT in the NHS: Progress since 2006. 2008, Available at: http://www.nao.org.uk/publications/0708/the_national_programme_for_it.aspx (Last accessed 25/05/2012)
Sheikh A, Cornford T, Barber N, Avery A, Takian A, Lichtner V, Petrakaki D, Crowe S, Marsden K, Robertson A, Morrison Z, Klecun E, Prescott R, Quinn C, Jani Y, Ficociello M, Voutsina K, Paton J, Fernando B, Jacklin A, Cresswell K: Implementation and adoption of nationwide electronic health records in secondary care in England: final qualitative results from prospective national evaluation in "early adopter" hospitals. BMJ. 2011, 343: d6054-10.1136/bmj.d6054.
Robertson A, Cresswell K, Takian A, Petrakaki D, Crowe S, Cornford T, Barber N, Avery A, Fernando B, Jacklin A, Prescott R, Klecun E, Paton J, Lichtner V, Quinn C, Ali M, Morrison Z, Jani Y, Waring J, Marsden K, Sheikh A: Implementation and adoption of nationwide electronic health records in secondary care in England: qualitative analysis of interim results from a prospective national evaluation. BMJ. 2010, 341: c4564-10.1136/bmj.c4564.
Murray SA, Sheikh A: Serial interviews for patients with progressive disease. Lancet. 2006, 368: 901-902. 10.1016/S0140-6736(06)69350-1.
Cornford T, Doukidis GI, Forster D: Experience with a structure, process and outcome framework for evaluating an information system. Omega, Int J Manage Sci. 1994, 22: 491-504. 10.1016/0305-0483(94)90030-2.
Crowe S, Cresswell K, Robertson A, Huby G, Avery A, Sheikh A: The case study approach. BMC Med Res Methodol. 2011, 27: 11-100.
Patton MQ: Qualitative research & evaluation methods. 2002, SAGE Publications, London
Takian A, Petrakaki D, Cornford T, Sheikh A, Barber N: Building a house on shifting sand: Methodological considerations when evaluating the implementation and adoption of national electronic health record systems. BMC Health Serv Res. 2012, 12: 105-10.1186/1472-6963-12-105.
Stake RE: The art of case study research. 1995, SAGE Publications, London
Yin R: Case study research, design and methods. 2009, SAGE Publications, London
Mays N, Pope C: Quality in qualitative health research: qualitative research in health care. 1999, BMJ Publication Group, London
Corbin J, Strauss A: Strategies for qualitative data analysis: basics of qualitative research. techniques and procedures for developing grounded theory. 2008, SAGE Publications, CA
Book Google Scholar
Petrakaki D, Cornford T, Klecun E: Sociotechnical changing in healthcare. Stud Health Technol Inform. 2010, 157: 25-30.
Butson R: Sociotechnical approach – STS. 2008, Available from: https://russell.wiki.otago.ac.nz/SocioTechnical_Approach_-_STS (last accessed: 21/05/2012)
Lorenzi NM, Riley RT: Managing change: an overview. JAMIA. 2000, 7: 116-124.
CAS PubMed PubMed Central Google Scholar
Ash JS, Gorman PN, Lavelle M, Stavri PZ, Lyman J, Fournier L, et al: Perceptions of physician order entry: results of a cross-site qualitative study. Methods Inf Med. 2003, 42 (4): 313-323.
CAS PubMed Google Scholar
Kling R, Scacchi W: The web of computing: computing technologies as social organization. Advances in Computers. Edited by: Yovits MC. 1982, Academic, New York
Berg M: The search for synergy: interrelating medical work and patient care information systems. Methods Inf Med. 2003, 42 (4): 337-344.
Massaro TA: Introducing physician order entry at a major academic medical center: I Impact on organizational culture and behavior. Acad Med. 1993, 68 (1): 20-25. 10.1097/00001888-199301000-00003.
Orlikowski WJ: Using technology and constituting structures: a practice lens for studying technology in organizations. Organization Sci. 2000, 11 (4): 404-428. 10.1287/orsc.11.4.404.14600.
Aarts J, Doorewaard H, Berg M: Understanding implementation: the case of a computerized physician order entry system in a large Dutch university medical center. JAMIA. 2004, 11 (3): 207-216.
PubMed PubMed Central Google Scholar
Berg M, Aarts J, Van Der Lei J: ICT in health care: sociotechnical approaches. Methods Inf Med. 2003, 42 (4): 297-301.
Clegg SR, et al: Learning/Becoming/Organizing. Organization. 2005, 12 (2): 147-167. 10.1177/1350508405051186.
Bijker WE: Of bicycles, bakelites and bulbs: Toward a theory of sociotechnical change. 1995, MIT Press, Cambridge, MA
Callen JL, Braithwaite J, Westbrook J: Contextual implementation model: a framework for assisting clinical information system implementations. JAMIA. 2008, 15 (2): 255-262.
Klecun E, Cornford T: A critical approach to evaluation. Eur J Inf Sys. 2005, 14: 229-243. 10.1057/palgrave.ejis.3000540.
Jones MR: Computers can land people on Mars, why can't they get them to work in a hospital? Implementation of an Electronic Patient Record System in a UK Hospital. Methods Inf Med. 2003, 42 (4): 410-415.
Cho S, Mathiassen L, Nilsson A: Contextual dynamics during health information systems implementation: an event-based actor-network approach. Eur J Inf Sys. 2008, 17: 614-630. 10.1057/ejis.2008.49.
Ash JS, Gorman PN, Lavelle M, Payne TH, Massaro TA, Frantz GL, et al: A cross-site qualitative study of physician order entry. JAMIA. 2003, 10 (2): 188-200.
Davidson E, Chiasson M: Contextual influences on technology use mediation: a comparative analysis of electronic medical records systems. Eur J Inf Sys. 2005, 14: 6-18. 10.1057/palgrave.ejis.3000518.
Lin A, Cornford T: Sociotechnical perspectives on emergence phenomena. The new sociotech: Graffiti on the long wall. 2000, Springer, Godalming, 51-60.
Chapter Google Scholar
Latour B: Science in Action: How to Follow Scientists and Engineers Through Society. New edition. 1988, Harvard University Press, Cambridge, MA
Berg M: Patient care information systems and health care work: a sociotechnical approach. Int J Med Inf. 1999, 55 (2): 87-101. 10.1016/S1386-5056(99)00011-8.
Article CAS Google Scholar
Law J: Organizing modernity: Social order and social theory. 1993, WileyBlackwell
Knight S: The NHS information management and technology strategy from a mental health perspective. Adv Psychiatr Treat. 1995, 8: 223-229.
Thiru K, Hassey A, Sullivan F: Systematic review of scope and quality of electronic patient record data in primary care. BMJ. 2003, 326: 1070-10.1136/bmj.326.7398.1070.
Gherardi S: Practice? It’s a matter of taste?. Manag Learn. 2009, 40: 535-10.1177/1350507609340812.
Mol A, Law J: Regions, networks and fluids: Anaemia and social topology. Soc Stud Sci. 1994, 24 (4): 641-671. 10.1177/030631279402400402.
Orlikowski WJ: Improvising organizational transformation over time: A situated change perspective. Inf Sys Res. 1996, 7 (1): 63-92. 10.1287/isre.7.1.63.
Ridsdale L, Hudd S: Computers in the consultation: the patient's view. Br J Gen Pract. 1994, 44: 367-369.
Roy D: Recording health care and sharing the information - more bureaucracy or a welcome challenge to prevailing practice?. Psychiatr Bull. 2004, 28: 33-35. 10.1192/pb.28.2.33.
Takian A: Envisioning electronic health record systems as change management: The experience of an English hospital joining the National Programme for Information Technology. Stud Health Technol Inform. 2012, 180: 901-905.
Greenhalgh T, Stones R: Theorising big IT programmes in healthcare: Strong structuration theory meets actor network theory. Soc Sci Med. 2010, 70: 1285-1294. 10.1016/j.socscimed.2009.12.034.
Latour B: Reassembling the Social: An Introduction to Actor-Network-Theory. 2007, Oxford University Press, USA
Berg M: Implementing information systems in health care organizations: myth and challenges. Int J Med Inf. 2001, 64: 143-156. 10.1016/S1386-5056(01)00200-3.
Lucas HC: Why information systems fail. 1975, Columbia University Press, New York
Sauer C: Why information systems fail: a case study approach. 1993, Henley-on-Thames: Alfred Waller, Oxfordshire, UK
Nandhra HS, Murray GK, Hymas N, Hunt N: Medical records: doctors' and patients' experiences of copying letters to patients. Psychiatr Bull. 2004, 28: 40-42. 10.1192/pb.28.2.40.
Meredith J: Electronic patient record evaluation in community mental health. Inf Pri Care. 2009, 17: 209-213.
Borzekowski DLG, Leith J, Medoff DR, Potts W, Dixon LB, Balis T, Hackman AL, Himelhoch S: Use of the internet and other media for health information among clinic outpatients with serious mental illness. Psychiatr Serv. 2009, 60: 1265-1268. 10.1176/appi.ps.60.9.1265.
Trivedi P, Wykes T: From passive subjects to equal partners. Qualitative review of user involvement in research. Br J Psychiatry. 2002, 181: 468-472. 10.1192/bjp.181.6.468.
Rose D, Sweeney A, Leese M, Clement S, Jones IR, Burns T, Catty J, Wykes T: Developing a user-generated measure of continuity of care: brief report. Acta Psychiatr Scand. 2009, 119: 320-324. 10.1111/j.1600-0447.2008.01296.x.
Takian A, Cornford T: NHS information: Revolution or evolution?. Health Policy and Technology. 2012, 1: 193-198. 10.1016/j.hlpt.2012.10.005.
Sittig DF, Classen DC: Safe electronic health record use requires a comprehensive monitoring and evaluation framework. JAMA. 2010, 303: 450-451. 10.1001/jama.2010.61.
Pre-publication history
The pre-publication history for this paper can be accessed here: http://www.biomedcentral.com/1472-6963/12/484/prepub
Download references
Acknowledgements
We are very grateful to Beta hospital for making this work possible and to all individuals who kindly gave their time. We thank Dr. Kathrin Cresswell for her comments on earlier draft of this manuscript and our colleagues on the NHS CRS Evaluation Team. This work was supported by the NHS Connecting for Health Evaluation Programme led by Professor Richard Lilford.
This paper is independent research commissioned by the NHS Connecting for Health Evaluation Programme. The views expressed in this publication are those of the authors and not necessarily those of the NHS, the National Institute for Health Research or the Department of Health.
Author information
Authors and affiliations.
Division of Health Studies, School of Health Sciences & Social Care, Brunel University London, Uxbridge, UB8 3PH, UK
Amirhossein Takian
eHealth Research Group, Centre for Population Health Sciences, The University of Edinburgh, Edinburgh, EH8 9DX, UK
Aziz Sheikh
Department of Practice and Policy, UCL School of Pharmacy, London, WC1H 9JP, UK
Amirhossein Takian & Nicholas Barber
You can also search for this author in PubMed Google Scholar
Corresponding author
Correspondence to Amirhossein Takian .
Additional information
Competing interests.
The authors declare that they have no competing interests.
Authors’ contributions
AT undertook data collection and analysis and drafted the first version of the manuscript with AS and NB, who all extensively contributed to several revisions and intellectual development of the article. All authors read and approved the final manuscript.
Electronic supplementary material
Additional file 1: interview topic guide: npfit & external stakeholders.(doc 30 kb), additional file 2: interview topic guide: healthcare professionals and managers.(doc 34 kb), additional file 3: interview topic guide: implementation teams.(doc 34 kb), rights and permissions.
This article is published under license to BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License ( http://creativecommons.org/licenses/by/2.0 ), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Reprints and permissions
About this article
Cite this article.
Takian, A., Sheikh, A. & Barber, N. We are bitter, but we are better off: case study of the implementation of an electronic health record system into a mental health hospital in England. BMC Health Serv Res 12 , 484 (2012). https://doi.org/10.1186/1472-6963-12-484
Download citation
Received : 29 May 2012
Accepted : 28 December 2012
Published : 31 December 2012
DOI : https://doi.org/10.1186/1472-6963-12-484
Share this article
Anyone you share the following link with will be able to read this content:
Sorry, a shareable link is not currently available for this article.
Provided by the Springer Nature SharedIt content-sharing initiative
- Electronic health records (EHR)
- Mental health
- ‘Sociotechnical changing’
- Implementation
BMC Health Services Research
ISSN: 1472-6963
- General enquiries: [email protected]
Log in using your username and password
- Search More Search for this keyword Advanced search
- Latest content
- BMJ Journals
You are here
- Volume 11, Issue 1
- Understanding challenges of using routinely collected health data to address clinical care gaps: a case study in Alberta, Canada
- Article Text
- Article info
- Citation Tools
- Rapid Responses
- Article metrics
- http://orcid.org/0000-0002-8800-6081 Taylor McGuckin 1 ,
- Katelynn Crick 1 ,
- Tyler W Myroniuk 2 ,
- Brock Setchell 1 ,
- Roseanne O Yeung 1 , 3 ,
- Denise Campbell-Scherer 1 , 4
- 1 Faculty of Medicine & Dentistry - Lifelong Learning & Physician Learning Program , University of Alberta , Edmonton , Alberta , Canada
- 2 Public Health , University of Missouri , Columbia , Missouri , USA
- 3 Division of Endocrinology & Metabolism, Faculty of Medicine and Dentistry , University of Alberta , Edmonton , AB , Canada
- 4 Department of Family Medicine, Faculty of Medicine and Dentistry , University of Alberta , Edmonton , AB , Canada
- Correspondence to Dr Denise Campbell-Scherer; denise.campbell-scherer{at}ualberta.ca
High-quality data are fundamental to healthcare research, future applications of artificial intelligence and advancing healthcare delivery and outcomes through a learning health system. Although routinely collected administrative health and electronic medical record data are rich sources of information, they have significant limitations. Through four example projects from the Physician Learning Program in Edmonton, Alberta, Canada, we illustrate barriers to using routinely collected health data to conduct research and engage in clinical quality improvement. These include challenges with data availability for variables of clinical interest, data completeness within a clinical visit, missing and duplicate visits, and variability of data capture systems. We make four recommendations that highlight the need for increased clinical engagement to improve the collection and coding of routinely collected data. Advancing the quality and usability of health systems data will support the continuous quality improvement needed to achieve the quintuple aim.
- quality improvement
- quality improvement methodologies
- data accuracy
- health services research
- healthcare quality improvement
This is an open access article distributed in accordance with the Creative Commons Attribution Non Commercial (CC BY-NC 4.0) license, which permits others to distribute, remix, adapt, build upon this work non-commercially, and license their derivative works on different terms, provided the original work is properly cited, appropriate credit is given, any changes made indicated, and the use is non-commercial. See: http://creativecommons.org/licenses/by-nc/4.0/ .
https://doi.org/10.1136/bmjoq-2021-001491
Statistics from Altmetric.com
Request permissions.
If you wish to reuse any or all of this article please use the link below which will take you to the Copyright Clearance Center’s RightsLink service. You will be able to get a quick price and instant permission to reuse the content in many different ways.
Introduction
A learning health system is foundational to achieving the quintuple aim of advancing patient care, population health, equity, cost-effectiveness, healthcare worker experience, and, ultimately, future goals such as precision health. 1–3 To be able to rapidly answer important clinical questions, the structure of, and data capture in, electronic medical records and health administrative databases needs to be improved. Alberta, Canada is a globally recognised jurisdiction for its health data infrastructure and capture. However, health service researchers have identified important limitations to its use. 4–8 Reasons for these limitations include the historic use of different health information systems across Alberta’s regions, 9 and the creation of administrative health databases for non-clinical functions such as payment. 10
The Physician Learning Program (PLP) 11 is a provincial programme that works to understand gaps in clinical practice, create clinically actionable information and cocreate sustainable solutions with physicians, allied health teams, patients and community, and health system partners to advance practice. Here, we share four examples of PLP projects on a range of rare to common medical conditions that highlight some of the current challenges of using routinely collected health data to inform real-world clinical problems and support quality improvement. These four projects demonstrate areas where we encountered limitations in data capture, which if rectified, would provide needed information to help advance care of Albertans. We offer guidance in improving routinely collected health data that is broadly relevant to health systems by addressing issues of data completeness, availability, missingness and duplication, and variability in capture. Improvements in these areas are necessary to increase the usability of data for healthcare, health services research, and, eventually, future applications of artificial intelligence and precision health.
The primary objective of this work was to capture, categorise, and label overarching and recurring problematic data patterns in electronic health records and administrative databases observed through work conducted at PLP. The four projects presented were conducted to understand gaps in clinical care and develop baseline data for quality improvement initiatives. Each project is described in table 1 , with notes on data sources in table 2 . For each project, a series of questions were co-created with clinicians to provide information of importance for clinical quality improvement. We identified whether secondary data from electronic medical records and administrative databases was available or whether primary data collection was necessary. Routinely collected health data from electronic medical records and other administrative databases was feasible and extracted for three projects: (1) Adult Diabetes; (2) Paediatric Diabetic Ketoacidosis, a serious complication of diabetes and (3) Adrenal Insufficiency, a rare, life-threatening hormonal disorder. For the Beta-Lactam Allergy and Surgical Prophylaxis project, the required clinical information was not routinely collected into an administrative database. Thus, primary data collection was required and included manually extracting information from paper charts.
- View inline
Description of the Physician Learning Program projects including purpose, representative questions, whether a challenge was encountered, and databases used
Descriptions of the data sources used to complete the projects
Figure 1 represents the iterative process used to identify, collect, clean and synthesise routinely collected health information needed for clinical quality improvement. Detailed methods and results of the four projects will be published elsewhere. The data collected and analysed for this paper is not the quantitative data of the four projects, but our observations while conducting them. Briefly, for the projects that used routinely collected health data, we formulated a data query to find and pull the raw data needed to answer each project question. A trained analyst employed by Alberta Health Services extracted the data. Once extracted, the raw data were cleaned and analysed using standard statistical software (Oracle SQL Developer, Python V.3.4, SAS V.9.4 and RStudio V.1.2.5033). The clinicians working on the project reviewed the results to assess the validity and completeness of the data in comparison with their knowledge of clinical workflow and processes. The results were compiled into various formats, including presentations, reports, infographics, and clinical tools, and then disseminated to relevant stakeholder groups. Their purpose ultimately is to inform clinical quality improvement and co-creation of interventions to address clinical gaps in care.
- Download figure
- Open in new tab
- Download powerpoint
The Physician Learning Program’s non-linear process of quality improvement using routinely collected health data. The key elements are: (1) cocreating clinical questions and identifying whether secondary data are available or if primary data collection is necessary; (2) gathering data from databases or completing primary data collection; (3) deep cleaning of the data; (4) conducting analyses and further data cleaning; and (5) effectively communicating findings that serve as the basis for quality improvement.
Systematic approach used to capture and categorise main challenges and identify root causes
Over a 2-year period, recurring difficulties arose when obtaining and analysing the administrative data needed to answer clinical questions for the four projects. We undertook a systematic approach to identify and capture whenever problems arose and then categorise them into main challenges . This systematic approach included: (1) capturing whenever a data problem occurred in a project; (2) discussing the problem within our interdisciplinary team of researchers and clinical experts; (3) discussing recurring issues and patterns through team meetings and with key informant discussions; and (4) synthesising them into main categories that spanned projects, healthcare settings, and health conditions. We identified and verified the root cause whenever possible by: (1) talking to clinical, administrative, and analytical staff within Alberta Health Services and Alberta Health (two regulatory government bodies that oversee the delivery of healthcare within the province of Alberta); (2) reading publicly available database documentation 12–14 and (3) talking to front-line healthcare staff with deep knowledge of the healthcare setting and clinical systems. Our systematic approach is summarised in box 1 .
Methods used to identify, collect and analyse the raw data (ie, problems arising in using administrative data to answer the clinical questions)
Methods to identify the raw data.
Observe when there was a problem while conducting each of the steps in figure 1 .
Verify if there was a challenge by checking against known published problems and discussing with data analysts and clinicians to see if it matches reality.
Methods to collect the raw data
Formally document the problem encountered and how it was verified .
Methods used to analyse the raw data
Discuss the problems from each project and collate and summarise them into overarching themes (main challenges).
Patient and public involvement
At the PLP, we have the mission to create “actionable clinical information and engage with physicians, teams and partners to cocreate sustainable solutions to advance practice.” 11 Inherent in this process is the involvement of broader networks outside of the project team including community physicians, physician networks, policy-makers, patients, researchers, and other healthcare professionals. Involvement of stakeholders starts at project conception with physicians and clinical teams cocreating project ideas with the PLP based on health system gaps. Engagement continues through to the dissemination of project outcomes where we integrate with networks to engage in knowledge translation activities, codesign sustainable solutions, and implement them with health system partners.
Through our systematic approach of capturing and categorising recurring problems, as outlined in detail above, we identified four broad challenges of using routinely collected health data to address real-world clinical questions. We present them here framed in four example projects. These four challenges and example project questions are summarised in table 3 .
Data challenges encountered while answering clinical questions
Description of challenges
Challenge 1: are the data field(s) needed to answer the clinical question available in administrative databases.
Not all information collected at a patient encounter has a corresponding data field in an administrative database; some information, although available, is not abstracted from the patient chart into a database. In the beta-lactam allergy and surgical prophylaxis project, 0 out of 3218 audited surgical cases contained allergy information in an available administrative database because there was no routinely populated data field for this information. However, for all cases, we found that allergies were recorded in paper charts. Importantly, inappropriate antibiotic prophylaxis, due to allergy status, is associated with a 50% increase odds in surgical site infections and increased costs to the system. 15 Assessing care using paper chart audits is sometimes justified but is not sustainable or scalable on a large basis because of its resource intensiveness. We are now working with health system delivery stakeholders to develop more sustainable solutions for this problem of antibiotic allergy and prophylaxis information not being electronically captured and available, specifically.
For the paediatric diabetic ketoacidosis project, only 28.6% of children’s admissions across Alberta contained data on medication, electrolyte, and fluid administration. Guideline concordance of care for this life-threatening condition cannot be assessed without this information. This information was only available for patients whose encounter was at a site that used Sunrise Clinical Manager, a specific Clinical Information System. Only five Alberta Hospitals and Health Centres, out of over 100 included in our project, used this system inhibiting the feasibility of assessing guideline concordant care across the whole system.
When assessing patient comorbidities in the adult diabetes project, we could not determine whether patients had a history of hyperosmolar hyperglycaemic state. Despite the International Classification of Diseases-9 (ICD-9) having a corresponding code for this condition, Alberta Health’s coding taxonomy, which is used to capture visit information to pay providers across the province, does not include all ICD-9 codes. 16 Thus, this comorbidity could not be assessed for any of the patients.
Challenge 2: if the data field needed to answer the clinical question is available, is the information complete and accurate?
The completeness of extracted data was problematic in two of our projects. When assessing lab results in the paediatric diabetic ketoacidosis project, we found that 46.6%, 94.5% and 12.6% of admissions at one of the children’s hospitals in the province had no results for blood pH, blood bicarbonate, and blood glucose, respectively. These laboratory results are central to guiding diabetes care and confirming a diagnosis of diabetic ketoacidosis. Through our root cause analysis, which included consulting with experts in the hospital laboratory, we uncovered that laboratory tests completed from capillary blood sources may not flow from bedside instruments to administrative databases; a historical legacy of funding restrictions when the system was developed. Additionally, we observed incomplete medication, fluid, and electrolyte administration data, which are all necessary for assessing quality of care in relation to established guidelines.
In the adult diabetes project, routinely collected health data were often missing for measures such as blood pressure, an important clinical assessment for predicting disease complications. In one clinic, 65.5% of visits did not have a blood pressure measurement recorded in a database. Through consultation, we determined that while front line staff are entering these measures into the electronic medical record, it does not flow into administrative databases.
Challenge 3: can the number of visits for a particular medical condition be accurately measured using administrative data?
We were unable to accurately estimate the number of outpatient visits for the treatment of adrenal insufficiency due to visits missing from the databases. Missing visits are a consequence of both imprecise codes used at the time of data submission (eg, visits coded as ‘follow-up’) and variation in data submission requirements in which not all visits are required to be submitted and thus captured. Variation in data submission requirements are a result of various payment structures (eg, alternative payments plans) across and within regions of the province. Thus, it is uncertain of how to compare regional data.
Furthermore, we encountered difficulty reconciling duplicate entries within and between databases housing different aspects of clinical visits. In this example, both Physician Claims and the National Ambulatory Care Reporting System (NACRS) database are used to capture outpatient visit data. They capture much of the same information but use different taxonomies to capture diagnostic information: one uses ICD-9 where the other uses ICD-10. Some visits are captured only in Physician Claims or NACRS, some in neither, and some in both. 12–14 17 There is no official reconciliation for visits captured in both. We found that at least 27% of adrenal insufficiency visits were likely duplicates. Of the 211 207 visits analysed, only 5.7% had a diagnostic code for adrenal insufficiency; clinical colleagues insisted this was implausibly low. This raised concerns that an indeterminate number of visits were missing from both databases, which may be due to visits for more than one medical condition not capturing all of the relevant diagnoses. In 78% of visits, there was only one code provided for the visit. Most codes used for the analysed visits were vague such as ‘general examination’ and ‘follow-up’, making it difficult to identify visits related to the treatment of adrenal insufficiency, which likely contributed to this discrepancy.
Challenge 4: can laboratory tests across the province be identified, harmonised, and analysed?
Three laboratory information systems are used across Alberta–a historical legacy of healthcare regionalisation. Laboratory codes are not harmonised across any of the laboratory databases within the province of Alberta. Each laboratory information system uses different laboratory codes, and thus, identifying and matching relevant codes across databases is not a trivial task. For example, haemoglobin A1c, a diabetes test, was found to be coded as HbA1c, ZHBA1C and HBA1X depending on where the lab test was completed. One major consequence was that 919 laboratory codes had to be reviewed to identify and harmonise the codes used in the paediatric diabetic ketoacidosis project. This was also problematic for the adult diabetes project ( online supplemental table 1 ).
Supplemental material
Strengths and limitations of the databases used.
Through completing these four projects, we identified both strengths of limitations of the administrative databases for informing clinical quality improvement projects. Strengths and limitations in relation to our example projects and questions specifically, and cautions for their use are summarised in table 4 . This is not a comprehensive overview of the strengths and limitations of these databases, but rather a summation of our experiences.
Strengths and limitations of the databases as elucidated by our example projects
Rapid access to clinically important information is crucial to building a powerful learning health system 3 in pursuit of the quintuple aim. Health data infrastructure that supports rapid access to clinically important information for evidence-informed care and clinical quality improvement is key to supporting practice reflection and innovations to meet patient needs. Our PLP projects illuminate four challenges of using routinely collected health data to achieve these aims. First, we found that not all information collected in a patient encounter has a corresponding data field in an administrative database; costly, time-consuming primary data collection is then needed to assess important clinical questions prohibiting the feasibility of continual monitoring. Second, when data fields are available, they may be absent or not uniformly populated. For instance, we observed this problem when clinical evaluations or readings from bedside instruments are used and the information does not flow to administrative databases. Third, establishing prevalence of medical conditions and number of visits was difficult due to missing records, complexity reconciling various databases that contain the same information, inconsistent diagnostic coding practices, and differing taxonomies used between databases. A key element of this challenge was that imprecise diagnostic codes, such as ‘follow-up’, did not permit clarity as to the topics addressed in the visit. The fourth challenge was the multiplicity of laboratory diagnostic codes used for the same test which made it difficult to develop data queries that capture all relevant tests.
The mission of the PLP is to create actionable clinical information and engage with physicians, teams, patients, and partners to cocreate sustainable solutions to advance practice. The creation of clinically actionable information from routinely available health data is hindered when there are substantial gaps in the information, as measuring improvement requires relevant baseline data and measurement over time to assess change. The strengths and limitations of administrative and electronic medical record health databases have been described extensively, for instance in the work of Burles et al , Clement et al and Edmondson and Reimer. 18–20 The inability to analyse data in real time is not a problem unique to the Canadian context, with challenges being documented in other jurisdictions including the USA. 21 The overarching issues relating to data capture, completeness, accuracy, and harmonisation, exist across healthcare systems and settings and challenges with data capture in clinical electronic medical records have been well documented. 22–27 Several of the databases outlined are available across Canada, including Discharge Abstract Database and NACRS, and thus these challenges are likely to exist across the country. Ongoing work is being conducted by the PLP and with relevant stakeholder groups to address the issues presented. We acknowledge the importance of collaborating with various stakeholders including data scientists, clinicians, and administrators to fully understand what the meaningful clinical data are and how to mobilise and act on them so that data-driven quality improvement is supported. Increased coordination and leveraging the opportunity of a new provincial acute care electronic medical record should continue to advance this work, particularly as efforts evolve across the care continuum.
Future directions
Advancing the quality of health systems data is crucial not only for current quality improvement projects, but also in realising the utility of precision health and artificial intelligence to advance healthcare in the future. 28–33 Health system data are necessary to meet the Federation of Medical Regulatory Authorities in Canada’s goal that all Canadian physicians participate in data-driven practice quality improvement. 33 The overarching purpose of these efforts is to support the development of a learning health system and to achieve improvements in the quintuple aim of improving population health, patients’ experience of care, equity, cost-effectiveness, and sustainability of healthcare workforce. 1–3 We strongly believe that the long-term benefits of improved data capture would significantly offset upfront investments. Importantly, supporting these efforts requires mobilising clinical information in a way that does not overwhelm the clinical workforce and contribute to physician burnout. 34
Addressing these four identified challenges is fundamental to creating a learning health system and to advancing healthcare delivery and health outcomes. We recommend the following:
To have more clinically important data available in readily extractable formats, we suggest expanding and harmonising mandatory data submission requirements with increased clinician engagement to ensure data that is captured is clinically meaningful.
To increase the quality and validity of the data available to assess patient care, we suggest the use of more specific codes and consistent taxonomies across the healthcare system to capture encounter diagnoses; standardisation of data entry processes with clear mechanisms of training and maintenance; and, ensuring the flow of clinically important information from bedside instruments, laboratory settings, and diagnostic imaging results to administrative databases in analysable formats.
To enhance efficiency and speed of data capture so that upgrading data quality, quantity, and structure is not at the cost of the clinical user, we suggest the incorporation of technologies like natural language processing, cross-platform interoperability, and application of human-centred design for workflow process improvement.
To promote real-time usability of data, we propose integrating technologies such as natural language processing and artificial intelligence to automate routinised functions to support appropriate real-time clinical decisions and reduce clinician burden.
Limitations
The challenges we identified in our routinely collected health data are specific to Alberta, Canada, however, they are commonly encountered in conducting QI and research work using administrative data and are generalisable internationally. 22–27 As information technology advances, integration into different health systems is variable leading to different local challenges in deriving solutions. We submit that the principles stated here may be of interest for consideration but additional factors will exist in different jurisdictions.
Through practical, real-world projects, we have identified four challenges in using administrative health and electronic medical record data to address clinical care gaps. Improving data infrastructure and quality will enable more nimble quality improvement efforts and real-world evidence studies. Improving this infrastructure, and the reliability and validity of data, is a necessary precondition for emergent technologies in precision health and artificial intelligence, and to developing a learning health system.

Ethics statements
Patient consent for publication.
Not applicable.
Ethics approval
This study was approved by Each project included the appropriate ethics approval from the Research Ethics Board-Health Panel at the University of Alberta, Edmonton, Alberta, Canada. The ethics approval numbers are as follows:Pediatric Diabetic Ketoacidosis-Pro00091652Diabetes Management-Pro00085385Beta-lactam Allergy and Surgical Prophylaxis-Pro00089593Adrenal Insufficienc-Pro00088478. Three of our projects included secondary retrospective analyses of routinely collected health data. The Physician Learning Programme only works with unidentified data. The third project was a paper chart audit and also was deidentified.
- Bodenheimer T ,
- R. Privitera M
- Friedman C ,
- Brown J , et al
- Hemmelgarn BR
- Simmonds K ,
- Usman HR , et al
- Guilcher SJT ,
- McKenzie N , et al
- Jolley RJ ,
- Jetté N , et al
- D'Souza AG , et al
- Donaldson C ,
- Sajobi T , et al
- ↵ Physician learning program , 2021 . Available: https://www.albertaplp.ca/ [Accessed 13 Jul 2021 ].
- Government of Alberta
- Alberta Health Services
- Canadian Institute of Health Information
- Blumenthal KG ,
- Li Y , et al
- Cunningham CT ,
- Topps D , et al
- Senior K , et al
- Clement FM ,
- Chin R , et al
- Edmondson ME ,
- Weiner MG ,
- Embi PJ , et al
- Madden JM ,
- Lakoma MD ,
- Rusinak D , et al
- Hickman T-TT , et al
- Malhotra S ,
- Barrón Y , et al
- Wang EC-H ,
- Cohoon TJ ,
- Bhavnani SP
- Setiyadi DBP ,
- Shaban-Nejad A ,
- Michalowksi M
- Maddox TM ,
- Rumsfeld JS ,
- Casalino LP ,
- Federation of Medicine Regulatory Authorities Canada
- Diabetes Canada clinical practice guidelines expert Committee
Supplementary materials
Supplementary data.
This web only file has been produced by the BMJ Publishing Group from an electronic file supplied by the author(s) and has not been edited for content.
- Data supplement 1
TM and KC are joint first authors.
Contributors TM, TWM ad KC conceived the project idea and drafted the manuscript. BS ensured data accuracy and contributed to the project methods. ROY and DC-S provided local clinical expertise. DC-S, TM, TWM, KC and ROY edited the manuscript.
Funding Supported by a financial contribution from the Government of Alberta via the Physician Learning Programme.
Disclaimer The views expressed herein do not necessarily represent the official policy of the Government of Alberta (no award/grant number).
Competing interests None declared.
Provenance and peer review Not commissioned; externally peer reviewed.
Supplemental material This content has been supplied by the author(s). It has not been vetted by BMJ Publishing Group Limited (BMJ) and may not have been peer-reviewed. Any opinions or recommendations discussed are solely those of the author(s) and are not endorsed by BMJ. BMJ disclaims all liability and responsibility arising from any reliance placed on the content. Where the content includes any translated material, BMJ does not warrant the accuracy and reliability of the translations (including but not limited to local regulations, clinical guidelines, terminology, drug names and drug dosages), and is not responsible for any error and/or omissions arising from translation and adaptation or otherwise.
Read the full text or download the PDF:
An official website of the United States government
The .gov means it’s official. Federal government websites often end in .gov or .mil. Before sharing sensitive information, make sure you’re on a federal government site.
The site is secure. The https:// ensures that you are connecting to the official website and that any information you provide is encrypted and transmitted securely.
- Publications
- Account settings
Preview improvements coming to the PMC website in October 2024. Learn More or Try it out now .
- Advanced Search
- Journal List
- J Med Internet Res
- v.22(10); 2020 Oct

Selecting Mobile Health Technologies for Electronic Health Record Integration: Case Study
1 Mobile App Gateway, Clinical & Translational Science Institute, Duke University, Durham, NC, United States
2 School of Nursing, Duke University, Durham, NC, United States
Marissa Stroo
Christopher fiander.
3 Duke Health Technology Solutions, Duke University Health System, Durham, NC, United States
Katlyn McMillan
Mobile health (mHealth) technologies, such as wearable devices and sensors that can be placed in the home, allow for the capture of physiologic, behavioral, and environmental data from patients between clinic visits. The inclusion of these data in the medical record may benefit patients and providers. Most health systems now have electronic health records (EHRs), and the ability to pull and send data to and from mobile devices via smartphones and other methods is increasing; however, many challenges exist in the evaluation and selection of devices to integrate to meet the needs of diverse patients with a range of clinical needs. We present a case report that describes a method that our health system uses, guided by a telehealth model to evaluate the selection of devices for EHR integration.
Introduction
Mobile health (mHealth) technologies, such as wearable devices and sensors that can be placed in the home, allow for the capture of physiologic, behavioral, and environmental data from patients between clinic visits. This patient-generated health data (PGHD) can help reveal underlying mechanisms of health by filling in gaps in the information, providing insights into the day-to-day activities of an individual, and allowing for better strategies to prevent and manage acute and chronic illnesses. Moreover, with the proliferation of smartphones rising to over 81% of the US population [ 1 ] and over 73% of households gaining in-home broadband internet [ 2 ], the ability to collect these data from diverse socioeconomic and geographic populations is growing. According to a 2018 survey conducted by Accenture, 75% of US consumers felt that technology was an important part of managing their health [ 3 ]. Rapid growth in the global digital health market, estimated to be over US $423 billion by 2024 [ 4 ], supports that sentiment. Because mHealth technologies tether to smartphones and Wi-Fi or have cellular-embedded chips, health data can be collected in near real time from patients in their daily environments.
In the United States, over 96% of all nonfederal acute care hospitals now possess a certified electronic health record (EHR) system [ 5 ], and over 9 in 10 office-based physician offices have adopted an EHR system [ 6 ]. As health care facilities move beyond EHR implementation, the integration of data from connected devices, including mHealth technologies, is gaining speed. Companies such as Apple Inc, for example, have enabled the ability for patients to aggregate their health records on an iPhone from multiple hospitals via authentication by health system patient portals, such as Epic’s MyChart [ 7 ]. It is also possible to integrate third-party data, such as patient-generated blood glucose levels, for example, into the EHR system via Apple HealthKit [ 8 ]. This capability is possible with many of the major EHR vendors, including Epic, Cerner, and Athena Health, among others. Furthermore, this capability is expanding to Android platforms as well with the use of Google Fit.
While these technologies afford much promise, many challenges exist for health systems and others in the selection of devices to integrate and recommend for equitable patient care. The Office of the National Coordinator for Health Information Technology published a white paper in 2018 highlighting some of the challenges of collecting and using PGHD [ 9 ]. These include the technical challenges related to accuracy of measurements, data provenance, and privacy and security concerns. They also explored the patients’ challenges and opportunities, which included the lack of internet or smartphone access as well as health and technology literacy deficits. A 2018 review by Reading and Merrill examined the needs of patients and providers around the use of PGHD in health care. Their review highlighted common needs for technology, including data quality, electronic integration, simple-to-understand actionable insights, and security.
The challenges and opportunities for PGHD are clear, but the path to moving forward remains undefined. One large obstacle is selecting the right devices from the ever-increasing number of consumer digital health devices on the market. Technology selection depends on the data of interest and the technology the patient, clinician, and health care system have ready access to and can use for clinical decision making or population health management. In this use case, we describe a method that Duke University Health System uses to evaluate and select devices for EHR integration.
Our team of researchers, clinicians, and informatics technology professionals met to identify factors involved with the selection of devices to integrate with the EHR system. These factors evolve based on feedback from stakeholders and the ever-growing digital health market. Key considerations included clinical validity of devices, patient satisfaction, and usability of both the connected device and the app interface associated with each smart, remote, monitoring device.
We use the Model for ASsessment of Telemedicine applications (MAST) [ 10 ] as a guide for device selection. This validated model is used by decision makers to aid in the choosing of the most appropriate telehealth technologies. We have modified the model to reflect variables needed in the selection of mHealth technologies for EHR integration. This includes, for example, details on US Food and Drug Administration (FDA) medical class and technological aspects, such as Bluetooth or Wi-Fi connection. The model includes three steps: Step 1: Preceding Considerations, Step 2: Multidisciplinary Assessment, and Step 3: Assessment of Transferability (see Figure 1 ) [ 10 ].

Process for evaluation of connected devices.
Step 1 involves determining the purpose of the connected device and relevant alternatives. The goal is to understand the primary outcomes and whether the device involves an upgraded or new technology. Next, several conditions are considered, including the following: legislation (ie, regulations, accreditations, and liability), reimbursement (ie, insurance vs hospital paid), maturity (ie, development time and resources needed over time to support the tool and how safe the tool is), and the number of patients involved to inform an economic analysis. Step 2 then involves a multidisciplinary assessment across eight domains. We added an eighth domain on technological aspects to reflect specific aspects of connected devices. The domains include the following: (1) health problem and description of the application, (2) safety, (3) clinical effectiveness, (4) patient perspectives, (5) economic aspects, (6) organizational aspects, (7) health equity [ 11 ], sociocultural, ethical, and legal aspects, and (8) technological aspects. Finally, in Step 3 an assessment is made as to the transferability of connected devices including interoperability (ie, Fast Healthcare Interoperability Resources) and the number of patients who will use the tool to determine costs per patient.
Figure 1 illustrates the three-step process for evaluating and selecting connected devices based upon a modified version of the MAST model. As we evaluate devices, our team maintains an internal working document of a table of devices. This document is refined and expanded as we make decisions and approach device integration. In order to create Figure 1 , we began by going through the process of selecting glucometers to recommend be integrated into our Epic-based EHR system (see Figure 2 ). This exercise allowed us to refine the process and add variables to Figure 1 . For example, because the evidence for many devices is limited we expanded to grey literature, including Consumer Reports and Amazon reviews, to gain perspective on patient usability and utility. Other examples include discovering the need to list technical requirements, such as Apple or Android capabilities, connection to Apple Health and Google Fit, and how data are collected and transmitted (ie, Bluetooth, Wi-Fi, and cellular). Of note, this case report focuses on evaluating device selection. Future work will evaluate clinical and institutional outcomes, as these tools are used as part of patient care delivery and research endeavors.

Example evaluation of noncontinuous glucometers: CONTOUR NEXT ONE.
We selected noncontinuous glucometers as our case study (see Figure 2 ) because of requests from clinician groups to retrieve glucometer data from patients and our experience integrating glucose data into our EHR system via Apple HealthKit [ 8 ]. As presented in Figure 2 , the exercise revealed that glucose is a data point of value for clinical care. Further, glucometers are considered FDA Class II medical devices and must demonstrate substantial equivalence to a predicate device. A review article by Klonoff et al investigated the accuracy of 18 marketed blood glucose monitors [ 12 ]. We searched the consumer-facing literature, such as Consumer Reports, to compare recommendations. The next steps in the evaluation process involved documenting the ability for each glucometer to be used on iOS and/or Android devices, integration with Apple HealthKit and Google Fit, costs, additional technical features, current integration with our EHR infrastructure and how data are retrieved, and if technical support is available. Results showed congruence across these measures and the CONTOUR NEXT ONE glucometer came out as the top contender.
Principal Findings
The proliferation of wireless and mobile technologies provides opportunities to connect information in real-world settings via wearable sensors and, when coupled with fixed sensors embedded in the environment, to produce continuous streams of data on an individual’s biology, psychology, behavior, and daily environment. These collected data have the potential to be analyzed and used in real time to prompt individuals to change behavior or their environmental exposures that can reduce health risks or to optimize health outcomes.
Selecting devices for integration requires many factors to be evaluated. These factors are technical, clinical, organizational, economic, and patient focused. Popular and currently well-known devices, such as the Fitbit and Apple Watch, are easier to identify due to their accessibility and widespread adoption. Evidence needed for activity trackers like these to be on the consumer market is also less stringent compared to evidence needed for a portable electrocardiogram or glucometer, which require FDA clearance. Devices that require FDA clearance provide an additional layer of evidence for safety and utility. This is in contrast to devices that do not require FDA clearance, such as sleep monitors.
Continued discussion with clinical and operational leaders suggests how broad the idea of technical support could be. Technical support can include such steps as configuring the device for the patient, providing support in person or remotely, and having staff available for ongoing troubleshooting. Other levels of technical support include patient support for managing the clinical data landing in the EHR system, with or without notification, along with addressing support related to the notifications specifically. Lastly, technical support should implicitly include the presentation of the data to providers so that they are actionable and accessible. Actionable and accessible data are essential for the provider or care manager to be able to intervene, while also not exacerbating provider burnout, which is more frequently reported since the large-scale implementation of EHRs. While these concepts are fundamental, they are also often frequent attributes referenced as potential barriers to inclusion by clinical and operational leaders.
A variety of devices should be selected for integration so that access to, and accessibility of, these tools is more equitable across patient populations. Patients have both iOS and Android devices and choosing one platform to focus on limits patient accessibility. Further, for devices that connect via a web portal or consume significant data through video, for example, patients may or may not have in-home broadband internet and are limited to internet access via their phone. This could be a limiting factor for patients based upon their geographic location or socioeconomic status. This is also important because the literature shows that devices are not always designed to be accurate across diverse populations. It was reported that the light sensor in some wearable devices was not usable on patients with darker skin tones due to the color of the optical sensor selected. While this has been addressed by many device manufacturers [ 13 ], it is a lesson in the importance of ensuring devices are usable across a variety of patient populations.
Future evaluation will also expand to include software platforms, such as those from Livongo, for example, which incorporate a variety of devices and provide personalized guidance to patients and clinicians with chronic illnesses. A third scenario is also necessary to consider regarding integration: there are applications and devices that offer their own portal for viewing data, but do not offer compatibility to iOS or Android, nor can these devices integrate into an aggregator inherently. Given this scenario, it becomes necessary to evaluate a device’s or platform’s capabilities through application programming interfaces (APIs) so that data can be aggregated productively and used in a clinical environment.
While mHealth technologies, specifically connected devices, hold promise to benefit patient care delivery and patient self-management, many challenges exist with their integration into health care. There is limited regulation, and rigorous scientific evaluation of many devices is lacking. There are many devices on the market, and every device must be tested for data quality, interoperability, and usefulness by patients and clinicians. Further, the rapid evolution of the connected device market requires frequent re-evaluation and system software updates. Finally, use of these tools in formal care delivery models is relatively new and, thus, understanding how to support patients and how to integrate and present the wealth of data from devices into actionable insights for clinical decision making continues to advance.
Conclusions
We present an example on how we recommend which mHealth devices should be integrated into a health system’s EHR system to collect PGHD. Many factors are involved, and it is important to conduct a thorough assessment to assess for clinical requirements, technical features, and patient-level factors such as usability and costs. Figures 1 and and2 2 can be used as templates for others to expand upon.
Acknowledgments
Support was provided, in part, by the Duke Clinical and Translational Science Institute, which is supported, in part, by a US National Institutes of Health Clinical and Translational Science Award (grant number UL1TR002553).
Abbreviations
| API | application programming interface |
| EHR | electronic health record |
| FDA | US Food and Drug Administration |
| MAST | Model for ASsessment of Telemedicine applications |
| mHealth | mobile health |
| PGHD | patient-generated health data |
Authors' Contributions: RS led the team and was responsible for all aspects of the project. MS, CF, and KM substantially contributed to the methods, data acquisition, results, and interpretation, and also participated in all aspects of writing the manuscript.
Conflicts of Interest: None declared.
This paper is in the following e-collection/theme issue:
Published on 15.7.2024 in Vol 10 (2024)
Development of Interoperable Computable Phenotype Algorithms for Adverse Events of Special Interest to Be Used for Biologics Safety Surveillance: Validation Study
Authors of this article:

Original Paper
- Ashley A Holdefer 1 , MS ;
- Jeno Pizarro 1 , BBA ;
- Patrick Saunders-Hastings 2 , PhD ;
- Jeffrey Beers 1 , MD ;
- Arianna Sang 1 , BA ;
- Aaron Zachary Hettinger 3, 4 , MS, MD ;
- Joseph Blumenthal 3 , BA ;
- Erik Martinez 1 , MBA ;
- Lance Daniel Jones 1 , MA ;
- Matthew Deady 1 , BA ;
- Hussein Ezzeldin 5 , PhD ;
- Steven A Anderson 5 , PhD
1 IBM Consulting, Bethesda, MD, United States
2 Accenture Inc, Ottawa, ON, Canada
3 Center for Biostatistics, Informatics and Data Science, MedStar Health Research Institute, Columbia, MD, United States
4 Department of Emergency Medicine, Georgetown University School of Medicine, Washington, DC, United States
5 Center for Biologics Evaluation and Research, United States Food and Drug Administration, Silver Spring, MD, United States
Corresponding Author:
Hussein Ezzeldin, PhD
Center for Biologics Evaluation and Research
United States Food and Drug Administration
10903 New Hampshire Avenue
Silver Spring, MD, 20993
United States
Phone: 1 2404028629
Email: [email protected]
Background: Adverse events associated with vaccination have been evaluated by epidemiological studies and more recently have gained additional attention with the emergency use authorization of several COVID-19 vaccines. As part of its responsibility to conduct postmarket surveillance, the US Food and Drug Administration continues to monitor several adverse events of special interest (AESIs) to ensure vaccine safety, including for COVID-19.
Objective: This study is part of the Biologics Effectiveness and Safety Initiative, which aims to improve the Food and Drug Administration’s postmarket surveillance capabilities while minimizing public burden. This study aimed to enhance active surveillance efforts through a rules-based, computable phenotype algorithm to identify 5 AESIs being monitored by the Center for Disease Control and Prevention for COVID-19 or other vaccines: anaphylaxis, Guillain-Barré syndrome, myocarditis/pericarditis, thrombosis with thrombocytopenia syndrome, and febrile seizure. This study examined whether these phenotypes have sufficiently high positive predictive value (PPV) to ensure that the cases selected for surveillance are reasonably likely to be a postbiologic adverse event. This allows patient privacy, and security concerns for the data sharing of patients who had nonadverse events can be properly accounted for when evaluating the cost-benefit aspect of our approach.
Methods: AESI phenotype algorithms were developed to apply to electronic health record data at health provider organizations across the country by querying for standard and interoperable codes. The codes queried in the rules represent symptoms, diagnoses, or treatments of the AESI sourced from published case definitions and input from clinicians. To validate the performance of the algorithms, we applied them to electronic health record data from a US academic health system and provided a sample of cases for clinicians to evaluate. Performance was assessed using PPV.
Results: With a PPV of 93.3%, our anaphylaxis algorithm performed the best. The PPVs for our febrile seizure, myocarditis/pericarditis, thrombocytopenia syndrome, and Guillain-Barré syndrome algorithms were 89%, 83.5%, 70.2%, and 47.2%, respectively.
Conclusions: Given our algorithm design and performance, our results support continued research into using interoperable algorithms for widespread AESI postmarket detection.
Introduction
The US Food and Drug Administration (FDA) Center for Biologics Evaluation and Research (CBER) is responsible for ensuring the safety, purity, potency, and effectiveness of biological products. This includes vaccines; allergenics; blood and blood products; and cell, tissue, and gene therapies for the prevention, diagnosis, and treatment of human diseases, conditions, or injuries [ 1 ]. The FDA’s history of safety surveillance for vaccines includes the creation and monitoring of the Vaccine Adverse Event Reporting System (VAERS). VAERS, jointly administered by the FDA and the Centers for Disease Control and Prevention (CDC), accepts spontaneous reports of suspected vaccine adverse events (AEs) after administration of any vaccine licensed in the United States.
VAERS has been successfully used as an early warning system to identify rare AEs; however, it has limitations. VAERS is a passive surveillance system that relies on individuals, patients, and clinical staff to send in reports, as opposed to automatically collecting them based on clinical data. This can lead to undercounting AEs. In addition, a causal relationship cannot be established using information from VAERS reports alone [ 2 ]. Because of VAERS’s limitations, more robust data systems are needed to enhance AE detection. These systems would be especially important for detecting the most severe AEs that require medical attention so that the FDA and CDC can offer guidance for these potentially life-threatening events and ensure that product labeling reflects known risks.
To address this gap, CBER established the Biologics Effectiveness and Safety Initiative (BEST) Initiative in 2017 to build data assets, analytics, and infrastructure for an active, large-scale, efficient postmarket surveillance system that can evaluate the safety and effectiveness of biologic products and develop innovative methods [ 3 ]. The BEST system is a collection of real-world data (RWD) sources: data related to patient health status and the delivery of health care that are routinely collected from several sources, such as electronic health record (EHR) or claims data [ 4 ]. EHR databases, specifically, are a rich source of information. They include data such as clinical notes, which can help address the limitations of VAERS. They also include entire populations of patients to identify whether cases are underreported. In addition, they may include patients’ entire clinical history, which can help establish a causal relationship for an AE. BEST has reached agreements with a limited number of foundational data partners. Access to these data partnerships does not fully address the possible undercounting of AEs of special interest (AESIs). However, these partnerships allow accelerated development and testing of AESI detection algorithms.
BEST is currently researching a system of distributed computable phenotype algorithms that could be applied at scale to many or all EHR systems across the United States to semiautomatically detect and report potential AESIs from RWD. Such a system could increase the speed and scope of AE surveillance beyond what is currently available to public health agencies through data partner agreements. To be candidate phenotypes for distributed surveillance use, the phenotypes need to identify probable AEs and avoid false detections. This reflects the need to balance the correct detection of AESIs with the protection of privacy and the reduction of burden on health provider systems. For the wider population of health providers to consider deploying such detection algorithms, these phenotype algorithms need to have reasonably high performance (measured by positive predictive value [PPV]) to ensure that the cases identified as AEs are likely to be verifiable cases with the outcome of interest. Toward this goal, the computable phenotypes in this study focus on existing EHR data reflecting a detected AE, which are reportable events for public health purposes. The algorithmic identification of undetected AEs or AEs that were not coded properly is beyond the scope of this study. Such research must include data from patients who had no AEs to fully evaluate the performance of a computable phenotype algorithm. Although scientifically desirable in the long term, the inclusion of non-AE cases falls outside of initial goals for a distributed surveillance system, which is assessing performance (measured by PPV) of the phenotypes for wide-scale surveillance purposes. The goal of distributing the phenotypes also poses limitations on designing the algorithms. Specifically, the components and complexity of the underlying algorithms need to take into account the current EHR standards and technology because they must be deployable and executable across EHR databases without imposing large overhead on health provider systems. If the phenotypes have sufficient PPV and are sufficiently easy to implement at health provider sites, the FDA could share the phenotypes to detect AESIs following vaccination in EHRs across the country, which could then be reported to the FDA for further review. The ability to detect AESIs using RWD could create an active surveillance system that enhances overall vaccine safety and helps make recommendations to minimize risks for postvaccination AESIs. The implementation of algorithmic detection and automated reporting of AESIs found in RWD has been shown to increase the odds of submitting a VAERS report by >30 times the preimplementation rate [ 5 ].
Although there is a history of studies around postvaccination AESIs, including those for influenza [ 6 - 8 ] and COVID-19 vaccines [ 9 - 13 ], there has been an increased interest in the analysis of vaccine safety and surveillance since the emergency use authorization (EUA) of 3 COVID-19 vaccines in the United States (Pfizer-BioNTech, Moderna, and Novavax) and their subsequent boosters (eg, bivalent boosters). The FDA hopes to contribute to this research through the development and performance validation of phenotypes for 5 postvaccination AESIs to identify potential vaccine safety events within EHR databases for this study. The 5 AESIs chosen include myocarditis/pericarditis, anaphylaxis, Guillain-Barré syndrome (GBS), intracranial or intra-abdominal thrombosis with thrombosis with thrombocytopenia syndrome (TTS), and febrile seizure. These AESIs were chosen because they are documented priorities of the CDC’s vaccine surveillance [ 14 ] for COVID-19 vaccine safety. In addition, several of these AESIs (anaphylaxis, GBS, and febrile seizure) are found following exposure to other vaccines, such as influenza; shingles; pneumococcal conjugate; and measles, mumps, and rubella. This study describes the methods to develop and validate these 5 computable phenotype algorithms on an EHR database and the validation results. It is part of the FDA’s efforts to improve postmarket surveillance and is valuable for public awareness, safety, and transparency.
Ethical Considerations
Ethical approval was not required for the study involving humans in accordance with the local legislation and institutional requirements. This study was part of the Sentinel activities conducted by the FDA as part of its postmarket surveillance duties. The Office of Human Research Protection (OHRP) in Health and Human Services (HHS) determined that the studies done under the Sentinel programs are not subject to regulation (45 CFR part 46) administered by OHRP. Written informed consent to participate in this study was not required from the participants or the participants’ legal guardians or next of kin in accordance with the national legislation and the institutional requirements.
Computable Phenotype Development
In total, 5 AESIs were selected to develop computable phenotypes for our validation study. The study’s main focus was detecting COVID-19 vaccine–related AESIs; therefore, we selected AESIs that the CDC specifically identified for monitoring after COVID-19 vaccination [ 14 ] or AESIs that have been reported for some subpopulations [ 15 ]. Given the uncertainty about the future use of COVID-19 seasonal boosters, the FDA also wanted to ensure that the AESIs selected had broad applicability to the safety surveillance of other widely used vaccines such as influenza; shingles; pneumococcal conjugate; diphtheria-tetanus-pertussis; and measles, mumps, and rubella. Three of our 5 selections met those criteria given in the CDC’s documented monitoring of anaphylaxis [ 16 ], GBS [ 17 ], and febrile seizures [ 18 ] for at least one of the vaccines listed.
The phenotype algorithms were designed to be relatively simple and interoperable so that any new health care organization’s IT department could translate and run them on their EHR database. They were built to query only structured data for interoperable, standard codes, such as Logical Observation Identifiers Names and Codes, Systematized Nomenclature of Medicine Clinical Terms, and RxNorm, so that the algorithm can be generalized or translated across different EHR systems. Historically, this has been a challenge for developing algorithms, since EHR databases often contain their own local code systems specific to the EHR vendor. For example, for this effort, we worked with the study partner to map Cerner Multum medication and observation codes to standard RxNorm and Logical Observation Identifiers Names and Codes, respectively.
Recent regulation now requires each EHR database to have an application programming interface (API) endpoint that translates any EHR data and many of the EHR’s proprietary codes to the United States Core Data for Interoperability (USCDI) implementation of the Fast Healthcare Interoperable Resources (FHIR) specification [ 19 ]. This specification requires the use of interoperable, published code lists [ 20 ] ( Table 1 ). These code systems cover almost all clinical events for the detection of AEs, such as medical diagnoses, medication prescriptions, laboratory tests or vital signs taken, and procedures performed. These APIs currently focus on supporting use cases where a single patient’s data are queried as opposed to aggregate searches across patients; therefore, we were unable to use them to identify the cohort that our phenotype would select. We were, however, able to use the FHIR API endpoints to pull data for each patient in our validation samples so that the participating clinicians could have data with the standard, interoperable code sets for their review.
| AESI | Description | Case definition reference |
| Myocarditis/pericarditis | Myocarditis and pericarditis are inflammatory processes involving the myocardium, pericardium, or both (myopericarditis). | Morgan et al [ ], 2008 |
| Anaphylaxis | Anaphylaxis is an acute hypersensitivity reaction with multiorgan system involvement that can present as, or rapidly progress to, a life-threatening reaction. It may occur following exposure to allergens from a variety of sources, including food, aeroallergens, insect venom, drugs, and immunizations. | Rüggeberg et al [ ], 2007 |
| GBS | GBS constitutes an important proportion of acute flaccid paralysis cases worldwide. It is a condition characterized by various degrees of weakness, sensory abnormalities, and autonomic dysfunction due to damage to peripheral nerves and nerve roots. | Sejvar et al [ ], 2011 |
| Intracranial or intra-abdominal TTS | Several cases of unusual thrombotic events and thrombocytopenia have developed after vaccination with the recombinant adenoviral vector encoding the spike protein antigen of SARS-CoV-2 (ChAdOx1 nCov-19, Astra Zeneca). More data were needed on the pathogenesis of this unusual clotting disorder [ ]. | Chen and Buttery Monashm [ ], 2021 |
| Febrile seizure | There is no Brighton Collaboration definition of febrile seizure, so we used both the fever and seizure case definitions. is defined as an elevation of body temperature above normal. It is usually caused by infection but can also be associated with several immunologic, neoplastic, hereditary, metabolic, and toxic conditions. Seizures are episodes of neuronal hyperactivity, most commonly resulting in sudden, involuntary muscular contractions. | Marcy et al [ ], 2004; Bonhoeffer et al [ ], 2004 |
a AESI: adverse event of special interest.
b GBS: Guillain-Barré syndrome.
c TTS: thrombosis with thrombocytopenia syndrome.
To facilitate health provider organizations’ ability to implement these queries on their EHR, phenotypes were rules based, used only certain types of structured data, and used common logic across AESIs. The general phenotype logic has been used previously for several postvaccination AESI studies at the FDA to identify potential AESI cases [ 28 , 29 ] and reuses concepts and methods from past literature from US-based collaborative health research groups, such as Observational Health Data Sciences and Informatics (OHDSI) [ 30 ], or from similar efforts in the United Kingdom [ 31 ] to develop computable phenotype libraries. A health organization only needs to write the general query logic once and then this logic would be able to detect different types of AESIs by referencing different lists of medical codes that represent the different medical events providing evidence that the various AESIs occurred. The logic common to all phenotypes is shown in Figure 1 . The code lists that we developed for necessary types of medical evidence are described in more detail below and listed in Table S1 in Multimedia Appendix 1 [ 21 - 23 , 25 - 27 ]. The circled items in Figure 1 represent a search for an FHIR resource element containing a code in one of the developed code lists. These were applied within the windows of time denoted by the brackets identifying windows of time before and after a condition diagnosis. The concepts in Figure 1 are described in additional detail below.

AESI Diagnoses and Problem List Items
The algorithm first looks for evidence of the AESI represented by a coded final or discharge diagnosis. Only final or discharge diagnoses are used since they best represent the ultimate determination of what was diagnosed during the patient’s care. The variability of admitting, working, and other diagnosis types lack the specificity required for the algorithm in this study.
Care Setting Filters
In addition, the care setting for every diagnosis was collected based on the medical encounter type for the diagnosis. All diagnosis care settings values were grouped into inpatient, outpatient, or emergency care setting types. Care setting was used to filter out diagnosis codes made during encounters with care settings unlikely to have the specific AESI diagnosis in the phenotype. The included care settings are defined by case definition and clinician input.
Clean Window
Next, a clean window (ie, a period before the coded diagnosis identified in step 1) is checked to ensure that the target diagnosis is the first known diagnosis of its type. This prevents the inclusion of historical or ongoing conditions. For all algorithms in this paper, the clean window is defined by all historical patient data in our data set. To make sure that all patient cases had at least a 1-year clean window, we pulled an additional historical year of data from our data partner before the study period. Cases where there were multiple occurrences of an AESI diagnosis suggested possible evidence of a chronic condition unrelated to vaccine exposure and thus were excluded.
Condition Window
Finally, the algorithm searches for sufficient supportive evidence within a condition window. The condition window is defined around the AESI diagnosis date and includes the entire medical encounter period when the condition was diagnosed, as well as 2 days before and 10 days after a condition is diagnosed. Clinical subject matter experts defined condition windows as the timeframe around a diagnosis that supportive evidence would likely present itself in the medical record.
Supporting Evidence
Within the condition window period, the algorithm may filter cases based on supporting evidence of an AESI. This filter looks for either laboratory test results found in observations, AESI treatment procedures, AESI treatment medications, or procedure or a combination of the 3 supporting evidence with a code that matches a code on to the phenotypes’ concepts code lists. These code lists in the lists aim to include all medical codes that could represent a particular concept, such as administration of epinephrine for an anaphylactic reaction. This AESI supporting evidence filter was applied to all phenotypes except for our febrile seizure AESI phenotype because a review of existing research [ 32 ] showed febrile seizure algorithms, in general, had the highest PPV among the selected AESIs. The concepts to build code lists for the supporting evidence were identified using case definitions. Following this, we prioritized improving specificity in the other AESI phenotypes by including filters requiring additional supporting evidence [ 32 - 35 ] and our clinician’s input.
Vaccine Exposure
In real-world operation, the algorithm would also include a vaccine exposure and risk window or a period surrounding vaccination in which diagnoses are searched. For the study’s purposes of having sufficient volume and statistical power to estimate operating characteristics of the algorithm, these exposure rules were not included.
Additional Details
Ideally, to assess whether these algorithms generalize to other sites, we would have a multisite validation study. Because of the high cost of data agreements, however, we only had data available for a single EHR site. To avoid overfitting and ungeneralizable results, we designed our algorithm development methods to only use our EHR data as a validation set and not use any of it to train, develop, or fine-tune the algorithm. While this does not remove the need for additional external validation, it reduces the likelihood of finding ungeneralizable results. To identify what medical concepts the algorithm should use as evidence, clinicians identified observations, medications, conditions, and procedure concepts from the AESI’s case definition, their relevant clinical experience, or other research from their literature review. A brief description of the AESI and the reference of the case definition used is captured in Table 1 , and additional information on the case definition is saved in Table S2 in Multimedia Appendix 1 [ 21 - 23 , 25 - 27 ].
An analyst completed a text search for a list of terms for these identified concepts, a list of which is captured in Table S2 in Multimedia Appendix 1 , to build the code lists of relevant codes from selected interoperable coding libraries ( Textbox 1 ) [ 21 - 23 , 25 - 27 ]. This was accomplished by searching the open-sourced OHDSI Observational Medical Outcomes Partnership concepts table and ATLAS tool (OHDSI community) [ 36 ], which is a collection of thousands of interoperable codes and their definitions and descriptions. The table was searched for any definition or description that matched the identified concept for the interoperable code systems that we listed in Textbox 1 and then was reviewed by a clinician for their suitability for the algorithm.
Clinical data and interoperable code lists used
- Diagnosis: International Classification of Diseases, Tenth Revision, Clinical Modification , Systematized Nomenclature of Medicine Clinical Terms
- Medication or immunization: National Drug Code, RxNorm
- Procedures: Current Procedural Terminology, International Classification of Diseases, 10th Revision Procedure Coding System
- Observations: Logical Observation Identifiers Names and Codes
The immunization and the diagnosis International Classification of Diseases, 10th Revision , clinical modification ( ICD-10 ) and Systematized Nomenclature of Medicine Clinical Terms code lists have been published on the Value Set Authority Center [ 37 ], and the additional observation, medication, and evidence code lists may be added in the future after this study is published.
For a surveillance use case, the algorithms need be run regularly (eg, daily or weekly) to collect batches of historical cases once all the data are available (as opposed to a real-time implementation to collect cases as they are happening). Because the algorithms were created to prioritize simplicity and interoperability rather than maximize total performance (eg, metrics beyond PPV such as sensitivity and, negative predictive power, etc), this study aimed for improved performance (measured by PPV) to existing AESI claims-based algorithms. Given our knowledge of how some crucial distinguishing information is part of unstructured clinical notes, which are not considered by the algorithms in this study, we expect further analysis is needed to improve accuracy [ 38 , 39 ]. Natural language processing techniques can improve algorithm performance but greatly increase the deployment complexity across health care organizations. Therefore, no natural language processing techniques were used for any phenotypes designed for this study.
Study Period
The study period spanned from January 1, 2018, through May 1, 2022, to ensure that the study’s data sampled patients both before and after the FDA issued the EUA and full licensure for COVID-19 vaccines. We also pulled at least 1 year of historical data for all patients; therefore, our data set includes historical information from January 1, 2017, to January 1, 2018, for all patients with medical encounters in the study period. Patients were included even if there were no clinical events in their historical period.
The study population came from a single academic health system in the United States, with EHR medical encounter data from >2.6 million patients and >20.7 million medical encounters for the study period. Table 2 shows the demographic breakdown for age, gender, race, and ethnicity of this population.
The entire EHR population during the study period was eligible to be selected by one of our developed phenotype algorithms. There were no age-related, medical condition–related, or other exclusions on the population for the algorithm to select cases. Clinical data necessary to select and validate cases selected by the algorithm were provided to the study team through a series of EHR data extracts for all patients in the study period. The algorithm required the following clinical data categories:
- demographic
- observation
| Category and demographic group | Patients, n (%) | |||
| <5 | 96,146 (3.6) | |||
| 5-17 | 224,941 (8.4) | |||
| 18-24 | 224,631 (8.4) | |||
| 25-44 | 840,395 (31.5) | |||
| 45-64 | 689,075 (25.8) | |||
| ≥65 | 591,497 (22.2) | |||
| Missing | 289 (0.01) | |||
| Male | 1,167,374 (43.8) | |||
| Female | 1,494,096 (56.1) | |||
| Missing | 5504 (0.2) | |||
| Black or African American | 748,746 (28.1) | |||
| American Indian or Alaska Native | 5834 (0.2) | |||
| Asian or Pacific Islander | 53,666 (2) | |||
| White | 1,030,834 (38.7) | |||
| Other | 198,265 (7.4) | |||
| Unknown | 629,608 (23.6) | |||
| Declined to answer | 21 (0) | |||
| Hispanic | 94,207 (3.5) | |||
| Non-Hispanic | 1,866,561 (70) | |||
| Unknown | 706,206 (26.5) | |||
EHR data extracts were mapped and loaded into an OHDSI Observational Medical Outcomes Partnership database [ 40 ]. Medication, observation, and procedure data extracts were requested and loaded into the database only for patients who would not be disqualified by other algorithm criteria. For patients selected to be in the validation sample, these data along with the clinical data for allergies, immunizations, and clinical notes were pulled from the EHR’s FHIR API endpoints, patient by patient, using a custom Python script to loop through the patients in the sample. The data were loaded into a Health Level 7 API (HAPI) FHIR server. We only pulled FHIR data for cases not initially disqualified by the vaccination and diagnosis filters to avoid unnecessary large data transfers and storage. The algorithm flagged potential AESIs that met the specified criteria. Samples of these cases were sent to physicians for validation.
Validation Sample
Once the algorithm identified cases, a random sample was drawn for each AESI for clinician adjudication. We used stratified sampling to ensure cases during pre– and post–COVID-19 EUA periods were represented ( Figure 2 ). This was due to concerns regarding potential confounding introduced by the COVID-19 vaccines, when attention to possible AESIs or medical charting of AESIs may have shifted. Where possible for each AESI, 100 cases were sampled from the pre–COVID-19 EUA period and 35 from the post–COVID-19 EUA period. If there were <100 or <35 cases during these periods, respectively, the sample would contain all cases the algorithm selected. Febrile seizure was the exception, as we believe the COVID-19 vaccine EUA should not affect the algorithm’s performance because febrile seizure AEs are usually associated with pediatric populations, and the COVID-19 vaccine was not approved for these populations during the study period [ 27 ].

Case counts sampled in each period were based on the incidence of diagnosis code occurrence within each period, as well as the period covered. In addition, we added negative controls selected randomly from every encounter in the period to establish a baseline comparison for the case validation process. We included negative controls as a quality control step to reduce the chance of quality issues with the data and to review the methods our clinicians were following and not for the purpose of making inferences about the phenotypes’ performance for non-AE cases (eg, through metrics such as sensitivity, negative predictive power, or an overall metric for performance). This study did not focus on the algorithmic identification of undetected AEs or AEs that were not coded properly. The focus of this study was to determine the phenotypes’ PPV. Given the expense of clinicians’ time for validations and the rarity of the AESIs, there would be minimum benefit to this study to have a negative control sample large enough to draw strong inferences. Furthermore, negative case controls would not further validate the utility of the phenotypes as tools for identifying probable AESIs through distributed surveillance. We added 20 negative controls from the pre–COVID-19 EUA period and 7 from the post–COVID-19 EUA period. Physicians were blinded to which cases were controls and which were not.
Chart Review Process
The sample of cases used to validate the algorithm was loaded into a chart review tool for clinician review. This allowed the clinicians to sort through the clinical information for a case and record the determination. Each case was assigned to 2 clinicians for review. The clinical validation used a patient’s full clinical history, which included EHR data, including all clinical notes for each case. The full EHR data used for clinician review included data unused by the detection algorithm described in the Computable Phenotype Development section, including different types of data (eg, allergies and clinical notes) and data filtered out (eg, admitting diagnosis and encounters with different care settings).
For each case, the clinician evaluated whether the clinical data evidence met the specified case definition criteria. Relevant patient data for the case window were available and presented to the clinicians in an easy-to-use, browser-based tool with a custom user interface. In the tool, clinicians were able to group items by type, search across all items and text, and request additional chart data to expand the window and access any available historical patient data, if desired.
All suspected AEs were validated using published case definitions [ 21 - 23 , 25 - 27 ] according to the levels of diagnostic certainty: level 1 (definite), level 2 (probable), and level 3 (possible). If a case did not meet one of the levels in the case definition, it was assigned as level 4 (doubtful) or level 5 (ruled out). “Ruled out” is distinct from “doubtful” in that “ruled out” cases have definitive evidence disqualifying them from being a correct diagnosis. If a case was determined to be “definite” or “probable,” it was considered a positive case of the AESI.
In the event of a disagreement between a positive and negative clinical review, a third clinician made a final determination by reviewing the case EHR data. If the clinicians found the structured or unstructured EHR data was insufficient, they marked this in their review by creating a level 3 (possible, insufficient evidence) designation, where an AESI could have occurred, but where there was not enough documentation to fulfill the requirements of the case definition.
Statistical Analysis
Ppv of algorithms.
Each algorithm’s PPV was the proportion of positive AEs the algorithm identified that were confirmed by clinical adjudication. PPVs were calculated for each AESI overall, as well as stratified by pre– and post–COVID-19 EUA periods and care setting (inpatient, emergency department, or outpatient). Sensitivity analyses were performed to evaluate the impact of medication use, different case definitions, and levels of evidence. PPVs were calculated in 2 different ways for each AESI algorithm. The first PPV calculated removed all possible cases with insufficient evidence from the denominator (cases labeled “definite” and “probable”/total cases minus any labeled “possible, insufficient evidence” by clinicians). PPV was then calculated with the cases with insufficient evidence added back into the denominator (cases labeled “definite” and “probable”/total cases). Reporting both PPV calculations can help with understanding the performance for different algorithm uses. Algorithm performance should ideally be compared with past literature of detection algorithms for the same AESI.
Because PPV is a binominal proportion, we calculated CIs for the PPV using the Agresti-Coull interval [ 41 ], which is the recommended method for estimating accurate CIs for binomial proportions such as PPV [ 42 ].
Interrater Reliability
Interrater reliability was used to measure the extent to which 2 physicians agreed in their AESI assessment. It was calculated using Cohen κ between the first 2 reviewers. Cohen κ measures the agreement between 2 raters classifying instances into mutually exclusive groups [ 43 ].
Stratification Analysis and Sensitivity Analysis
After validation was completed, we conducted a stratification and sensitivity analysis. We selected 2 stratification variables that could reasonably impact the generalizability of the results. First, we stratified the data by pre- and post-EUA date to confirm that the algorithm behavior did not change for AESIs after the COVID-19 vaccine was approved and administered to a large portion of the population. Ideally, the algorithms would perform consistently across these eras, but there are multiple factors that could impact the performance over these time periods. We also stratified the data by the care setting of the AE diagnosis, given that care settings may be associated with varying EHR data elements (eg, emergency departments compared with inpatient settings). Algorithm performance was computed using PPV within each stratum.
We also completed a post hoc sensitivity analysis where we investigated whether the algorithm could be improved, as measured by PPV, through small changes to it or by updating the process for evaluation. These changes were based on insights from clinicians or data analysts reviewing validation results, so results may not generalize to other data sets. However, we did attempt to limit our analysis to decisions that could have been feasibly made without postvalidation insights. The changes to the algorithms were either removing medications, observations, procedures, or diagnosis codes that are not specific enough to the AESI in question or adding logic to further filter out cases by requiring more supporting evidence ( Table 3 ).
The stratification or sensitivity analyses are meant as exploratory analyses to prompt additional research, but subgroups often have too small a sample size that have narrow enough CIs for meaningful results.
We also completed a sensitivity analysis on the GBS algorithm to calculate the PPV if we relaxed some of the specific case definition evaluation criteria and if more general evidence was available. We found that the 2 pieces of evidence that the case definition required were often missing in the chart review tool: lack of cerebrospinal fluid (CSF) white blood cell (WBC) count in cases of elevated CSF protein and limited or inconsistent documentation of diminished or absent reflexes. In some of these cases, we saw evidence that a neurologist was consulted and felt there was strong suspicion of GBS despite the missing documentation for these tests. This could be explained by 2 mechanisms.
First, and most likely, this could be due to data loss during the delivery or translation of EHR data to our chart review tool. Because we did not have direct access to the data, our process for obtaining, translating to different common data models or standards, and presenting the data to clinicians using the chart review tool could cause the data for these tests to be incorrectly mapped.
| AESI | Data type | Sensitivity analysis | Reasoning |
| Myocarditis/pericarditis | Medication | Removal of NSAIDs from our list of qualifying medication supporting evidence | NSAIDs are medications that can be used to treat many different conditions besides myocarditis and pericarditis. |
| Myocarditis/pericarditis | Diagnostic code | Stratification by diagnostic code (myocarditis vs pericarditis) | Diagnostic criteria differ for these related conditions and may lead to different performance. |
| GBS | Medication | Removal of gabapentin from our list of qualifying medication supporting evidence | Gabapentin was originally used as supporting evidence of a GBS episode due to its use for nerve pain associated with GBS events [ ]. However, it is also used for a variety of other conditions with neuropathic pain and is not specific to GBS. |
| GBS | Case definition | Update case definition criteria to allow for a case to be validated as positive if there is a missing documentation for absent or diminished reflexes in the weak limbs, CSF WBC count with neurology consult, or clinical note indicating evidence of the test result of GBS more generally | Documentation required for definite or probable GBS as defined by the case definition diagnosis was often missing from our data set due to failure to capture in EHR or failure to translate to our data set and can be supplemented by an expert’s judgment (eg, a neurologist). |
| Febrile seizure | Medication | Addition of medications used to treat fever | The original febrile seizure algorithm did not filter out cases without suggested evidence, but we believed adding suggested evidence could improve PPV . |
| Febrile seizure | Observation | Addition of observation of clinician describing the symptoms of seizure activity | The original febrile seizure algorithm did not filter out cases without suggested evidence, but we believed adding suggested evidence could improve PPV. |
| TTS | Diagnostic code | Stratification by most prevalent diagnostic code I81 versus all other codes | Diagnostic criteria differ for these related conditions and may lead to different performance. |
a NSAID: nonsteroidal anti-inflammatory drug.
c CSF: cerebrospinal fluid.
d WBC: white blood cell.
e EHR: electronic health record.
f PPV: positive predictive value.
g TTS: thrombosis with thrombocytopenia syndrome.
Second, case definition requirements for GBS are extremely strict, and physicians in this study believed that some of these might have represented valid GBS cases while not meeting every requirement. For example, several of the cases with missing CSF WBC count did mention cytoalbuminologic dissociation (or similar); in the presence of such a clinical statement, we might infer that CSF WBC count was performed and acceptable to meet the case definition criteria despite a missing test result.
Furthermore, in cases where a neurologist felt strongly that GBS was a likely diagnosis, along with other supporting evidence, it may be acceptable to rely on documented progressive and significant muscle weakness, especially with conflicting reflex findings. In these instances, we placed more weight on the clinician review (which may account for any unforeseen difficulties in data processing and the strictness of the case definition), not relying solely on the available (nonmissing) data types of the algorithm for assigning case diagnostic certainty.
Population Sample
Figure 2 illustrates the identification of the study populations and validation sample. From the study population of 20.7 million medical encounters for 2,666,974 patients over the study period, the algorithm selected 1195 (0.04%) cases of myocarditis/pericarditis, 550 (0.02%) of anaphylaxis, 123 (0.005%) of GBS, 626 (0.02%) of febrile seizure, and 395 (0.01%) of TTS. Of these patient cases (n=2,666,974), a stratified, random sample of 135 (0.01%) cases each was selected from myocarditis/pericarditis, 135 (0.01%) from anaphylaxis, and 135 (0.01%) from TTS populations. All 75 pre-EUA cases of GBS and a random sample of 35 post-EUA cases were selected to be validated. A random selection of 100 cases from the pre-EUA period were sampled to validate febrile seizure. An additional 27 negative control cases were sampled for each algorithm from the roughly 20.7 million medical encounters not selected by the algorithm in our study period. In total, 20 of these cases were sampled from the period before the COVID-19 vaccine EUA, and the remaining 7 came from the period after the EUA.
Overall PPV and Interrater Reliability Results
Table 4 presents algorithm performance measured by PPV for each of the 5 AESIs using cases that had sufficient evidence and all cases (ie, including cases unable to be confirmed as positive by clinicians due to insufficient evidence). Counts for the number of cases included in each PPV calculation can be found in Table S3 in Multimedia Appendix 1 [ 21 - 23 , 25 - 27 ].
Overall PPVs, when removing all cases with insufficient evidence, were highest for anaphylaxis (93.3%, 95% CI 86.4%-97%) and febrile seizure (89%, 95% CI 80%-94.4%), followed by myocarditis/pericarditis (83.5%, 95% CI 74.9%-89.6%) and TTS at unusual sites (70.2%, 95% CI 61.4%-77.6%). The lowest was for GBS (47.2%, 95% CI 35.8%-58.9%). All negative control cases across the 5 phenotypes were correctly classified by the algorithms.
The PPV results from the chart reviews of the validation sample for each AESI are reported for all cases as well as for only cases with sufficient evidence to make a clear by chart reviewers. The frequencies and percentages for insufficient evidence are presented with the stratification results in Table 5 . The interrater reliability scores for clinician chart reviews all showed substantial agreement between the clinicians ( Table 6 ). Interrater reliability, measured by Cohen κ, suggests substantial reliability when the value is >0.61, with many similar texts recommending a higher threshold of 0.80 [ 43 ].
| AESI and metric | Detected cases, PPV % (95% CI) | ||
| Cases with sufficient evidence only | 83.5 (74.9-89.6) | ||
| All cases | 63.7 (55.2-71.4) | ||
| Cases with sufficient evidence only | 93.3 (86.4-97) | ||
| All cases | 72.6 (64.4-79.5) | ||
| Cases with sufficient evidence only | 47.2 (35.8-58.9) | ||
| All cases | 30.9 (22.9-40.3) | ||
| Cases with sufficient evidence only | 70.2 (61.4-77.6) | ||
| All cases | 64.4 (55.9-72.1) | ||
| Cases with sufficient evidence only | 89 (80-94.4) | ||
| All cases | 89 (80-94.4) | ||
| AESI and metric | Detected cases | Pre-EUA period | Post-EUA period | Inpatient | Outpatient | Emergency department | |||||||
| Total cases, n | 135 | 100 | 35 | 91 | 26 | 18 | |||||||
| Total TP cases, n (PPV %; 95% CI) | 86 (63.7; 55.2-71.4) | 68 (68.0; 58.1-76.5) | 18 (51; 35-68) | 72 (79; 69-86) | 10 (38; 21-59) | 4 (22; 7-48) | |||||||
| Total cases with sufficient evidence, n (PPV % for TP cases with sufficient evidence; 95% CI) | 103 (83.5; 74.9-89.6) | 79 (86; 76-92) | 24 (75; 53-89) | 79 (91; 82-96) | 16 (63; 36-84) | 8 (50; 15-85) | |||||||
| Total cases, n | 135 | 100 | 35 | 27 | — | 108 | |||||||
| Total TP cases, n (PPV %; 95% CI) | 98 (72.6; 64.4-79.5) | 70 (70; 60.2-78.3) | 28 (80; 63-90.9) | 17 (63; 42.9-79.7) | — | 81 (75; 65.8-82.4) | |||||||
| Total cases with sufficient evidence, n (PPV %; 95% CI) | 105 (93.3; 86.4-97) | 74 (94.6; 86.2-98.4) | 31 (90.3; 73.4-98) | 19 (89.5; 65.6-99.7) | — | 86 (94.2; 86.6-97.9) | |||||||
| (n=110) | |||||||||||||
| Total cases, n (%) | 110 | 65 | 45 | 110 | — | — | |||||||
| Total TP cases, n (PPV %; 95% CI) | 34 (30.9; 22.9-40.3) | 24 (40; 25.9-49.5) | 20 (44; 30.4-59.4) | 34 (30.9; 22.8-40.3) | — | — | |||||||
| Total cases with sufficient evidence, n (PPV %; 95% CI) | 72 (47.2; 35.8-58.9) | 52 (46.2; 32.9-60) | 20 (50; 28.1-71.9) | 72 (47.2; 35.8-58.9) | — | — | |||||||
| (n=135) | |||||||||||||
| Total cases, n | 135 | 100 | 35 | 133 | 1 | 1 | |||||||
| Total TP cases, n (PPV %; 95% CI) | 87 (64.4; 55.9-72.1) | 64 (64; 54-72.9) | 23 (66; 48.2- 80) | 86 (64.7; 56.1-72.4) | 1 (100; 0-100) | 0 (0; 0-100) | |||||||
| Total cases with sufficient evidence, n (PPV %; 95% CI) | 124 (70.2; 61.4-77.6) | 91 (70.3; 60-78.9) | 33 (70; 51.6-83.5) | 122 (70.5; 61.7-78) | 1 (100; 0-100) | 1 (100; 0-100) | |||||||
| Total cases, n | 100 | 100 | — | 1 | — | 99 | |||||||
| Total TP cases, n (PPV %; 95% CI) | 73 (73; 63.3-80.9) | 73; (73; 63.3-80.9) | — | 0 (0; 0-100) | — | 73 (74; 64.1-81.6) | |||||||
| Total cases with sufficient evidence, n (PPV %; 95% CI) | 83 (88; 78.8-93.6) | 83 (88; 78.8-93.6) | — | 1 (0; 0-100) | — | 82 (89; 80-94.4) | |||||||
b EUA: emergency use authorization.
c TP: true positive.
d PPV: positive predictive value.
e Not applicable.
f GBS: Guillain-Barré syndrome.
| AESI | Total cases validated, n | Interrater reliability |
| Myocarditis/pericarditis | 162 | 0.814 |
| Anaphylaxis | 162 | 0.770 |
| GBS | 137 | 0.832 |
| TTS at unusual sites | 162 | 0.851 |
| Febrile seizure | 120 | 0.965 |
Stratification
To evaluate consistency across pre- and post-EUA periods and care settings, we reported true positive and PPV results for each stratum ( Table 5 ).
None of the algorithms had notable differences between the pre- and post-EUA periods since all 95% CIs had some overlap. However, there were some differences between the PPVs for the 2 periods that could be significant with a larger validation sample. The difference in PPV for myocarditis/pericarditis varied from 68% in the pre-EUA period to 51.4% in the post-EUA period, while anaphylaxis showed the opposite pattern with a 70% PPV in the pre-EUA period that increased to 80% PPV in the post-EUA period.
We also reported stratified results by care setting ( Table 5 ). For myocarditis/pericarditis, the PPV of cases with an inpatient care setting (79.1%, 95% CI 69.4%-86.4%) was notably higher than that from the outpatient (38.5%, 95% CI 21.2%-58.8%) or emergency department (22.2%, 95% CI 6.7%-47.9%) care settings.
Anaphylaxis did not have a large difference across care settings, as the 95% CIs overlapped between the 2 care settings. However, they did show better performance with cases in an emergency department (PPV 75%, 95% CI 65.8%-82.4%) care setting over cases with an inpatient care setting (PPV 63%, 95% CI 42.9%-79.7%). The other AESI algorithms filtered for only 1 care setting or had a vast majority of cases in 1 care setting.
Sensitivity Analysis
Medication and observation algorithm changes.
We analyzed whether changes to medication code lists for the myocarditis/pericarditis and GBS algorithms could improve performance. For the myocarditis/pericarditis algorithm, removal of nonsteroidal anti-inflammatory drugs from the medication code lists showed no change in PPV at 83.5% ( Table 7 ), but PPV values were higher for cases selected with the pericarditis instead of myocarditis ICD-10 codes.
For the GBS algorithm, when cases were removed where gabapentin (used for post-GBS pain management) was the only supporting evidence, PPV increased to 38.1% (95% CI 28.2%-49.1%) from 30.9% (95% CI 22.9%-40.3%; Table 8 ).
Our initial febrile seizure algorithm did not use any supporting evidence to filter out possible false positives since we believed we could get adequate PPV without it.
For our sensitivity analysis, we tested requiring supporting evidence in the condition period, such as the presence of medications for reducing fever such as acetaminophen, observation evidence when the patient’s chief complaint was related to fever or seizure, or the presence of both. When filtered to only cases with either medication or observation evidence, febrile seizure PPV increased significantly to 93.3% (95% CI 84.7%-97.6%) from the original algorithm PPV of 73% (95% CI 63.3%-80.9%), with no overlap in 95% CIs and a P value of <.001 ( Table 9 ). When the algorithm required both medication and observation evidence, it performed even better (PPV 96.9%, 95% CI 88.5%-99.9%).
| AESI and sensitivity analysis | Total TP cases, n | Selected cases, n (change, n) | PPV , % (95% CI; change) | Selected cases with sufficient evidence, n (change, n) | PPV, % (95% CI; change) |
| Removal of NSAIDs | 86 | 135 (0) | 63.7 (55.2-71.4; 0) | 103 (0) | 83.5 (74.9-89.6; 0) |
| Pericarditis diagnosis | 59 | 82 (–53) | 72 (61.1-80.8; +8.3) | 67 (–36) | 88.1 (77.6-94.3; +4.6) |
| Myocarditis diagnosis | 27 | 53 (–82) | 50.9 (37.4-64.3; –12.8) | 36 (–67) | 75 (57.9- 87.1; –8.5) |
b TP: true positive.
c PPV: positive predictive value.
d Values in parentheses reflect the change due to the modified algorithm features.
e NASID: nonsteroidal anti-inflammatory drug.
f All International Classification of Diseases, Tenth Revision, Clinical Diagnosis codes that the algorithm used were broken into 2 groups: myocarditis (I40.0 infective myocarditis, I40.1 isolated myocarditis, I40.8 other acute myocarditis, I40.9 acute myocarditis, unspecified, and I51.4 Viral myocarditis) and pericarditis (B33.22 viral pericarditis, B33.23 acute rheumatic pericarditis, I30.0 acute nonspecific idiopathic pericarditis, I30.1 infective pericarditis, I30.8 other forms of acute pericarditis, I30.9 acute pericarditis, unspecified, I32 pericarditis in diseases classified elsewhere, and I41 meningococcal pericarditis).
| AESI and sensitivity analysis | Total TP cases | Selected cases (change) | PPV , % (95% CI; change) | Selected cases with sufficient evidence (change) | PPV, % (95% CI; change) |
| Removal of gabapentin | 33 | 86 (–24) | 38.4 (28.6-49.2; 7.5) | 53 (–19) | 62.3 (48.3-74.5; +15) |
| Adjusted case definition | 49 | 110 (0) | 44.5 (35.4-54; +13.6) | 72 (0) | 68.1 (56.3-78; +20.8) |
| Adjusted case definition+removal of gabapentin | 49 | 86 (–26) | 57.1 (46.2-67.4; +26.2) | 68 (–4) | 72.1 (60-81.6; +24.8) |
| AESI and sensitivity analysis | Total TP cases, n | Selected cases, n (change, n) | PPV , % (95% CI; change) | Selected cases with sufficient evidence, n (change, n) | PPV , % (95% CI; change) |
| Cases with either medication or observation | 70 | 75 (–25) | 93.3 (84.7-97.6; +20.3) | 73 (–10) | 95.9 (87.9-99.2; +7.9) |
| Cases with both medication and observation evidence | 63 | 65 (–35) | 96.9 (88.5-99.9; +23.9) | 63 (–20) | 100 (92.8-100; +12) |
c Values in parentheses reflect the change due to the modified algorithm features.
Diagnostic Code List Changes
We also analyzed if changing diagnostic codes that were used to identify the AESI might lead to higher performance for the myocarditis/pericarditis and TTS algorithms.
For myocarditis/pericarditis, we found that an algorithm only looking for the myocarditis code (PPV 50.9%, 95% CI 37.4%-64.3%) underperformed an algorithm with just pericarditis codes (PPV 72%, 95% CI 61.1%-80.8%; Table 7 ). For TTS, we found that the main ICD-10 code I81 for “portal vein thrombosis” (73.5%, 95% CI 64%-81.3%) outperformed all other codes in our code list, including G08 (intracranial and intraspinal phlebitis and thrombophlebitis), I82.0 (Budd-Chiari syndrome), I82.3 (embolism and thrombosis of renal vein), and I82.890 (acute embolism and thrombosis of other specified veins), with a PPV of 36.4% (95% CI 21.3%-54.4%; Table 10 ).
| AESI and sensitivity analysis | Total TP cases, n | Selected cases, n (change, n) | PPV , % (95% CI; change) | Selected cases with sufficient evidence, n (change, n) | PPV, % (95% CI; change) |
| I81 | 75 | 102 (–33) | 73.5 (64-81.3; +9.1) | 96 (–28) | 78.1 (68.6-85.4; +8) |
| All other TTS codes | 12 | 33 (–102) | 36.4 (21.3-54.4; –28) | 28 (–96) | 42.9 (25.4-62.1; –27.3) |
e ICD: International Classification of Diseases.
f All other TTS ICD codes include G08, I82.0, I82.3, and I82.890.
Case Definition Validation Criteria
Finally, we analyzed whether a small update to our case definition criteria for the GBS algorithms described in the Stratification Analysis and Sensitivity Analysis section would improve reported performance in Table 7 . When we applied both changes, the validation criteria change to the algorithm and removal of gabapentin, as discussed in the Medication and Observation Algorithm Changes section, the algorithm achieved a PPV of 57.1% (95% CI 46.2%-67.4%).
Principal Findings
The results of this study show that for 4 out of 5 AESIs, we can build an interoperable computable phenotype with comparable or increased performance to algorithms in the existing literature. These algorithms are developed using a rules-based approach to facilitate their application and increase the generalizability of performance across EHR databases. For the phenotypes with poorer performance, the issues were often that the case definition required documentation of a test that was lost in our data pipeline, or was not completed, or was not recorded by the treating physician or nurse. While these cases are marked as false positives based on our methodology, they may be true AEs that are lacking the documentation to meet the case definition. Some small updates to the algorithms or the case definition evaluation method could be made to potentially improve the algorithms’ performances, but a more important next step would be to validate our algorithms on other data partners to ensure generalizability of the original algorithms and any updates. Given the need for active AE surveillance, this study is still an important first step toward building an algorithm that can be distributed and implemented on health provider EHR databases and can accurately detect AEs.
The PPV results of the phenotypes, negative control groups, and stratification and sensitivity analysis are discussed in more detail in the following sections. Note our negative control groups and many of the stratification and sensitivity analyses have sample sizes too small to draw strong conclusions as illustrated by the width of the 95% CIs for those results. These were exploratory analyses completed as a supplement to the main findings of the study around the PPV of the algorithms.
Myocarditis/Pericarditis
The myocarditis/pericarditis algorithm showed strong PPV performance using cases with sufficient evidence. The literature appears to lack good comparison studies against which to evaluate this algorithm’s performance. A meta-analysis from 2013 reviewed myocarditis/pericarditis algorithm studies and found that none of them evaluated their algorithm by calculating PPV [ 45 ].
When myocarditis/pericarditis was segmented via care settings, algorithm performance was highest for inpatient settings, with a PPV of 79.1%. This can be attributed to the availability of supporting clinical data needed for accurate case detection in such settings. Given that inpatient testing is necessary to meet the criteria of the case definition, the algorithm performance matches clinical expectations and adds to its public health importance.
In emergency care settings, myocarditis/pericarditis is often diagnosed for patients with a history of inpatient visits to one or more other health systems. This increases the probability of these patients having additional documentation necessary to meet the case definition. This highlights the role of health information exchanges in supporting public health use cases, improving AE reporting, and enhancing postmarket surveillance.
Myocarditis/pericarditis had a notable difference in PPVs for pre- and post-EUA date. The post-EUA date strata of the sample had a higher percentage of cases coming from the emergency department, which had few cases before EUA. This could be explained by patients being diagnosed during previous inpatient stays in other health systems and a lower threshold to provide a preliminary diagnosis with limited information. This category had a lower PPV on average for myocarditis/pericarditis, likely due to less documentation in an emergency care setting than in an inpatient care setting. This highlights the need for further validation of the algorithm in these settings for an effective public health benefit and to gain confidence that our algorithm is fit for purpose. Because the aim of the algorithms is postvaccination AESI detection in support of public health safety surveillance, any potential degradation in performance in the post-EUA period is a concern. If performance decrease in the post-EUA period is driven by postvaccination myocarditis/pericarditis being more likely to have confounding physical findings that could affect how quickly and in which care setting it gets diagnosed, the PPV from this study may not be applicable to a postvaccination version of the phenotype. There is a small overlap in the 2 periods’ PPV 95% CI, and a 2-sample proportion test returns a P value of .08. This suggests that the difference could also be due to statistical noise. However, given the importance of the post-EUA period to the algorithm’s future task and the size of the difference, we suggest validating additional cases in the post-EUA period to confirm whether the algorithm is actually less effective.
Anaphylaxis
In cases with sufficient evidence, our anaphylaxis algorithm performed strongly with a PPV score of 93.3% (95% CI 86.4%-97%). This shows a possible slight improvement over previous anaphylaxis research, although both results were within the 95% CI [ 33 , 34 ]. When stratified by care setting, the algorithm performed better in emergency department care settings. This can be explained due to the anaphylaxis symptoms and treatment being more likely to be well-documented in this setting. Availability of additional evidence increases the PPV of the algorithm. Since anaphylaxis cases related to vaccination are more likely to culminate in visits to the emergency department, the better performance of the algorithm would provide a better public health benefit.
Overall, the performance of the algorithm was moderate compared with that seen in literature. With no obvious avenues for improvement available, no additional sensitivity analyses were applied.
GBS Algorithm
Our initial GBS algorithm showed weak performance for GBS with a PPV of 47.2% (95% CI 35.8%-58.9%). Given existing research on GBS validations, this result is not surprising, since our result is comparable with a study result showing GBS algorithm validation PPV of 29% (95% CI 24%-34%) [ 35 ]. We hoped that our algorithm would improve on this study’s results, allowing us to meet the “moderate” performance threshold defined in the Methods section, given that we added additional logic to require suggested evidence and filter out historical diagnoses. However, we believe that the algorithm’s performance could be improved based on the sensitivity analysis results.
An increase in performance was observed when adjusting the case definition interpretation of GBS to allow for more general written clinical notes or neurology consult evidence to replace specific documented test results. The lack of standardization in laboratory results is fraught with challenges such as inconsistent data. The observed improvement in the GBS phenotype highlighted the need for further standardization to have a better impact on public health benefit.
Furthermore, the performance of the GBS algorithm was improved by the exclusion of nonspecific medications such as gabapentin, increasing its public health benefit. Gabapentin is often used to treat generalized neuropathic pain for a variety of conditions other than GBS, including diabetes, and can confound the results.
With both case definition and medication adjustments to the algorithm, the PPV rose to be closer to the moderate performance threshold and an increase over the cited historical study [ 35 ]. Because these changes were informed by the cases in the validation study post hoc, they might be overfitted to this validation sample and may not be generalizable. They should be tested in other EHR systems.
The GBS algorithm performed slightly better in the post-EUA period, but the performance of both periods was well within the 95% CI of the other period. The GBS algorithm only applies to the inpatient care setting; therefore, no care setting stratification analysis was performed.
Febrile Seizure
Our febrile seizure algorithm performed strongly, with a PPV score of 89% using cases with sufficient evidence. This performance is in line with existing febrile seizure algorithm validation research [ 32 ], where a febrile seizure validation study on the FDA Sentinel database showed a PPV of 70% (95% CI 64%-76%). Our sensitivity analysis suggests that even better performance could possibly be achieved by adding additional filters to select cases with supporting medication and observation evidence, which are well-documented in EHRs. The better performance of the algorithm provides better public health benefits and further supports the use of EHRs in the detection of AEs. For cases that met either or both criteria, the PPV increased. Since these changes to the algorithm happened after the validation was completed, they overstate the general performance increases when applied to a new EHR setting but offer avenues for a future validation study. Future research can test whether stronger performance is possible with these filters and focus on reviewing the algorithm’s application to AEs following pediatric vaccinations.
TTS Algorithm
The TTS algorithm showed moderate performance for PPV at 70.2% which is similar to a separate FDA TTS validation study which estimated the performance at 76.1% (95% CI 67.2-83.2%) [ 29 ]. TTS had consistent performance across both pre- and post-EUA periods and did not have enough cases in the outpatient and emergency department care settings for any defensible findings around diagnosis care setting stratification. Our sensitivity analysis revealed that when the AESI was diagnosed with the ICD-10 code I81 (portal vein thrombosis), the algorithm showed a significant increase when compared with the performance of all other ICD codes (73.5%, 95% CI 64%-81.3%, compared with 36.4%, 95% CI 21.3%-54.4%). Although if an increase to specificity is desired at the cost of some sensitivity, the TTS algorithm could be limited to only select the higher performing I81 diagnosis code.
Limitations
There are several limitations to this study. First, it only evaluates general AESIs and not postvaccination AESIs specifically since the algorithms do not require the evidence of vaccine administration as criteria. While this was necessary due to the rareness of the postvaccination AESIs in our data, it is possible that the algorithms perform worse detecting postvaccination AESIs specifically since they will often present slightly differently in different populations when occurring after a vaccine administration. For example, the major presenting symptoms appeared to resolve faster in cases of myocarditis after COVID-19 vaccination than in typical viral cases of myocarditis [ 9 ]. To guard against this, we included both pre– and post–COVID-19 EUA data with the hope that post-EUA cases would include some postvaccination AESIs. However, we did not have enough post-EUA cases available to build a large enough sample size for a comparison with sufficient statistical power to provide definitive evidence on this topic. Another limitation in this vein is the general small sample size for all stratification, sensitivity, and negative control analyses. We make sure to state that these analyses are exploratory in nature, and the reader should not form strong conclusions from them given their small samples size and large CI range. Future research could address these concerns by identifying a data source with enough postvaccination AESI cases to complete a comparably large validation study.
An additional limitation of this study is that it only measures algorithms’ PPVs instead of investigating other metrics that could give a better picture of the algorithm’s holistic performance such as sensitivity and specificity. Specifically, these other metrics would estimate how many of the total positive cases are being identified and how well the algorithm is able to identify cases without the AESIs. However, we believe that this limitation is necessary for the following reasons: (1) the main purpose of this study was to assess the PPV of phenotypes because it answers the most relevant public health question, if the algorithms will generate a quality detected set of AE cases for the public health surveillance and (2) a much higher cost and more extensive data sharing are needed to properly estimate sensitivity and specificity because of the required validation sample size necessary for a negative control group. To calculate PPV, one only needs a sample of the cases selected by the algorithm. To estimate the sensitivity and specificity, however, it would be necessary to also validate an extremely large negative control group sample since the AESI conditions that the algorithms try to detect are often rare events. We would expect it to be even more rare for these conditions of interest for AESIs to happen and not be recorded with types of structured data elements that are being used in the phenotypes. In fact, the lack of structured data elements in some negative control cases led to a clinician asking the research team if something was wrong because their case had no relevant charted events to be reviewed. A much larger validation study would also expose clinicians to a larger set of patient data for cases that have a low likelihood of having an AE. This approach limits the interaction with protected health information data until the algorithms’ PPVs support continued research with broader samples and methodologies.
Another limitation is that although they were designed to be simple to deploy, the algorithms are still time-consuming to apply to different EHR systems. Although a hallmark of this algorithm is its interoperability, the algorithm logic still must be applied to the EHR common data model or extracted and translated into another common data model as was done for this study. Interoperable codes should be available for all patients, given the requirement to provide patient data in an interoperable FHIR standard. However, given the recency of the requirement, they might not be available in all systems and require some code translation on the health organization side, especially when analyzing at the population level. In addition, since the interoperable codes will only be available through a FHIR API, this adds another data pull and integration with the EHR system to obtain these codes for the algorithm.
In the future, the evolving landscape of health IT may facilitate the public health use cases of detecting and reporting postvaccination AESIs in a safe and secure manner that protects patient privacy. This could be achieved by EHRs supporting secure querying of patient cohorts with probable postvaccination AESIs using clinical query language [ 46 ] or other interoperable query language. Reducing the burden of automatic detection of postvaccination AESIs would help public health organizations improve AE surveillance with minimal additional burden to health care organizations and providers.
A final limitation of this study is that the algorithms were only applied to 1 site. Going forward, algorithm performance should be validated at other sites to ensure their generalizability. Although the algorithms were generated without prior input from the data, the study is still limited to 1 health care organization, and this method could have different operating characteristics (PPV, sensitivity, etc) at a second location.
Future research can be performed to improve algorithm accuracy and as stated previously would require additional partner EHR data systems. To create a better performing algorithm, machine learning techniques could be used to train the model to identify specific patterns of data instead of relying on rules-based methods that incorporate published case definition criteria and clinical subject matter expert experience. When given enough data, machine learning approaches generally outperform rules-based approaches across domains, and some prior research suggests that this is true in the medical domain as well [ 47 ].
However, machine learning methods will not generalize across EHR systems because the data patterns that machine learning identifies could be specific to an individual health care organization. Trying to build a large data set that combines multisite data is extremely difficult and costly due to concerns over infrastructure, regulations, privacy, and data standardization. A method such as federated learning could be explored to alleviate this problem. Federated learning allows multiple sites to collaboratively train a global model without directly sharing data and has been used to train machine learning algorithms at EHR sites previously [ 48 ].
Conclusions
In summary, this study presents strong initial evidence that creating simple, interoperable, rules-based phenotypes can detect AESIs on a new data source and that the phenotypes outperform the PPV outcomes for historical validations studies for these conditions. The study validates 5 different AESIs to prove that this approach can work for a broad range of AESIs, while also highlighting where the approach might be less successful. For example, the GBS algorithm was built using ICD-10 codes that previous validation studies have demonstrated are not accurate predictors of a GBS case that meets case definition criteria; subsequently, our GBS algorithm performed poorly. The validation study sample sizes for all AESIs allowed for adequate precision to evaluate algorithm PPV against historical studies.
An active surveillance system can enhance vaccine safety and aid in the development and use of safer vaccines and recommendations to minimize the AE risks after vaccination [ 49 ]. The algorithms were developed using a method that should be able to be applied to and generalize performance for new EHR databases, but more research is needed to confirm this. If the methodology can be successfully used to detect postvaccination AESI cases across EHR databases, these algorithms could be deployed widely to inform FDA decision-making, promote public safety, and improve public confidence. Going forward, further research and investigation are needed to enhance algorithm performance and integrate the algorithms across health care organizations for active surveillance in the interest of public health.
Acknowledgments
Development of the manuscript benefitted from significant engagement with the Food and Drug Administration (FDA) Center for Biologics Evaluation and Research (CBER) team members and their partners. The authors thank them for their contributions. Additional feedback on the manuscript was provided by IBM Consulting (Stella Muthuri and Brian Goodness), Accenture Consulting (Shayan Hobbi), and Korrin Bishop (writing and editing). This research was funded through the FDA CBER Biologics Effectiveness and Safety Initiative. Several coauthors hold commercial affiliations with Accenture, IBM Consulting, and MedStar Health Research Institute. Accenture (PSH); IBM Consulting (AAH, JP, JB, AS, EM, LDJ, and MD); and MedStar Health Research Institute (AZH and JB) provided support in the form of salaries for authors but did not have any additional role in the study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Data Availability
The data sets generated during and analyzed during this study are not publicly available, and were only made available to the Food and Drug Administration for the purpose of evaluating algorithms for adverse events of special interest outcomes. For inquiries or questions regarding the data, individual queries should be directed to the corresponding author.
Conflicts of Interest
Authors AAH, JP, JB, AS, EM, LDJ, and MD are or were employed by IBM while participating in the study. PSH is employed by Gevity Consulting, Inc, a part of Accenture. Authors AZH and HJB are employed by MedStar Health Research Institute, and AZH holds an appointment with Georgetown University School of Medicine. These authors have delivered clinical and epidemiology consulting engagement for public and private sector partners. These affiliations did not impact the study design, data collection and analysis, decision to publish, or preparation of the manuscript and do not alter our adherence to JMIR policies on sharing data and materials. The opinions expressed are those of the authors and do not necessarily represent the opinions of their respective organizations. The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Search terms and code lists for 5 developed phenotypes and detailed case definitions.
- CBER vision and mission. U.S. Food and Drug Administration. Sep 25, 2019. URL: https://www.fda.gov/about-fda/center-biologics-evaluation-and-research-cber/cber-vision-mission [accessed 2024-02-22]
- Vaccine adverse event reporting system (VAERS). Centers for Disease Control and Prevention. URL: https://www.cdc.gov/vaccinesafety/ensuringsafety/monitoring/vaers/index.html#anchor_1616772696807 [accessed 2024-02-22]
- CBER biologics effectiveness and safety (BEST) system. U.S. Food and Drug Administration. URL: https://www.fda.gov/vaccines-blood-biologics/safety-availability-biologics/cber-biologics-effectiveness-and-safety-best-system [accessed 2024-02-22]
- Corrigan-Curay J. Framework for FDA’s real-world evidence program. U.S. Food and Drug Administration. Mar 15, 2019. URL: https://www.fda.gov/media/123160/download [accessed 2024-02-22]
- Baker MA, Kaelber DC, Bar-Shain DS, Moro PL, Zambarano B, Mazza M, et al. Advanced clinical decision support for vaccine adverse event detection and reporting. Clin Infect Dis. Sep 15, 2015;61(6):864-870. [ FREE Full text ] [ CrossRef ] [ Medline ]
- Polakowski LL, Sandhu SK, Martin DB, Ball R, Macurdy TE, Franks RL, et al. Chart-confirmed Guillain-Barre syndrome after 2009 H1N1 influenza vaccination among the Medicare population, 2009-2010. Am J Epidemiol. Sep 15, 2013;178(6):962-973. [ CrossRef ] [ Medline ]
- Martin D, Menschik D, Bryant-Genevier M, Ball R. Data mining for prospective early detection of safety signals in the Vaccine Adverse Event Reporting System (VAERS): a case study of febrile seizures after a 2010-2011 seasonal influenza virus vaccine. Drug Saf. Jul 2013;36(7):547-556. [ CrossRef ] [ Medline ]
- Erlewyn-Lajeunesse M, Bonhoeffer J, Ruggeberg JU, Heath PT. Anaphylaxis as an adverse event following immunisation. J Clin Pathol. Jul 2007;60(7):737-739. [ FREE Full text ] [ CrossRef ] [ Medline ]
- Oster ME, Shay DK, Shimabukuro TT. Myocarditis cases after mRNA-based COVID-19 vaccination in the US-reply. JAMA. May 24, 2022;327(20):2020-2021. [ CrossRef ] [ Medline ]
- Le Vu S, Bertrand M, Jabagi MJ, Botton J, Drouin J, Baricault B, et al. Age and sex-specific risks of myocarditis and pericarditis following Covid-19 messenger RNA vaccines. Nat Commun. Jun 25, 2022;13(1):3633. [ FREE Full text ] [ CrossRef ] [ Medline ]
- Hanson KE, Goddard K, Lewis N, Fireman B, Myers TR, Bakshi N, et al. Incidence of Guillain-Barré syndrome after COVID-19 vaccination in the vaccine safety datalink. JAMA Netw Open. Apr 01, 2022;5(4):e228879. [ FREE Full text ] [ CrossRef ] [ Medline ]
- Kim JE, Min YG, Shin JY, Kwon YN, Bae JS, Sung JJ, et al. Guillain-Barré syndrome and variants following COVID-19 vaccination: report of 13 cases. Front Neurol. Jan 27, 2021;12:820723. [ FREE Full text ] [ CrossRef ] [ Medline ]
- MacIntyre CR, Veness B, Berger D, Hamad N, Bari N. Vaccine. Aug 09, 2021;39(34):4784-4787. [ FREE Full text ] [ CrossRef ] [ Medline ]
- Selected adverse events reported after COVID-19 vaccination. Centers for Disease Control and Prevention. URL: https://www.cdc.gov/coronavirus/2019-ncov/vaccines/safety/adverse-events.html [accessed 2024-02-22]
- COVID-19 vaccine safety in children and teens. Centers for Disease Control and Prevention. URL: https://www.cdc.gov/coronavirus/2019-ncov/vaccines/vaccine-safety-children-teens.html [accessed 2024-02-22]
- Preventing and managing adverse reactions: general best practice guidelines for immunization. Centers for Disease Control and Prevention. URL: https://www.cdc.gov/vaccines/hcp/acip-recs/general-recs/adverse-reactions.html [accessed 2024-02-22]
- Guillain-Barré syndrome and vaccines. Centers for Disease Control and Prevention. URL: https://www.cdc.gov/vaccinesafety/concerns/guillain-barre-syndrome.html [accessed 2024-02-22]
- Febrile seizures and childhood vaccines. Centers for Disease Control and Prevention. URL: https://www.cdc.gov/vaccinesafety/concerns/febrile-seizures.html [accessed 2024-02-22]
- Standards-based application programming interface (API) certification criterion. The Office of the National Coordinator for Health Information Technology. 2020. URL: https://www.healthit.gov/sites/default/files/page2/2020-03/APICertificationCriterion.pdf [accessed 2024-02-22]
- US core data for interoperability USCDI. Health Level 7 Fast Health Interoperability Resources. URL: https://build.fhir.org/ig/HL7/US-Core/uscdi.html [accessed 2024-02-22]
- Morgan J, Roper MH, Sperling L, Schieber RA, Heffelfinger JD, Casey CG, et al. Myocarditis, pericarditis, and dilated cardiomyopathy after smallpox vaccination among civilians in the United States, January-October 2003. Clin Infect Dis. Mar 15, 2008;46 Suppl 3(s3):S242-S250. [ CrossRef ] [ Medline ]
- Rüggeberg JU, Gold MS, Bayas JM, Blum MD, Bonhoeffer J, Friedlander S, et al. Anaphylaxis: case definition and guidelines for data collection, analysis, and presentation of immunization safety data. Vaccine. Aug 01, 2007;25(31):5675-5684. [ CrossRef ] [ Medline ]
- Sejvar JJ, Kohl KS, Gidudu J, Amato A, Bakshi N, Baxter R, et al. Guillain-Barré syndrome and Fisher syndrome: case definitions and guidelines for collection, analysis, and presentation of immunization safety data. Vaccine. Jan 10, 2011;29(3):599-612. [ CrossRef ] [ Medline ]
- Greinacher A, Thiele T, Warkentin TE, Weisser K, Kyrle PA, Eichinger S. Thrombotic thrombocytopenia after ChAdOx1 nCov-19 vaccination. N Engl J Med. Jun 03, 2021;384(22):2092-2101. [ FREE Full text ] [ CrossRef ] [ Medline ]
- Chen R, Buttery Monashm J. DRAFT - TTS: case definition and guidelines for data collection, analysis, and presentation of immunization safety data. Zenodo. Nov 11, 2021. URL: https://zenodo.org/records/6697333 [accessed 2024-07-01]
- Marcy SM, Kohl KS, Dagan R, Nalin D, Blum M, Jones MC, et al. Fever as an adverse event following immunization: case definition and guidelines of data collection, analysis, and presentation. Vaccine. Jan 26, 2004;22(5-6):551-556. [ CrossRef ] [ Medline ]
- Bonhoeffer J, Menkes J, Gold MS, de Souza-Brito G, Fisher MC, Halsey N, et al. Generalized convulsive seizure as an adverse event following immunization: case definition and guidelines for data collection, analysis, and presentation. Vaccine. Jan 26, 2004;22(5-6):557-562. [ CrossRef ] [ Medline ]
- Shoaibi A, Lloyd PC, Wong HL, Clarke TC, Chillarige Y, Do R, et al. Evaluation of potential adverse events following COVID-19 mRNA vaccination among adults aged 65 years and older: two self-controlled studies in the U.S. Vaccine. Jul 19, 2023;41(32):4666-4678. [ FREE Full text ] [ CrossRef ] [ Medline ]
- Hobbi S, Saunders-Hastings P, Zhou CK, Beers J, Srikrishnan A, Hettinger A, et al. Development and validation of an algorithm for thrombosis with thrombocytopenia syndrome (TTS) at unusual sites. Int J Gen Med. Jun 15, 2023;16:2461-2467. [ FREE Full text ] [ CrossRef ] [ Medline ]
- Kostka K. Chapter 10: defining cohorts. The Book of OHDSI. 2021. URL: https://ohdsi.github.io/TheBookOfOhdsi/Cohorts.html [accessed 2024-02-22]
- Denaxas S, Gonzalez-Izquierdo A, Direk K, Fitzpatrick NK, Fatemifar G, Banerjee A, et al. UK phenomics platform for developing and validating electronic health record phenotypes: CALIBER. J Am Med Inform Assoc. Dec 01, 2019;26(12):1545-1559. [ FREE Full text ] [ CrossRef ] [ Medline ]
- Kawai AT, Martin D, Henrickson SE, Goff A, Reidy M, Santiago D, et al. Validation of febrile seizures identified in the sentinel post-licensure rapid immunization safety monitoring program. Vaccine. Jul 09, 2019;37(30):4172-4176. [ CrossRef ] [ Medline ]
- Walsh KE, Cutrona SL, Foy S, Baker MA, Forrow S, Shoaibi A, et al. Validation of anaphylaxis in the food and drug administration's mini-sentinel. Pharmacoepidemiol Drug Saf. Nov 2013;22(11):1205-1213. [ FREE Full text ] [ CrossRef ] [ Medline ]
- Bann MA, Carrell DS, Gruber S, Shinde M, Ball R, Nelson JC, et al. Identification and validation of anaphylaxis using electronic health data in a population-based setting. Epidemiology. May 01, 2021;32(3):439-443. [ CrossRef ] [ Medline ]
- Funch D, Holick C, Velentgas P, Clifford R, Wahl PM, McMahill-Walraven C, et al. Algorithms for identification of Guillain-Barré syndrome among adolescents in claims databases. Vaccine. Apr 12, 2013;31(16):2075-2079. [ CrossRef ] [ Medline ]
- Schuemie M, DeFalco F. 8.3 ATLAS. The Book of OHDSI. 2021. URL: https://ohdsi.github.io/TheBookOfOhdsi/OhdsiAnalyticsTools.html#atlas [accessed 2024-02-22]
- Value set authority center. National Institute for Health National Library of Medicine. URL: https://vsac.nlm.nih.gov/ [accessed 2024-02-22]
- Whitaker B, Pizarro J, Deady M, Williams A, Ezzeldin H, Belov A, et al. Detection of allergic transfusion-related adverse events from electronic medical records. Transfusion. Oct 2022;62(10):2029-2038. [ CrossRef ] [ Medline ]
- Deady M, Ezzeldin H, Cook K, Billings D, Pizarro J, Plotogea AA, et al. The food and drug administration biologics effectiveness and safety initiative facilitates detection of vaccine administrations from unstructured data in medical records through natural language processing. Front Digit Health. Dec 22, 2021;3:777905. [ FREE Full text ] [ CrossRef ] [ Medline ]
- Standardized data: the OMOP common data model. Observational Health Data Sciences and Informatics. URL: https://www.ohdsi.org/data-standardization/ [accessed 2024-02-22]
- Agresti A, Coull BA. Approximate is better than “exact” for interval estimation of binomial proportions. Am Stat. 1998;52(2):119-126. [ CrossRef ]
- Brown LD, Cai TT, DasGupta A. Interval estimation for a binomial proportion. Statist Sci. May 2001;16(2):101-133. [ CrossRef ]
- McHugh ML. Interrater reliability: the kappa statistic. Biochem Med (Zagreb). 2012;22(3):276-282. [ FREE Full text ] [ Medline ]
- Pandey CK, Bose N, Garg G, Singh N, Baronia A, Agarwal A, et al. Gabapentin for the treatment of pain in guillain-barré syndrome: a double-blinded, placebo-controlled, crossover study. Anesth Analg. Dec 2002;95(6):1719-23, table of contents. [ CrossRef ] [ Medline ]
- Idowu RT, Carnahan R, Sathe NA, McPheeters ML. A systematic review of validated methods to capture myopericarditis using administrative or claims data. Vaccine. Dec 30, 2013;31 Suppl 10:K34-K40. [ CrossRef ] [ Medline ]
- CQL - clinical quality language. eCQI Resource Center. URL: https://ecqi.healthit.gov/cql?qt-tabs_cql=0 [accessed 2024-02-22]
- Ong MS, Klann JG, Lin KJ, Maron BA, Murphy SN, Natter MD, et al. Claims‐based algorithms for identifying patients with pulmonary hypertension: a comparison of decision rules and machine‐learning approaches. J Am Heart Assoc. Sep 29, 2020;9(19):e016648. [ CrossRef ]
- Dang TK, Lan X, Weng J, Feng M. Federated learning for electronic health records. ACM Trans Intell Syst Technol. Jun 21, 2022;13(5):1-17. [ CrossRef ]
- Manual for the surveillance of vaccine-preventable diseases. Centers for Disease Control and Prevention. URL: https://www.cdc.gov/vaccines/pubs/surv-manual/chpt21-surv-adverse-events.html [accessed 2024-02-22]
Abbreviations
| adverse event |
| adverse event of special interest |
| application programming interface |
| Biologics Effectiveness and Safety Initiative |
| Center for Biologics Evaluation and Research |
| Centers for Disease Control and Prevention |
| cerebrospinal fluid |
| electronic health record |
| emergency use authorization |
| Food and Drug Administration |
| Fast Healthcare Interoperable Resources |
| Guillain-Barré syndrome |
| Health Level 7 application programming interface |
| International Classification of Diseases, 10th Revision, clinical modification |
| Observational Health Data Sciences and Informatics |
| positive predictive value |
| real-world data |
| thrombosis with thrombocytopenia syndrome |
| United States Core Data for Interoperability |
| Vaccine Adverse Event Reporting System |
| white blood cell |
Edited by A Mavragani, T Sanchez; submitted 09.06.23; peer-reviewed by B Ru, AS Bhagavathula; comments to author 20.01.24; revised version received 24.02.24; accepted 26.05.24; published 15.07.24.
©Ashley A Holdefer, Jeno Pizarro, Patrick Saunders-Hastings, Jeffrey Beers, Arianna Sang, Aaron Zachary Hettinger, Joseph Blumenthal, Erik Martinez, Lance Daniel Jones, Matthew Deady, Hussein Ezzeldin, Steven A Anderson. Originally published in JMIR Public Health and Surveillance (https://publichealth.jmir.org), 15.07.2024.
This is an open-access article distributed under the terms of the Creative Commons Attribution License (https://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work, first published in JMIR Public Health and Surveillance, is properly cited. The complete bibliographic information, a link to the original publication on https://publichealth.jmir.org, as well as this copyright and license information must be included.
This paper is in the following e-collection/theme issue:
Published on 27.6.2024 in Vol 12 (2024)
Data Flow Construction and Quality Evaluation of Electronic Source Data in Clinical Trials: Pilot Study Based on Hospital Electronic Medical Records in China
Authors of this article:

- Yannan Yuan 1 , MS ;
- Yun Mei 2 , MS ;
- Shuhua Zhao 1 , MS ;
- Shenglong Dai 3 , MS ;
- Xiaohong Liu 1 , MS ;
- Xiaojing Sun 3 , MA ;
- Zhiying Fu 1 , MS ;
- Liheng Zhou 3 , MS ;
- Jie Ai 2 , MS ;
- Liheng Ma 3 , MD ;
- Min Jiang 4 , MS
1 Key Laboratory of Carcinogenesis and Translational Research (Ministry of Education/Beijing), National Drug Clinical Trial Center, Peking University Cancer Hospital & Institute, , Beijing, , China
2 Yidu Tech Inc, , Beijing, , China
3 Pfizer (China) Research & Development Co, , Shanghai, , China
4 State Key Laboratory of Holistic Integrative Management of Gastrointestinal Cancers, Beijing Key Laboratory of Carcinogenesis and Translational Research, National Drug Clinical Trial Center, Peking University Cancer Hospital & Institute, , Beijing, , China
Corresponding Author:
Min Jiang, MS
Background: The traditional clinical trial data collection process requires a clinical research coordinator who is authorized by the investigators to read from the hospital’s electronic medical record. Using electronic source data opens a new path to extract patients’ data from electronic health records (EHRs) and transfer them directly to an electronic data capture (EDC) system; this method is often referred to as eSource. eSource technology in a clinical trial data flow can improve data quality without compromising timeliness. At the same time, improved data collection efficiency reduces clinical trial costs.
Objective: This study aims to explore how to extract clinical trial–related data from hospital EHR systems, transform the data into a format required by the EDC system, and transfer it into sponsors’ environments, and to evaluate the transferred data sets to validate the availability, completeness, and accuracy of building an eSource dataflow.
Methods: A prospective clinical trial study registered on the Drug Clinical Trial Registration and Information Disclosure Platform was selected, and the following data modules were extracted from the structured data of 4 case report forms: demographics, vital signs, local laboratory data, and concomitant medications. The extracted data was mapped and transformed, deidentified, and transferred to the sponsor’s environment. Data validation was performed based on availability, completeness, and accuracy.
Results: In a secure and controlled data environment, clinical trial data was successfully transferred from a hospital EHR to the sponsor’s environment with 100% transcriptional accuracy, but the availability and completeness of the data could be improved.
Conclusions: Data availability was low due to some required fields in the EDC system not being available directly in the EHR. Some data is also still in an unstructured or paper-based format. The top-level design of the eSource technology and the construction of hospital electronic data standards should help lay a foundation for a full electronic data flow from EHRs to EDC systems in the future.
Introduction
Source data are the original records from clinical trials or all information recorded on certified copies, including clinical findings, observations, and records of other relevant activities necessary for the reconstruction and evaluation of the trial [ 1 ]. Electronic source data are data initially recorded in an electronic format (electronic source data or eSource) [ 2 , 3 ].
The traditional clinical trial data collection process requires a clinical research coordinator (CRC) who is authorized by the investigators to read from the hospital’s electronic medical record and other clinical trial–related data from the hospital information system and then manually enter the patient’s data into the electronic data capture (EDC) system. After data entry, the clinical research associate visits the site to perform source data verification and source data review. The drawbacks of collecting data by manual transcription are that data quality and timeliness cannot be guaranteed and that it is a waste of human and material resources. Using electronic source data opens a new path to extract patients’ data from electronic health records (EHRs) and transfer it directly to EDC systems (often the method is referred to as eSource) [ 4 ]. eSource technology in a clinical trial data flow can improve data quality without compromising timeliness [ 5 ]. At the same time, improved data collection efficiency reduces clinical trial costs [ 6 ].
eSource can be divided into two levels. The first level is to enable the hospital information system to obtain complete data sets; the second level is to allow direct data transfer to EDC systems based on the clinical trial patients’ electronic data in hospitals to avoid the electronic data being transcribed manually again, which is the core purpose of eSource [ 7 ]. This project will explore the use of eSource technology to extract clinical trial data from EHRs, send it to the sponsor data environment, and discuss the issues and challenges occurring in its application process.
Ethics Approval
This study was approved by the Ethics Committee and Human Genetic Resource Administration of China (2020YW135). During the ethical review process, the most significant challenges were patients’ informed consent, privacy protection, and data security. The B7461024 Informed Consent Form (Version 4) states that “ interested parties may use subjects’ personal information to improve the quality, design, and safety of this and other studies,” and “Is my personal information likely to be used in other studies? Your coded information may be used to advance scientific research and public health in other projects conducted in future.” This project is an exploration of using electronic source data technology instead of traditional manual transcription in the process of transferring data from hospital EHRs to EDC systems, which will improve the data quality of clinical trials and will improve the data flow in the future. Therefore, this project is within the scope of the informed consent form for study B7461024, which was approved by the ethics committee after clarification.
Project Information
This project was conducted from December 15, 2020, to November 19, 2021, which was before China’s personal information protection law and data security law were introduced. The data for this project were obtained from an ongoing phase 2, multicenter, open-label, dual-cohort study to evaluate the efficacy and safety of Lorlatinib (pf-06463922) monotherapy in anaplastic lymphoma kinase (ALK) inhibitor–treated locally advanced or metastatic ALK-positive non–small cell lung cancer patients in China (B7461024), registered by the sponsor on the Drug Clinical Trials Registration and Disclosure Platform (CTR20181867). The data extraction involved 4 case report form (CRF) data modules: demographics, concomitant medication, local lab, and vital signs, which were collected in the following ways:
- Demographics: Originally entered directly into the hospital EHR then manually transcribed by the CRC to the sponsor’s EDC system
- Local lab: Laboratory data collected by the hospital laboratory information management system (LIMS) and then manually transcribed by the CRC into the EDC system
- Vital signs: Hospital uses paper-based tracking form provided by the sponsor to record patients’ vital signs and investigators transcribe the vital signs data into the hospital medical record
- Concomitant medication: Similar to vital signs, hospital uses the paper tracking form provided by the sponsor to record the adverse reactions and concomitant medication; investigator might also transfer the concomitant medication data into the hospital EHR, but there was no mandatory requirement to transfer these data into patients’ medical records
All information was collected from 6 patients in a total of 29 fields ( Textbox 1 ).
Demographics
- Date of birth
Concomitant medication
- Combined drug name
- Whether for the treatment of adverse reactions
- Adverse event number
- Combined drug start date
- Combined drug end date
- Currently still in use
Vital signs
- Date of vital signs collection
- Weight unit
- Body temperature
- Height unit
- Location of temperature measurement
- Systolic blood pressure
- Diastolic blood pressure
- Laboratory inspection name
- Laboratory name and address
- Sponsor number
- Laboratory number
- Incomplete laboratory inspection
- Sample collection data
- Inspection results
Data Process Workflow
The study chosen in our project used the traditional manual data entry method to transcribe patients’ CRF data into the EDC system. This project proposes testing the acquisition of data directly from the hospital EHR, deidentification of the patients’ electronic data on the hospital medical data intelligence platform, mapping and transforming the data based on the sponsor’s EDC data standard, and transferring the data into the sponsor’s environment. The data was transferred from the hospital to the sponsor’s data environment and compared to data that was captured by traditional manual entry methods to verify the availability, completeness, and accuracy of the eSource technology.
In the network environment of this project, the technology provider accessed the hospital network through a virtual private network (VPN) and a bastion host, and processed the data of this project as a private cloud, thus ensuring the security of the hospital data.
Data Integration
The hospital information system involved in this project has reached the national standards of “Level 3 Equivalence,” “Electronic Medical Record Level 5,” and “Interoperability Level 4.” The medical data intelligence platform in this project is deployed in a hospital intranet, isolated from external networks. Integrated data from different information systems, including the hospital information system, LIMS, picture archiving and communication system, etc, were deidentified from the platform and transferred to a third-party private cloud platform for translation and data format conversion after authorization by the hospital through a VPN.
The scope of data collection in this project was limited to patients who signed Informed Consent Form (Version 4) for study B7461024. The structured data of four CRF data modules (demographic, concomitant medications, local lab, and vital signs) were extracted from the source data in hospital systems, and data processing was completed.
Three-Layer Deidentification of Data
In this project, three layers of deidentification were performed on the electronic source data to ensure data security. The first layer of deidentification was performed before the certified copy of data was loaded to the hospital’s medical data intelligence platform. The second layer of deidentification follows the Health Insurance Portability and Accountability Act (HIPAA) by deidentifying 18 data fields at the system level. A third layer of deidentification was performed when mapping and transforming third-party databases for the clinical trial data (demographics, concomitant medications, laboratory tests, and vital signs) collected for this study, as required by the project design.
Collected data did not contain any sensitive information with personal identifiers of the patients, and all deidentification processes were conducted in the internal environment of the hospital. In addition to complying with the relevant laws and regulations, we followed the requirements of Good Clinical Practice regarding patient privacy and confidentiality, and further complied with the requirements of HIPAA to deidentify the 18 basic data fields. Data fields outside the scope of HIPAA will be deidentified and processed in accordance with the TransCelerate guidelines published in April 2015 to ensure the security of patients’ personal information and to eliminate the possibility of patient information leakage [ 8 ].
The general rules for the third layer of deidentification were as follows:
- Time field: A specific time point is used as the base time, and the encrypted time value is the difference between the word time and the base time
- ID field: Categorized according to the value and only shows the category
- Age field: Categorized according to the value and only shows the category
- Low-frequency field: set to null
In addition, all data flows keep audit trails throughout and are available for audit.
Data Normalization and Information Extraction
After three layers of deidentification, the data was transferred from a hospital to a third-party private cloud platform through a VPN, where translation from Chinese to English and data format conversion were implemented. The whole transfer process was performed for the data that was collected for the clinical trial of this study. Standardization of data is a crucial task during the data preparation phase. This process involves consolidating data from different systems and structures into a consistent, comprehensible, and operable format. First, a thorough examination of data from various systems is necessary. Understanding the data structure, format, and meaning of each system is essential. The second step involves establishing a data dictionary that clearly outlines the meaning, format, and possible values of each data element. Next, selecting a data standard is necessary to ensure consistency and comparability. In this study, we adopted the Health Level 7 (HL7) standard. Additionally, data cleansing and transformation are needed to meet standard requirements, including handling missing data, resolving mismatched data formats, or performing data type conversions. Extract, transform, and load tools were used to integrate data from different systems. Data security must be ensured throughout the data integration process. This includes encrypting sensitive information and strictly managing data permissions. Data verification and validation steps were then performed by professional staff on the translated data. The data from the hospital’s medical data intelligence platform were then converted from JSON format to XML and Excel formats. The processed data was transferred back to the hospital via a VPN to a designated location for final adjudication before loading to the sponsor’s environment.
One-Time Data Push and Quality Assessment
After the hospital received the processed data, it was then pushed by the hospital to the sponsor’s secure and controlled environment ( Figure 1 ). All data deidentification processes were conducted in the hospital’s environment, and none of the data obtained by the sponsor can be traced back to patients’ personal information to ensure their privacy and information security.
The data quality of this project was assessed using industry data quality assessment rules [ 9 ], which are shown in Table 1 .

| Data validation methods | Dimension | Method description | Cases |
| Data availability verification | Field dimension | The ratio of the total number of data fields in the clinical trial CRF available in the hospital EHR to the total number of data fields required in the electronic CRF: EHR /CRF × 100% | Based on the electronic CRF, 6 data fields in the demography need to be captured, and 3 of them have records in the EHR. Data availability: 3/6 × 100% = 50% |
| Data availability verification | Field dimension | The ratio of the total number of data fields in the clinical trial CRF (eSource) that can be transmitted electronically in the hospital’s EHR to the total number of data fields required in the electronic CRF: eSource /CRF × 100% | Based on the electronic CRF, 6 data fields in the demography need to be captured, and 2 data fields can be captured by the eSource method. Data availability: 2/6 × 100% = 33.33% |
| Data completeness verification | Numerical dimension | The ratio of the total number of nonnull data (eSourceV) captured (processed and sent to the sponsor) via the eSource method to the total number of data fields requested on the electronic CRF: eSourceV /CRF × 100% | Based on the clinical trial design, 38 concomitant medication pages need to be collected: 7 pages were collected via eSource and 2 fields was entered per page. Data completeness: 7 × 2/(2 × 38) × 100% = 18.42% |
| Data accuracy verification | Numerical dimension | Matching of data field values in the hospital’s EHR with data field values that can be captured by eSource (data fields that are processed and sent to the sponsor) | 4 fields of demography were successfully transmitted through eSource, with 4 data points in each. After comparing with the data in the electronic data capture system, there were no mismatches for one data point. Data accuracy: 8/(2 × 4) × 100% = 100% |
a CRF: case report form.
b EHR: electronic health record.
c Total number of data fields in the hospital’s EHR.
d Total number of data fields requested in the electronic CRF.
e Total number of data fields captured (processed and sent to the sponsor) through the eSource method.
f Total number of nonempty data fields captured (processed and sent to the sponsor) through the eSource method.
In this project, we collected patients’ demographics, vital signs information, local laboratory data, and concomitant medication data from EHRs, successfully pushed the data directly to the designated sponsor environment, and evaluated the data quality from three perspectives including availability, completeness, and accuracy ( Table 2 ).
- The eSource-CRF availability score, which is used to evaluate the ratio of fields in EHR that can be collected by eSource and used for CRF, was low for demographics, blood tests, and urine sample tests but higher for vital signs and concomitant medications.
- Data completeness, defined as the ratio of the total number of nonnull data captured by eSource to the total number of data fields required in the electronic CRF, was used to evaluate the ratio of nonnull data fields in the CRF that can be captured by eSource. In this study, the completeness score of the vital signs module was only 1.32%, and the concomitant medications and laboratory test modules also had poor performance in the data completeness evaluation.
- Data accuracy, defined as the compatibility between the data field values in the hospital EHR and the data field values that can be collected using eSource, was 100% for all modules.
- EHR-CRF availability, which is used to evaluate the ratio of fields in the EHR that can be used for the CRF, was 50%, 60%, and 66.67% for demographics, blood tests, and urine sample tests, respectively, in this study, and the rest of the data were 100% available.
| CRF domain | CRF-EHR data availability, n/N (%) | CRF-eSource data availability, n/N (%) | Data completeness (preliminary findings), n/N (%) | Data accuracy (preliminary findings), n/N (%) | |
| Definition | Study CRF data elements available in hospital EHR | Study CRF data elements available in hospital EHR and able to be electronically transferred through eSource technology | Study CRF data elements available and entered into hospital EHR and transferred through eSource technology | Study CRF data elements available and entered into hospital EHR and transferred through eSource technology with expected result (eg, matches what was entered directly in form) | |
| Demographics | 3/6 (50.00) | 2/6 (33.33) | 12/12 (100.00) | 12/12 (100.00) | |
| Vital signs | 10/10 (100.00) | 9/10 (90.00) | 24/1812 (1.32) | 20/20 (100.00) | |
| Blood biochemical tests | 6/10 (60.00) | 5/10 (50.00) | 12,968/13,540 (95.78) | 7767/7767 (100.00) | |
| Urine sample tests | 6/9 (66.67) | 5/9 (55.56) | 15/40 (37.56) | 15/15 (100.00) | |
| Concomitant medication | 10/10 (100.00) | 9/10 (90.00) | 14/76 (18.42) | 6/6 (100.00) | |
c Checks were made with the relevant clinical research associates (CRAs) regarding the original data collection and CRF completion methods for the following reasons: vital signs were obtained using paper tracking forms provided by the sponsor as the original data source, and the data may not be transcribed into the hospital information system (HIS) by the researcher. Therefore, data from many visits are not available in the HIS.
d A total of 2708 blood biochemistry tests were involved.
e Concomitant medication uses tracking forms to record adverse event and ConMed (a paper source), and data may not be transcribed into the HIS. As confirmed by the CRA, the percentage of paper ConMed sources was approximately 80%.
Although EHRs have been widely used, the degree of structure of EHR data varies substantially among different data modules. In EHRs, demographics, vital signs, local lab data, and concomitant medications are more structured than patient history or progress notes and often contain unstructured text [ 10 ]. Therefore, we selected these 4 well-structured data modules for exploration in this project.
For demographics data, among the 6 required fields (subject ID, date of birth, sex, ethnicity, race, and age), subject ID (subject code number/identifier in the trial, not the patient code number/identifier in the EHR system), ethnicity, and race were not available in the EHR, so the EHR-CRF availability score was 50%. Since this was an exploratory project, the date of birth field was also deidentified and thus could not be collected based on our deidentification rule, so the eSource-CRF availability score was 33%. In the future, the availability score can reach close to 100% by bidirectional design of the EHR and CRF under the premise of obtaining compliance for industrial-level applications.
The low availability score of local laboratory data on EHR-CRFs is due to the lack of required fields in the hospital system; “Lab ID” and “Not Done” do not exist in the LIMS, and for the “Clinically Significant” field, the meaning of laboratory test results needs to be manually interpreted by an investigator, so they cannot be transcribed directly. The availability score of eSource-CRFs was further decreased because the field “Laboratory Name and Address” is not an independent structured field in the EHR. The completeness score of urine sample test data was only 37.56% because during the actual clinical trial, especially amid the COVID-19 pandemic period, patients completed study-related laboratory tests at other sites, and those test results were collected via paper-based reports, so the complete data sets cannot be extracted from the site’s system.
To improve data availability in future applications, clinical trial–specific fields need to be added to EHR designs for those data that require an investigator’s interpretation such as “Clinically Significant,” and data transfer and mapping processes for the determination of the scope of data collection also needs to be optimized. Based on these two conditions, the completeness score can be improved to over 90%.
The availability and accuracy of vital signs data are ideal. However, since not all vital signs data collection was recorded by the electronic system during the actual study visit, many vital signs data were collected in “patient diary” and other types of paper-based documents during the study, resulting in a serious limitation in data completeness. With the development of more clinical trial–related electronic hardware and enhancements in products intelligence, more vital signs data will be directly collected by electronic systems, and the completeness of vital signs data transferred from EHR to EDC will be greatly improved in the future.
In the concomitant medication module, there was a good score for availability and accuracy because the standardization and structuring of prescriptions are well done in this hospital system. However, the patient’s medication use period during hospitalization is recorded in unstructured text, so the data could not be captured for this study, resulting in a low completeness score of 18.42% for concomitant medication.
In summary, the accuracy score of eSource data in this study was high (100% for all fields). A study by Memorial Sloan Kettering Cancer Center and Yale University confirmed that the error rate of automatic transcription reduced from 6.7% to 0% compared to manual transcription [ 10 ]. However, data availability and completeness have not reached a good level. Data availability varies widely across studies, ranging from 13.4% in the Retrieving EHR Useful Data for Secondary Exploitation (REUSE) project [ 11 ] to 75% in The STARBRITE Proof-of-Concept Study [ 12 ], mainly related to the coverage and structure of the EHR.
National drug regulatory agencies (eg, US Food and Drug Administration [FDA], European Medicines Agency, Medicines and Healthcare products Regulatory Agency, and Pharmaceuticals and Medical Devices Agency) have developed guidelines to support the application of eSource to clinical trials [ 3 , 13 - 15 ]. The new Good Clinical Practice issued by the Center for Drug Evaluation in 2020 encourages investigators to use clinical trials’ electronic medical records for source data documentation [ 1 ]. Despite this, we still encountered challenges, including ethical review and data security, during this study’s implementation process. Without knowing the precedents, the project team decided to follow the requirements for clinical trials to control the quality of the study. There were no existing regulatory policies or national guidance on eSource in China at the time of this study. The project team provided explanations for inapplicable documents and communicated several times to ensure the approval of relevant institutional departments before finally becoming the first eSource technology study to be approved by the Ethics Committee and Human Genetic Resource Administration of China.
In the absence of regulatory guidelines, our eSource study, the first in China’s International Multi-center Clinical Trial, navigated challenges in data deidentification. We adopted HIPAA and TransCelerate’s guidelines [ 8 ]. Securing approval under “China International Cooperative Scientific Research Approval for Human Genetic Resources,” we answered queries and achieved unprecedented recognition. For transferring data from the hospital to the sponsor’s environment, we prioritized security and obtained necessary approvals. Iterative revisions ensured a robust data flow design. Challenges in mapping hospital EHR to EDC standards highlighted the need for a scalable mechanism. This study pioneers eSource tech integration in China, emphasizing the importance of seamless data mapping. In the process of executing data standardization, several challenges may arise, including inconsistent data definitions. Data from different systems may use different definitions due to the independent development of these systems, leading to varied interpretations of even identical concepts. To address this issue, establishing a unified data dictionary is crucial to ensure consensus on the definition of each data element. Different systems might also use distinct data formats such as text encodings. Preintegration format conversion is required, and extract, transform, and load tools or scripts can assist in standardizing these formats. During the integration of data from multiple systems, it is possible to discover data in one system that is not present in another. In the data standardization process, considerations must be made on how to handle missing data, which may involve interpolation, setting default values, etc. Quality issues like errors, duplicates, or inaccuracies may exist in data from different systems. Data cleansing, involving deduplication, error correction, logical validation, etc, is necessary to address these quality issues. Different systems may generate data based on diverse business rules and hospital use scenarios. In data standardization, unifying these rules requires collaboration with domain experts to ensure consistency.
Internationally, multiple research studies and publications have been released on regulations, guidelines, and validation of eSource. The FDA provided guidance on the use of electronic source data in clinical trials in 2013 that aims to address barriers to capturing electronic source data for clinical trials, including the lack of interoperability between EHRs and EDC systems. The European-wide Electronic Health Records for Clinical Research (EHR4CR) project was launched in 2011 to explore technical options for the direct capture of EHR data within 35 institutions, and the project was completed in 2016 [ 16 ]. The second phase of the project connected the EHRs to EDC systems [ 17 ] and aimed to realize the interoperability of EHRs and EDC systems. The US experience focuses more on improving and standardizing the existing EHRs to make them more uniform.
In Europe, the experience focuses on breaking down the technical barrier of interoperability between EHRs and EDC systems. In China, the current industry trends focus on the governance of existing EHR data in the hospital and the building of clinical data repository platforms [ 7 ]. Clinical data repository platforms focus on data integration and cleaning between EHRs and other systems in hospital environments and on unstructured data normalization and standardization by natural language processing and other AI technology [ 18 ]. At the national level, China is also actively promoting the digitization of medical big data and is committed to the formation of regional health care databases [ 19 ], which lays the foundation for the future implementation of eSource in China [ 20 ].
This study evaluates the practical application value of eSource in terms of availability, completeness, and accuracy. To improve availability, the structure of the CRF needs to be designed according to the information of the EHR data at the design stage of clinical trials. Even so, since EHRs are designed for the physicians to conduct daily health care activities, certain fields in clinical trials (eg, judgment of normal or abnormal values of laboratory tests and judgment of correlations of adverse events and combined medications) are still not available, and clinical trial–specific fields need to be added to EHR designs for those data that require investigators’ interpretation to improve data availability. Completeness could be improved by the development of hospital digitalization that ensures patients’ data is collected electronically rather than on paper. Additionally, 2708 blood test records were successfully collected from only 6 patients via eSource in this study, which indicates that laboratory tests often contain large amounts of highly structured data that are suitable for eSource. EHR-EDC end-to-end automatic data extraction by eSource is suitable for laboratory examinations and can improve the efficiency and accuracy of data extraction significantly as well as reduce redundant manual transcriptions and labor costs. Processing unstructured or even paper-based data in eSource is still a big challenge. Using machine learning tools (eg, natural language processing tools) for autostructuring can be explored in the future. The goal is to have common data standards and better top-level design to facilitate data integrity, interoperability, data security, and patient privacy protection in eSource applications. During deidentification, we processed certain data with a specific logic to protect privacy. The accuracy assessment was performed during the deidentification step to ensure that the data was still sufficiently accurate while meeting privacy requirements. Reversible methods need to be used when performing deidentification as well as providing controlled access mechanisms to the data so that the raw data can be accessed when needed. It is worth noting that different regions and industries may have different privacy regulations and compliance requirements. When deidentifying, you need to ensure that you are compliant with the relevant regulations and understand the limitations of data use. This may require working closely with a legal team.
In the future, we can consider adding performance analysis, including an assessment of data import performance. This involves evaluating the speed and efficiency of data import to ensure it is completed within a reasonable timeframe. Additionally, analyzing data query performance is crucial in practical applications to ensure that the imported data meets the expected query performance in the application. For long-term applications involving a larger size of patients, it is advisable to consider adding analyses related to maintainability and cost-effectiveness. This includes implementing detailed logging and monitoring mechanisms to promptly identify and address potential issues. Furthermore, for the imported data, establishing a version control mechanism is essential for tracing and tracking changes in the data. Simultaneously, for overall resource use, evaluating the resources required during the data import process ensures completion within a cost-effective framework. It is also important to consider the value of imported data for clinical trial operations and related decision-making, providing a comparative analysis between cost and value.
Acknowledgments
This research was supported by the Capital's Funds for Health Improvement and Research (grant No. CFH2022-2Z-2153), and the Beijing Municipal Science & Technology Commission (grant No. Z211100003521008).
Conflicts of Interest
None declared.
- Good clinical practice for drug clinical trial (GCP) 2020[EB/OL]. National Medical Products Administration. Apr 26, 2020. URL: https://www.nmpa.gov.cn/xxgk/fgwj/xzhgfxwj/20200426162401243.html [Accessed 2024-06-07]
- Sheng Q, Wang B, Chen J, et al. Classification and application of electronic source data in clinical trials [Article in Chinese]. Chin Food Drug Admin Magazine. 2021;3:36-43. URL: https://cnki.net/KCMS/detail/detail.aspx?dbcode=CJFD&dbname=CJFDLAST2021&filename=YPJD202103004&uniplatform=OVERSEA&v=TYNESF_EvmJSkJJzsO1qJ78umjfdRnU-44Oah6CO-x2-bUH1PAzstPQztpONnrV5 [Accessed 2024-06-13]
- Guidance for industry: electronic source data in clinical investigations. Food and Drug Administration. Sep 2013. URL: https://www.fda.gov/media/85183/download [Accessed 2024-06-07]
- Safran C, Bloomrosen M, Hammond WE, et al. Toward a national framework for the secondary use of health data: an American Medical Informatics Association white paper. J Am Med Inform Assoc. 2007;14(1):1-9. [ CrossRef ] [ Medline ]
- Garza M, Myneni S, Fenton SH, Zozus MN. eSource for standardized health information exchange in clinical research: a systematic review of progress in the last year. J Soc Clin Data Manag. 2021;1(2). [ CrossRef ]
- Beresniak A, Schmidt A, Proeve J, et al. Cost-benefit assessment of using electronic health records data for clinical research versus current practices: contribution of the electronic health records for clinical research (EHR4CR) European project. Contemp Clin Trials. Jan 2016;46:85-91. [ CrossRef ] [ Medline ]
- Dong C, Yao C, Gao S, et al. Strengthening clinical research source data management in hospitals to promote data quality of clinical research in China. Chin J Evid Based Med. 2019;19:11-1261. URL: https://cnki.net/KCMS/detail/detail.aspx?dbcode=CJFD&dbname=CJFDLAST2019&filename=ZZXZ201911002&uniplatform=OVERSEA&v=sYN9IctSf5JwG0pw2UoXAuBVRb9i4a9eqPVo2yeMDC4RQl7-JB3UGvM9tMvP2h-7 [Accessed 2024-06-13]
- Data de-identification and anonymization of individual patient data in clinical studies: a model approach. TransCelerate BioPharma. 2015. URL: http://www.transceleratebiopharmainc.com/wp-content/uploads/2015/04/TransCelerate-Data-De-identification-and-Anonymization-of-Individual-Patient-Data-in-Clinical-Studies-1.pdf [Accessed 2024-06-13]
- Nordo A, Eisenstein EL, Garza M, Hammond WE, Zozus MN. Evaluative outcomes in direct extraction and use of EHR data in clinical trials. Stud Health Technol Inform. 2019;257:333-340. [ Medline ]
- Vattikola A, Dai H, Buckley M, Maniar R. Direct data extraction and exchange of local LABS for clinical research protocols: a partnership with sites, biopharmaceutical firms, and clinical research organizations. J Soc Clin Data Manag. Mar 2021;1(1). [ CrossRef ]
- El Fadly A, Rance B, Lucas N, et al. Integrating clinical research with the healthcare enterprise: from the RE-USE project to the EHR4CR platform. J Biomed Inform. Dec 2011;44 Suppl 1:S94-S102. [ CrossRef ] [ Medline ]
- Kush R, Alschuler L, Ruggeri R, et al. Implementing single source: the STARBRITE proof-of-concept study. J Am Med Inform Assoc. 2007;14(5):662-673. [ CrossRef ] [ Medline ]
- Reflection paper on expectations for electronic source data and data transcribed to electronic data collection tools in clinical trials. European Medicines Agency. Jun 9, 2010. URL: https://www.ema.europa.eu/en/documents/regulatory-procedural-guideline/reflection-paper-expectations-electronic-source-data-data-transcribed-electronic-data-collection_en.pdf [Accessed 2024-06-07]
- Technical conformance guide on electronic study data submissions. Pharmaceuticals and Medical Devices Agency. Apr 27, 2015. URL: https://www.pmda.go.jp/files/000206449.pdf [Accessed 2024-06-07]
- MHRA position statement and guidance: electronic health records. MHRA. Sep 16, 2015. URL: https://assets.publishing.service.gov.uk/government/uploads/system/uploads/attachment_data/file/470228/Electronic_Health_Records_MHRA_Position_Statement.pdf [Accessed 2024-06-07]
- McCowan C, Thomson E, Szmigielski CA, et al. Using electronic health records to support clinical trials: a report on stakeholder engagement for EHR4CR. Biomed Res Int. Oct 2015;2015:707891. [ CrossRef ] [ Medline ]
- Sundgren M, Ammour N, Hydes D, Kalra D, Yeatman R. Innovations in data capture transforming trial delivery. Appl Clin Trials. Aug 12, 2021;30(7/8). URL: https://www.appliedclinicaltrialsonline.com/view/innovations-in-data-capture-transforming-trial-delivery [Accessed 2024-06-17]
- Wang Q, Yingping Y. Governance and application of big data in clinical healthcare. J Med Inform. 2018;39:2-6. [ CrossRef ]
- Guidance from the general office of the state council on promoting and regulating the development of health care big data applications 2016[EB/OL]. Gov.CN. 2016. URL: https://www.gov.cn/gongbao/content/2016/content_5088769.htm [Accessed 2024-06-07]
- Wang B, Lai J, Liao X, Jin F, Yao C. Challenges and solutions in implementing eSource technology for real-world studies in China: qualitative study among different stakeholders. JMIR Form Res. Aug 10, 2023;7:e48363. [ CrossRef ] [ Medline ]
Abbreviations
| anaplastic lymphoma kinase |
| clinical research coordinator |
| case report form |
| electronic data capture |
| electronic health record |
| Electronic Health Records for Clinical Research |
| Food and Drug Administration |
| Health Insurance Portability and Accountability Act |
| Health Level 7 |
| laboratory information management system |
| Retrieving EHR Useful Data for Secondary Exploitation |
| virtual private network |
Edited by Christian Lovis; submitted 19.09.23; peer-reviewed by Hareesh Veldandi, Yujie Su; final revised version received 20.12.23; accepted 18.04.24; published 27.06.24.
© Yannan Yuan, Yun Mei, Shuhua Zhao, Shenglong Dai, Xiaohong Liu, Xiaojing Sun, Zhiying Fu, Liheng Zhou, Jie Ai, Liheng Ma, Min Jiang. Originally published in JMIR Medical Informatics (https://medinform.jmir.org), 27.6.2024.
This is an open-access article distributed under the terms of the Creative Commons Attribution License ( https://creativecommons.org/licenses/by/4.0/ ), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work, first published in JMIR Medical Informatics, is properly cited. The complete bibliographic information, a link to the original publication on https://medinform.jmir.org/ , as well as this copyright and license information must be included.
A Case Study on Impact of Electronic Health Records System (EHRS) on Healthcare Quality at Asamankese Government Hospital
- September 2022
- This person is not on ResearchGate, or hasn't claimed this research yet.
Abstract and Figures

Discover the world's research
- 25+ million members
- 160+ million publication pages
- 2.3+ billion citations
- Kennedy Addo
- Pabbi Kwaku Agyepong

- Ronald Nakaka
- HEALTH INF MANAG J

- Razzaq Adetona Adio

- Open Transport J

- Phil Johnson
- Murray Clark
- Sara L. Gill

- Robert B. McDaniel

- James M Hoffman
- Recruit researchers
- Join for free
- Login Email Tip: Most researchers use their institutional email address as their ResearchGate login Password Forgot password? Keep me logged in Log in or Continue with Google Welcome back! Please log in. Email · Hint Tip: Most researchers use their institutional email address as their ResearchGate login Password Forgot password? Keep me logged in Log in or Continue with Google No account? Sign up
- Research article
- Open access
- Published: 10 July 2024
Consistency, completeness and external validity of ethnicity recording in NHS primary care records: a cohort study in 25 million patients’ records at source using OpenSAFELY
- The OpenSAFELY Collaborative ,
- Colm D. Andrews 1 ,
- Rohini Mathur 2 , 4 ,
- Jon Massey 1 ,
- Robin Park 1 ,
- Helen J. Curtis 1 ,
- Lisa Hopcroft 1 ,
- Amir Mehrkar 1 ,
- Seb Bacon 1 ,
- George Hickman 1 ,
- Rebecca Smith 1 ,
- David Evans 1 ,
- Tom Ward 1 ,
- Simon Davy 1 ,
- Peter Inglesby 1 ,
- Iain Dillingham 1 ,
- Steven Maude 1 ,
- Thomas O’Dwyer 1 ,
- Ben F. C. Butler-Cole 1 ,
- Lucy Bridges 1 ,
- Chris Bates 3 ,
- John Parry 3 ,
- Frank Hester 3 ,
- Sam Harper 3 ,
- Jonathan Cockburn 3 ,
- Ben Goldacre 1 ,
- Brian MacKenna 1 ,
- Laurie A. Tomlinson 2 ,
- Alex J. Walker 1 &
- William J. Hulme 1
BMC Medicine volume 22 , Article number: 288 ( 2024 ) Cite this article
199 Accesses
Metrics details
Ethnicity is known to be an important correlate of health outcomes, particularly during the COVID-19 pandemic, where some ethnic groups were shown to be at higher risk of infection and adverse outcomes. The recording of patients’ ethnic groups in primary care can support research and efforts to achieve equity in service provision and outcomes; however, the coding of ethnicity is known to present complex challenges. We therefore set out to describe ethnicity coding in detail with a view to supporting the use of this data in a wide range of settings, as part of wider efforts to robustly describe and define methods of using administrative data.
We describe the completeness and consistency of primary care ethnicity recording in the OpenSAFELY-TPP database, containing linked primary care and hospital records in > 25 million patients in England. We also compared the ethnic breakdown in OpenSAFELY-TPP with that of the 2021 UK census.
78.2% of patients registered in OpenSAFELY-TPP on 1 January 2022 had their ethnicity recorded in primary care records, rising to 92.5% when supplemented with hospital data. The completeness of ethnicity recording was higher for women than for men. The rate of primary care ethnicity recording ranged from 77% in the South East of England to 82.2% in the West Midlands. Ethnicity recording rates were higher in patients with chronic or other serious health conditions. For each of the five broad ethnicity groups, primary care recorded ethnicity was within 2.9 percentage points of the population rate as recorded in the 2021 Census for England as a whole. For patients with multiple ethnicity records, 98.7% of the latest recorded ethnicities matched the most frequently coded ethnicity. Patients whose latest recorded ethnicity was categorised as Other were most likely to have a discordant ethnicity recording (32.2%).
Conclusions
Primary care ethnicity data in OpenSAFELY is present for over three quarters of all patients, and combined with data from other sources can achieve a high level of completeness. The overall distribution of ethnicities across all English OpenSAFELY-TPP practices was similar to the 2021 Census, with some regional variation. This report identifies the best available codelist for use in OpenSAFELY and similar electronic health record data.
Peer Review reports
Ethnicity is known to be an important determinant of health inequalities, particularly during the COVID-19 outbreak where a complex interplay of social and biological factors resulted in increased exposure, reduced protection and increased severity of illness in particular ethnic groups [ 1 , 2 ]. The UK has a diverse ethnic population (The 2021 Office for National Statistics (ONS) Census estimated 9.6% Asian, 4.2% Black, 3.0% Mixed, 81.0% White, 2.2% Other [ 3 ]), which can make health research conducted in the UK generalisable to countries. Complete and consistent recording of patients’ ethnic group in primary care can support efforts to achieve equity in service provision and reduces bias in research [ 4 , 5 ]. Ethnicity recording for new patients registering with general practice across the UK has improved following Quality and Outcomes Framework (QOF) financial incentivisation between 2006/07 and 2011/12 [ 6 , 7 ]. As a result, ethnicity is now being captured for the majority of the population in routine electronic healthcare records and is comparable to the general population [ 6 ]. The uptake and utilisation of healthcare services still varies across ethnic groups, and the recently established NHS Race and Health Observatory have led calls for a dedicated drive by NHS England and NHS Digital to emphasise the importance of collecting and reporting ethnicity data [ 8 ].
OpenSAFELY is a secure health analytics platform created by our team on behalf of NHS England. OpenSAFELY provides a secure software interface allowing analysis of pseudonymised primary care patient records from England in near real-time within highly secure data environments.
In primary care data, patient ethnicity is recorded via clinical codes, similar to how any other clinical condition or event is recorded. In OpenSAFELY-TPP, both Clinical Terms Version 3 (CTV3 (Read)) codes and Systematised Nomenclature of Medicine Clinical Terms (SNOMED CT) codes are used. SNOMED CT is an NHS standard, widely used across England.
Ethnicity is also recorded in secondary care, when patients attend emergency care, inpatient or outpatient services, independently of ethnicity in the primary care record. This is available via NHS England’s Secondary Uses Service (SUS) [ 9 ]. It is common practice in OpenSAFELY to supplement primary care ethnicity, where missing, with ethnicity data from SUS [ 10 , 11 ]. Throughout this paper, we refer to ethnicity rather than race as recommended by the ONS: ‘The word “race” places people into categories based on physical characteristics, whilst ethnicity is self-defined and includes aspects such as culture, heritage, religion and identity’. However, we recognise that the distinction between and use of these terms may differ in different settings.
In this paper, we study the completeness, consistency and representativeness of routinely collected ethnicity data in primary care.
Study design
Retrospective cohort study across 25 million patients registered with English general practices in OpenSAFELY-TPP.
Data sources
This study uses data from the OpenSAFELY-TPP database, covering around 40% of the English population. The database includes primary care records of patients in practices using the TPP SystmOne patient information system and is linked to other NHS data sources, including in-patient hospital records from NHS England’s Secondary Use Service (SUS), where ethnicity is also recorded independently of ethnicity in the primary care record.
All data were linked, stored and analysed securely within the OpenSAFELY platform https://opensafely.org/ . Data include pseudonymized data such as coded diagnoses, medications and physiological parameters. No free text data are included. All code is shared openly for review and re-use under MIT open licence (opensafely/ethnicity-short-data-report at notebook). Detailed pseudonymised patient data is potentially re-identifiable and therefore not shared.
Study population
Patients were included in the study if they were registered at an English general practice using TPP on 1 January 2022.
Ethnicity ascertainment
In primary care data, there is no categorical ‘ethnicity’ variable to record this information. Rather, ethnicity is recorded using clinical codes—entered by a clinician or administrator with a location and date—like any other clinical or administrative event, with specific codes relating to each ethnic group [ 12 , 13 , 14 ]. This means ethnicity can be recorded by the practice in multiple, potentially conflicting, ways over time.
We created a new codelist, SNOMED:2022 [ 13 ], by identifying relevant ethnicity SNOMED CT codes and ensuring completeness by comparing the codelist to the following: another OpenSAFELY created codelist (CTV3:2020) [ 13 ], a combined ethnicity codelist from SARS-CoV2 COVID19 Vaccination Uptake Reporting Codes published by Primary Care Information Services (PRIMIS) [ 12 , 15 ] and a codelist from General Practice Extraction Service (GPES) Data for Pandemic Planning and Research (GDPPR) [ 16 ]. Codes which relate to religion rather than ethnicity (e.g. ‘Muslim—ethnic category 2001 census’) and codes which do not specify a specific ethnicity (e.g. ‘Ethnic group not recorded’) were excluded. In total, 258 relevant ethnicity codes were identified. We then created a codelist categorisation based on the 2001 UK Census categories, which are the NHS standard for ethnicity [ 17 ], and cross referenced it against the CTV3, PRIMIS and GDPPR codelists. The ‘Gypsy or Irish Traveller’ and ‘Arab’ groups were not specifically listed in 2001 however we categorised them as `White` and `Other` respectively as per the 2011 Census grouping [ 18 ]).
The codelist categorisation consists of two ethnicity groupings based on the 2001 census (Table 1 ): all analyses used the 5-group categorisation unless otherwise stated.
If a SNOMED:2022 ethnicity code appeared in the primary care record on multiple dates, the latest entry was used unless otherwise stated.
In OpenSAFELY, the function ethnicity_from_sus combines SUS ethnicity data from admitted patient care statistics (APCS), emergency care (EC) and outpatient attendance (OPA) and selects the most frequently used ethnicity code for each patient. In hospital records from SUS, recorded ethnicity is categorised as one of the 16 categories on the 2001 UK census. This accords with the 16-level grouping described above.
We looked at the completeness of ethnicity coding in the whole population and across each of the following demographic and clinical subgroups:
Patient age was calculated as of 1 January 2022 and grouped into 5-year bands, to match the ONS age bands.
We used categories ‘male’ and ‘female’, matching the ONS recorded categories; patients with any other/unknown sex were excluded.
Deprivation
Overall deprivation was measured by the 2019 Index of Multiple Deprivation (IMD) [ 19 ] derived from the patient’s postcode at lower super output area level. IMD was divided by quintile, with 1 representing the most deprived areas and 5 representing least deprived areas. Where a patient’s postcode cannot be determined the IMD is recorded as unknown.
Region was defined as the Nomenclature of Territorial Units for Statistics (NUTS 1) region derived from the patient’s practice postcode.
As the rate of ethnicity recording would be expected to be lower in patients with fewer clinical interactions, and therefore fewer opportunities for ethnicity to be recorded, completeness was also compared in the clinical subgroups of dementia, diabetes, hypertension and learning disability which are more likely to require additional clinical interactions. Clinical subgroups were defined as the presence or absence of relevant SNOMED CT codes in the GP records for dementia [ 20 ], diabetes [ 21 ], hypertension [ 22 ] and learning disabilities [ 23 ] as of 1 January 2022.
Statistical methods
Completeness and distribution of ethnicity recording.
The proportion of patients with either (i) primary care ethnicity recorded (that is, the presence of any code in the SNOMED:2022 codelist in the patient record) or (ii) primary care ethnicity supplemented, where missing, with ethnicity data from secondary care [ 24 ] was calculated. Completeness was reported overall and within clinical and demographic subgroups.
Amongst those patients where ethnicity was recorded, the proportion of patients within each of the 5 groups was calculated, within each clinical and demographic subgroup. We also calculated the distribution of complete ethnicity recording across practices with at least 1000 registered patients.
Consistency of ethnicity recording within patients over time
Discrepancies may arise due to errors whilst entering the data or if a patient self-reports a different ethnic group from their previously recorded ethnic group. We calculated the proportion of patients with any ethnicity recorded which did not match their ‘latest’ recorded grouped ethnicity for each of the five ethnic groups.
We also calculated the proportion of patients whose latest recorded ethnicity did not match their most frequently recorded ethnicity for each of the five ethnic groups.
Consistency of ethnicity recording across data sources (primary care versus secondary care)
We calculated the proportion of patients whose latest recorded ethnicity in primary care matched their ethnicity as recorded in secondary care for each of the five ethnic groups, where both primary and secondary care are recorded.
External validation against the 2021 UK census population
The UK Census collects individual and household-level demographic data every 10 years for the whole UK population. Data on ethnicity were obtained from the 2021 UK Census for England. The most recent census across the UK was undertaken on 27 March 2021. Ethnic breakdowns for the population of England were obtained via NOMIS [ 25 ].
The ethnic breakdown of the census population was compared with our OpenSAFELY-TPP population and the relative difference was calculated using the ONS value as the baseline proportion and OpenSAFELY as the comparator. In the 2021 UK Census, the Chinese ethnic group was included in the Asian ethnic group, whereas in the 2001 census, it was included in the Other ethnic group [ 26 ]. In order to provide a suitable comparison with primary care data, we regrouped the 2021 census data as per the 2001 groups. As an additional analysis, we also compared the primary care data with the census data using the 2021 census categories.
Completeness of ethnicity data
19,618,135 of the 25,102,210 patients (78.2%) registered in OpenSAFELY-TPP on 1 January 2022 had a recorded ethnicity, rising to 92.5% when supplemented with secondary care data (Fig. 1 , Additional file 1: Table S1).
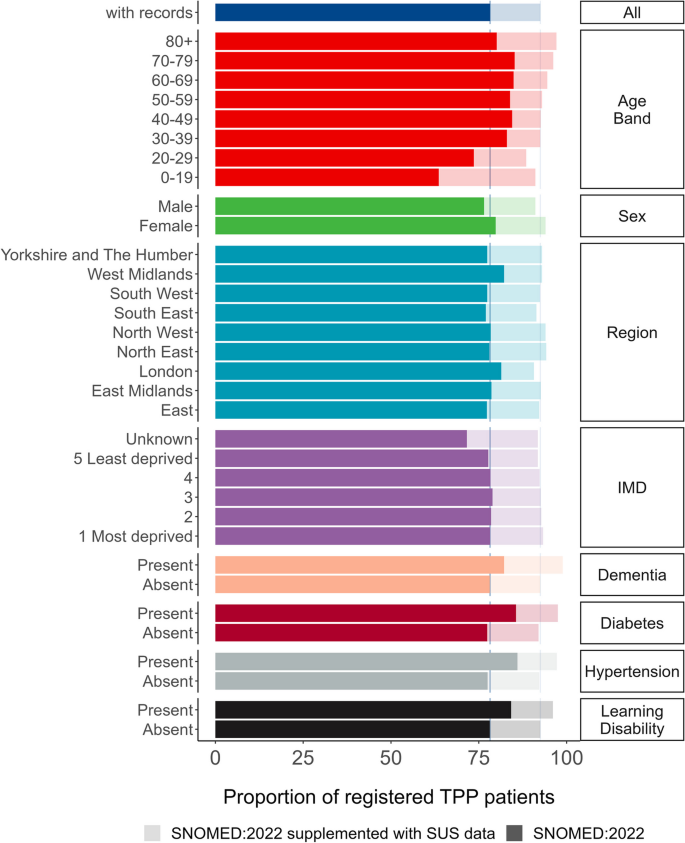
Bar plot showing proportion of registered TPP population with a recorded ethnicity by clinical and demographic subgroups, based on primary care records (solid bars) and when supplemented with secondary care data (pale bars)
Primary care ethnicity recording completeness was lowest for patients aged over 80 years (80.1%) and under 30, whereas ethnicity recording was highest in those over 80 when supplemented with secondary care data (97.1%). Women had a higher proportion of recorded ethnicities than men (79.8% and 76.5% respectively, 94% and 91.1% when supplemented with secondary care data). The completeness of primary care ethnicity recording ranged from 77% in the South East of England to 82.2% in the West Midlands. IMD was within 1.2 percentage points for known values (77.7% in the least deprived group 5 to 78.9% in group 3) and was lowest for the unknown group (71.6%). Primary care ethnicity recording was at least 4 percentage points higher in all of the clinical subgroups compared to the general population.
Distribution of ethnicity
Using ethnicity recorded in primary care only, 6.8% of the population were recorded as Asian, 2.3% Black, 1.5% Mixed, 65.6% White and 1.9% Other, and ethnicity was not recorded for 21.8%. When supplementing with hospital-recorded ethnicity data, corresponding percentages were 7.8% Asian, 2.6% Black, 1.9% Mixed, 77.9% White, 2.3% Other and 7.5% not recorded, representing a percentage point increase ranging from 0.3% in the Black group to 12.3% in the White group.
Older patients tended to have a higher rate of recorded White ethnicity (e.g. 76.3% in the 80 + group vs 50.0% in the 0–19 group), whereas younger patients had a higher rate of recording for Asian, Black, Mixed and Other groups. The higher proportion of women with recorded ethnicity was reversed in the Asian group where men (7.0% and 8.0% with secondary care data) had a higher proportion of recording than women (6.6% and 7.6% with secondary care data). The proportion of ethnicity reporting was lower for patients with dementia, hypertension or learning disabilities in every ethnic group other than White (Fig. 2 /Additional file 1: Table S2). The breakdown by 16 group ethnicity is shown in Additional file 1: Table S3. There was considerable variation in the completeness of ethnicity recording across practices with at least 1000 registered patients (Fig. 3 ).
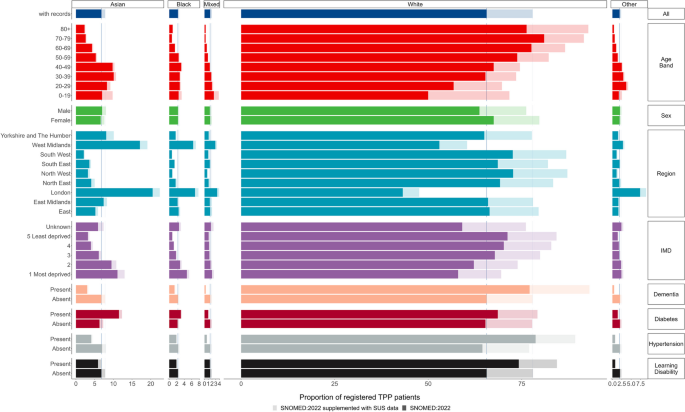
Boxplot showing the 5th, 25th, 50th, 75th and 95th percentiles of completeness of ethnicity recording across practices with at least 1000 registered patients
Consistency of ethnicity recording within patients
3.1% [260, 611] of the 19,618,135 patients with a recorded ethnicity had at least one ethnicity record that was discordant with the latest recorded ethnicity (Table 3 ). Patients whose latest recorded ethnicity was categorised as Mixed were most likely to have a discordant ethnicity recording (32.2%, 118,560), of whom 17.0% (62,565) also had a recorded ethnicity of White. 5.7% (33,205) of the 583,770 patients with the latest recorded ethnicity of Black also had a recorded ethnicity of White (Table 2 ).
Overall, for 19,364,120 (98.7%) of patients, their latest recorded ethnicity in primary care matched their most frequently recorded ethnicity in primary care (Table 3 ). 16,390,425 (99.5%) patients with the most recent ethnicity ‘White’ had matching most frequently recorded ethnicity. Other was the least concordant group, just 81.6% (399,440) of patients with the most recent ethnicity ‘Mixed’ had matching most frequently recorded ethnicity. 0.9% (5450) of patients with latest ethnicity ‘Black’ had the most frequently recorded ethnicity ‘White’ (Additional file 1: Table S4).
Of the 19.6 million total patients with a primary care ethnicity record, 12.9 million (66.0%) also had a secondary care ethnicity record. The proportion of patients with no secondary care coded ethnicity ranged from 31.9% in the White group to 58.6% in the Other group (Additional file 1: Table S5). SNOMED:2022 and secondary care coded ethnicity matched for 93.5% of patients with both coded ethnicities, ranging from 34.8% in the Mixed group to 96.9% in the White group (Fig. 4 , Additional file 1: Table S6).
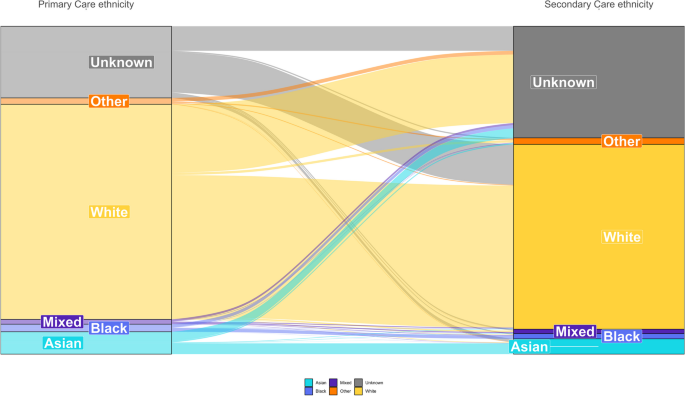
Sankey plot comparing the categorisation of ethnicity in primary care and secondary care
Comparison with the 2021 UK census population
The proportion of patients in each ethnicity group based on primary care records as of January 2022 was within 2.9 percentage points of the 2021 Census estimate (amended to the 2001 grouping) for the same ethnicity group across England as a whole (Asian: 8.7% primary care, 8.8% Census, relative difference (RD) − 1.5; Black: 3.0%, 4.2%, RD − 29.4; Mixed: 1.9%, 3.0% RD − 36.5; White: 84.0%, 81.0% RD 3.6; Other: 2.5%, 2.9%, RD − 15.1). When supplemented secondary care data, this increased to 3.2% (Fig. 5 , Additional file 1: Table S7). In primary care records, the White population was underrepresented in all regions other than the North West (7.1% percentage points higher than Census estimates), South East (2.8%) and South West (0.6%) and was most severely underestimated in the West Midlands (− 12.5%). The Asian population was overrepresented in all regions other than the North West (− 3.6%) and South East (− 1.6%) (Fig. 6 , Additional file 1: Table S8). We also compared the primary care data to the 2021 Census estimates using 2021 rather than 2001 ethnicity groups (Additional file 1: Figs. S1 and S2 and Additional file 1: Table S9).

Bar plot showing the proportion of 2021 Census and primary care populations per ethnicity grouped into 5 groups (excluding those without a recorded ethnicity (21.8% SNOMED:2020 and 7.5% supplemented with ethnicity data from secondary care)). Data labels indicate the percentage point difference between 2021 Census and TPP populations
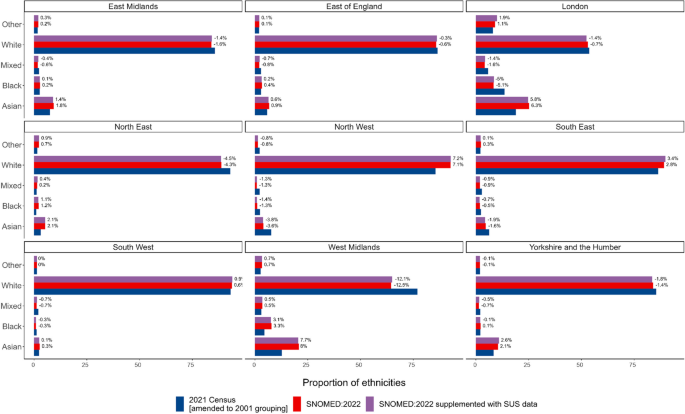
Bar plot showing the proportion of 2021 Census and TPP populations in each ethnicity group by region (excluding those without a recorded ethnicity (21.8% in primary care and 7.5% supplemented with ethnicity data from secondary care)). Data labels indicate percentage point difference between 2021 Census and TPP populations
This study reported ethnicity recording quality in around 25 million patients registered with a general practice in England and available for analysis in the OpenSAFELY-TPP database. Over three quarters of all patients had at least one ethnicity record in primary care data. When supplemented with hospital records, ethnicity recording was 92.5% complete, which is consistent with previously reported England-wide primary care data sources [ 27 , 28 ]. 98.7% of patients’ latest and most frequently recorded ethnicity matched. As the latest recorded ethnicity is computationally more efficient within OpenSAFELY, we recommend the use of the latest recorded ethnicity. The reported concordance of primary and secondary care records of 93.5% is consistent with those previously reported [ 29 ]. Despite regional variations, the overall ethnicity breakdown across all English OpenSAFELY-TPP practices was similar to the 2021 Census; however, larger relative differences were observed, in particular for the Mixed and Black groups. Therefore, relative to the size of certain ethnic groups, discrepant ethnicity recording practices may be a concern.
Strengths and weaknesses
This study provides a breakdown of primary care coding in OpenSAFELY-TPP by key clinical and demographic characteristics. The key strengths of this study are the use of large Electronic Health Record (EHR) datasets representing roughly 40% of the population of England registered with a GP, which enabled us to assess the quality of ethnicity data against a variety of important clinical characteristics.
Practices may utilise differing strategies for collecting ethnicity information from patients. Typically ethnicity is self-reported by the patient at registration or during consultation [ 30 ] but may not always be self-reported and may reflect an assumption made by the person entering the data. OpenSAFELY-TPP was missing ethnicity for 21.8% of patients, and the missingness of ethnicity data in EHRs may not be random [ 6 ].
This study focussed on the 5 Group ethnicity of the SNOMED:2022 codelists categorisation. However, there can be important variations in clinical care within these broad categories, as seen in COVID vaccine uptake [ 31 , 32 ]. More detailed categorisations, alternative coding systems and codelists have been further explored in the OpenSAFELY-TPP Ethnicity short data report.
It is common for OpenSAFELY-TPP studies to supplement the primary care recorded ethnicity, where missing, with ethnicity data from secondary care [ 10 , 11 , 33 ]. The representativeness of the CTV3:2020 coded ethnicity supplemented with SUS data has been reported previously [ 33 ]. However, secondary care data is only available for people attending hospital within the time period that data were available (currently April 2019 onwards in OpenSAFELY). The population who still have no ethnicity record after supplementation are likely very different to the wider population, for example having a much lower chance of having been admitted to hospital, or interacting with healthcare services generally.
This study represents a snapshot of ethnicity recording as of 1 January 2022 and does not provide insights into temporal trends in ethnicity recording. Trends in ethnicity recording over time are difficult to investigate due to loss of record date during transfer of clinical records when patients register with a new practice (Additional file 1: Fig. S4). Therefore, we are unable to assess the impact of QOF financial incentives being rescinded in 2011/12.
The most up-to-date formal estimates of England’s population by ethnic group currently available are from the 2021 Census. Accuracy of the 2021 Census ethnicity estimates may vary by region. The 2021 census response rate was not even between regions, ranging from 95% in London to 98% in the South East, South West and East of England [ 34 ]. The 2021 census used multiple imputation to account for missing ethnicity [ 35 ]; the percentage of eligible persons who had an ethnicity value imputed or edited was not even between regions. Imputation rate was highest in London (2.0%) and lowest in the North East (1.0%) [ 34 ].
There are limitations in comparing the GP-registered population with the census population as differences naturally arise. For example, patients registered with a GP may have left the country some years ago and hence not be counted in the census; certain populations are less likely to be registered with a GP (such as Gypsy, Roma and Traveller communities [ 36 ] and migrants [ 37 , 38 ]); not everyone responds to the census but some may be registered with a GP; and regional differences occur, for example due to students moving to cities during term-time. We looked at the GP-registered population in January 2022, whereas the census was taken in March 2021; therefore, some small changes in population also may have occurred during this time.
Findings in context
Over 20 studies have been conducted using the OpenSAFELY framework. It is important to understand the data issues with using ethnicity in OpenSAFELY. Whilst ethnicity data has been shown to be more complete for the CTV3:2020 codelist than the SNOMED:2022 codelist [ 13 ], the CTV3:2020 codelist included codes such as ‘XaJSe: Muslim—ethnic category 2001 census’ which relate to religion rather than ethnicity and were, therefore, excluded from the SNOMED:2022 codelist. The common practice of supplementing CTV3:2020 coded ethnicity with either secondary care data or the PRIMIS codelists could lead to inconsistent classification as both secondary care data and PRIMIS codelists follow the 2001 census categories.
Recording ethnicity is not straightforward. Indeed, despite often being used as a key variable to describe health, the idea of ‘ethnicity’ has been disputed [ 39 ]. Ethnicity is a complex mixture of social constructs, genetic make-up and cultural identity [ 40 ]. Self-identified ethnicity is not a fixed concept and evolving socio-cultural trends could contribute to changes in a person’s self-identified ethnic group, particularly for those with mixed heritage [ 41 ]. It is therefore perhaps not surprising to see lower levels of concordance between latest ethnicity and most common ethnicity in those with latest ethnicity coded as ‘mixed’. It is not clear to what extent this would explain all the discordance we identified or whether other factors such as data error are involved. Our findings agree with previous literature, both from the US and UK [ 5 , 41 ], which suggest that the consistency of ethnicity information tends to be highest for white populations, and lowest for Mixed or Other racial/ethnic groups [ 42 ].
The 2001 census categories are the NHS standard for ethnicity [ 17 ], but we have not been able to find any explanation for the continued use of the 2001 census categories as the standard.
Due to the significant differences experienced by ethnic groups in terms of health outcomes, accurate ethnicity coding to the most granular code possible is crucial. Although we have focussed on codelist categorisations based on the 2001 census categories, ethnicity can be extracted for each of the component codes (Additional file 1: Table S8), so researchers have the option to use custom categorisations as required.
We believe that the SNOMED:2022 codelist and codelist categorisation provides a more consistent representation of ethnicity as defined by the 2001 census categories than the CTV3:2020 codelist and should be the preferred codelist and categorisation for primary care ethnicity.
Policy implications and interpretation
This paper is principally to inform interpretation of the numerous current and future analyses completed and published using OpenSAFELY-TPP and similar UK electronic healthcare databases. The practice of supplementing primary care ethnicity with secondary care ethnicity from SUS can, depending on the study design, introduce bias and should be used with caution. For example, patients who have more clinical interactions are more likely to have a recorded ethnicity and therefore patients with a recorded ethnicity in secondary care data may tend to be sicker than the general population. Ethnicity recording has been found to be more complete for patients who died in hospital compared with those discharged [ 5 ].
This report describes the completeness and consistency of primary care ethnicity in OpenSAFELY-TPP and suggests the adoption of the SNOMED:2022 codelist and codelist categorisation as the best standard method.
Availability of data and materials
Access to the underlying identifiable and potentially re-identifiable pseudonymised electronic health record data is tightly governed by various legislative and regulatory frameworks, and restricted by best practice. The data in OpenSAFELY is drawn from General Practice data across England where TPP is the Data Processor. TPP developers (CB, JC, JP, FH and SH) initiate an automated process to create pseudonymised records in the core OpenSAFELY database, which are copies of key structured data tables in the identifiable records. These are linked onto key external data resources that have also been pseudonymised via SHA-512 one-way hashing of NHS numbers using a shared salt. Bennett Institute for Applied Data Science developers and PIs (BG, CEM, SB, AJW, KW, WJH, HJC, DE, PI, SD, GH, BBC, RMS, ID, KB, EJW and CTR) holding contracts with NHS England have access to the OpenSAFELY pseudonymised data tables as needed to develop the OpenSAFELY tools. These tools in turn enable researchers with OpenSAFELY Data Access Agreements to write and execute code for data management and data analysis without direct access to the underlying raw pseudonymised patient data and to review the outputs of this code. All code for the full data management pipeline—from raw data to completed results for this analysis—and for the OpenSAFELY platform as a whole is available for review at github.com/OpenSAFELY.
Abbreviations
Admitted patient care statistics
Clinical Terms Version 3
Emergency care
Electronic health record
General Practice Extraction Service Data for Pandemic Planning and Research
General practitioner
General Practice Extraction Service
Index of Multiple Deprivation
Nomenclature of Territorial Units for Statistics
Office for National Statistics
Outpatient attendance
Primary Care Information Services
Quality and Outcomes Framework
Systematised Nomenclature of Medicine Clinical Terms
Secondary Uses Service
Irizar P, Pan D, Kapadia D, Bécares L, Sze S, Taylor H, et al. Ethnic inequalities in COVID-19 infection, hospitalisation, intensive care admission, and death: a global systematic review and meta-analysis of over 200 million study participants. EClinicalMedicine. 2023;57:101877.
Article PubMed PubMed Central Google Scholar
Mathur R, Rentsch CT, Morton CE, Hulme WJ, Schultze A, MacKenna B, et al. Ethnic differences in SARS-CoV-2 infection and COVID-19-related hospitalisation, intensive care unit admission, and death in 17 million adults in England: an observational cohort study using the OpenSAFELY platform. Lancet. 2021;397(10286):1711–24.
Article CAS PubMed PubMed Central Google Scholar
Garlick S. Ethnic group, England and Wales - Office for National Statistics. Office for National Statistics; 2022. Available from: https://www.ons.gov.uk/peoplepopulationandcommunity/culturalidentity/ethnicity/bulletins/ethnicgroupenglandandwales/census2021 . Cited 2023 May 24.
Knox S, Bhopal RS, Thomson CS, Millard A, Fraser A, Gruer L, et al. The challenge of using routinely collected data to compare hospital admission rates by ethnic group: a demonstration project in Scotland. J Public Health. 2020;42(4):748–55.
Article CAS Google Scholar
Scobie S, Spencer J, Raleigh V. Ethnicity coding in English health service datasets. Available from: https://www.nuffieldtrust.org.uk/files/2021-06/1622731816_nuffield-trust-ethnicity-coding-web.pdf . Cited 2023 Feb 12.
Mathur R, Bhaskaran K, Chaturvedi N, Leon DA, vanStaa T, Grundy E, et al. Completeness and usability of ethnicity data in UK-based primary care and hospital databases. J Public Health. 2014;36(4):684–92.
Article Google Scholar
Contract changes 2011/12. Available from: https://web.archive.org/web/20110504084616/http://www.nhsemployers.org/PayAndContracts/GeneralMedicalServicesContract/GMSContractChanges/Pages/Contract-changes-2011-12.aspx . Cited 2023 May 24.
Kapadia, Zhang, Salway, Nazroo, Booth. Ethnic inequalities in healthcare: a rapid evidence review. NHS Race and Health. 2022. Available from: https://www.nhsrho.org/research/ethnic-inequalities-in-healthcare-a-rapid-evidence-review-3/ . Cited 2024 June 27
NHS Digital. Secondary Uses Service (SUS). Available from: https://digital.nhs.uk/services/secondary-uses-service-sus . Cited 2023 May 16.
Fisher L, Hopcroft LEM, Rodgers S, Barrett J, Oliver K, Avery AJ, et al. Changes in English medication safety indicators throughout the COVID-19 pandemic: a federated analysis of 57 million patients’ primary care records in situ using OpenSAFELY. BMJ Med. 2023;2(1):e000392. https://doi.org/10.1136/bmjmed-2022-000392 .
Nab L, Parker EP, Andrews CD, Hulme WJ, Fisher L, Morley J, Mehrkar A, et al. Changes in COVID-19-Related Mortality across Key Demographic and Clinical Subgroups in England from 2020 to 2022: A Retrospective Cohort Study Using the OpenSAFELY Platform. Lancet Public Health. 2023;8(5):e364–77.
OpenCodelists: Ethnicity codes. Available from: https://www.opencodelists.org/codelist/primis-covid19-vacc-uptake/eth2001/v1/ . Cited 2022 Sep 13.
OpenCodelists: ethnicity (SNOMED). Available from: https://www.opencodelists.org/codelist/opensafely/ethnicity-snomed-0removed/2e641f61/ . Cited 2022 Sep 13.
OpenCodelists: Ethnicity. Available from: https://www.opencodelists.org/codelist/opensafely/ethnicity/2020-04-27/ . Cited 2022 Sep 13.
PRIMIS develops the national Covid-19 Vaccination Uptake Reporting Specification. Available from: https://www.nottingham.ac.uk/primis/about/news/newslisting/primis-develops-the-national-covid-19-vaccination-uptake-reporting-specification.aspx . Cited 2022 Aug 19.
NHS Digital. General Practice Extraction Service (GPES) Data for pandemic planning and research: a guide for analysts and users of the data. Available from: https://digital.nhs.uk/coronavirus/gpes-data-for-pandemic-planning-and-research/guide-for-analysts-and-users-of-the-data . Cited 2022 Aug 19.
Ethnic Category. Available from: https://www.datadictionary.nhs.uk/data_elements/ethnic_category.html?hl=ethnic . Cited 2022 Aug 22.
Gypsy, Roma and Irish Traveller ethnicity summary. Available from: https://web.archive.org/web/20220213182343/https://www.ethnicity-facts-figures.service.gov.uk/summaries/gypsy-roma-irish-traveller . Cited 2023 Jun 6.
McLennan D, Noble S, Noble M, Plunkett E, Wright G, Gutacker N. The English Indices of Deprivation 2019 : technical report. 2019. Available from: https://dera.ioe.ac.uk/id/eprint/34259 . Cited 2022 Aug 4.
OpenCodelists: Dementia (SNOMED). Available from: https://www.opencodelists.org/codelist/opensafely/dementia-snomed/2020-04-22/ . Cited 2022 Sep 13.
OpenCodelists: Diabetes (SNOMED). Available from: https://www.opencodelists.org/codelist/opensafely/diabetes-snomed/2020-04-15/ . Cited 2022 Sep 13.
OpenCodelists: Hypertension (SNOMED). Available from: https://www.opencodelists.org/codelist/opensafely/hypertension-snomed/2020-04-28/ . Cited 2022 Sep 13.
OpenCodelists: Wider learning disability. Available from: https://www.opencodelists.org/codelist/primis-covid19-vacc-uptake/learndis/v1/ . Cited 2022 Sep 13.
Variable reference. Available from: https://docs.opensafely.org/study-def-variables/ . Cited 2022 Nov 18.
Mortality statistics - underlying cause, sex and age - Nomis - Official Labour Market Statistics. Available from: https://www.nomisweb.co.uk/datasets/mortsa . Cited 2022 Jan 28.
List of ethnic groups. Available from: https://www.ethnicity-facts-figures.service.gov.uk/style-guide/ethnic-groups . Cited 2023 Apr 17.
Wood A, Denholm R, Hollings S, Cooper J, Ip S, Walker V, et al. Linked electronic health records for research on a nationwide cohort of more than 54 million people in England: data resource. BMJ. 2021;7(373):n826.
Pineda-Moncusí M, Allery F, Delmestri A, Bolton T, Nolan J, Thygesen JH, Handy A, et al. Ethnicity Data Resource in Population-Wide Health Records: Completeness, Coverage and Granularity of Diversity. Sci Data. 2024;11(1):221.
Shiekh SI, Harley M, Ghosh RE, Ashworth M, Myles P, Booth HP, et al. Completeness, agreement, and representativeness of ethnicity recording in the United Kingdom’s Clinical Practice Research Datalink (CPRD) and linked Hospital Episode Statistics (HES). Popul Health Metr. 2023;21(1):3.
Hull SA, Mathur R, Badrick E, Robson J, Boomla K. Recording ethnicity in primary care: assessing the methods and impact. Br J Gen Pract. 2011;61(586):e290–4.
Watkinson RE, Williams R, Gillibrand S, Sanders C, Sutton M. Ethnic inequalities in COVID-19 vaccine uptake and comparison to seasonal influenza vaccine uptake in greater Manchester, UK: a cohort study. PLoS Med. 2022;19(3):e1003932.
Curtis HJ, Inglesby P, Morton CE, MacKenna B, Green A, Hulme W, et al. Trends and clinical characteristics of COVID-19 vaccine recipients: a federated analysis of 57.9 million patients’ primary care records in situ using OpenSAFELY. Br J Gen Pract. 2022;72(714):e51–62.
Article PubMed Google Scholar
Andrews C, Schultze A, Curtis H, Hulme W, Tazare J, Evans S, et al. OpenSAFELY: representativeness of electronic health record platform OpenSAFELY-TPP data compared to the population of England. Wellcome Open Res. 2022;18(7):191.
Measures showing the quality of Census 2021 estimates. Available from: https://www.ons.gov.uk/peoplepopulationandcommunity/populationandmigration/populationestimates/methodologies/measuresshowingthequalityofcensus2021estimates . Cited 2023 Feb 16.
Wardman L, Aldrich S, Rogers S. Census item edit and imputation process. Disponible en ligne:[06/01/2015]. http://www.ons.gov.uk/ons/guide-method/census/2011/census-data/2011-census-userguide/quality-and-methods/quality/quality-measures/response-and-imputation-rates/item-edit-andimputation-process.pdf . 2011. Cited 2022 Feb 3.
Tackling inequalities faced by Gypsy, roma and traveller communities. Available from: https://publications.parliament.uk/pa/cm201719/cmselect/cmwomeq/360/full-report.html . Cited 2023 May 25.
Kang C, Tomkow L, Farrington R. Access to primary health care for asylum seekers and refugees: a qualitative study of service user experiences in the UK. Br J Gen Pract. 2019;69(685):e537–45.
Knights F, Carter J, Deal A, Crawshaw AF, Hayward SE, Jones L, et al. Impact of COVID-19 on migrants’ access to primary care and implications for vaccine roll-out: a national qualitative study. Br J Gen Pract. 2021;71(709):e583–95.
Bradby H. Ethnicity: not a black and white issue. A research note. Sociol Health Illn. 1995;17(3):405–17.
Lee C. “Race” and “ethnicity” in biomedical research: how do scientists construct and explain differences in health? Soc Sci Med. 2009;68(6):1183–90.
Saunders CL, Abel GA, El Turabi A, Ahmed F, Lyratzopoulos G. Accuracy of routinely recorded ethnic group information compared with self-reported ethnicity: evidence from the English Cancer Patient Experience survey. BMJ Open. 2013;3(6):e002882.
Arday SL, Arday DR, Monroe S, Zhang J. HCFA’s racial and ethnic data: current accuracy and recent improvements. Health Care Financ Rev. 2000;21(4):107–16.
CAS PubMed PubMed Central Google Scholar
Aitken M, Tully MP, Porteous C, Denegri S, Cunningham-Burley S, Banner N, et al. Consensus statement on public involvement and engagement with data intensive health research. Int J Popul Data Sci. 2019;4(1):586.
PubMed PubMed Central Google Scholar
NHS Digital. BETA – Data Security Standards - NHS Digital. Available from: https://digital.nhs.uk/about-nhs-digital/our-work/nhs-digital-data-and-technology-standards/framework/beta---data-security-standards . Cited 2020 Apr 30.
NHS Digital. Data Security and Protection Toolkit - NHS Digital. Available from: https://digital.nhs.uk/data-and-information/looking-after-information/data-security-and-information-governance/data-security-and-protection-toolkit . Cited 2020 Apr 30.
NHS Digital. ISB1523: Anonymisation Standard for Publishing Health and Social Care Data. Available from: https://digital.nhs.uk/data-and-information/information-standards/information-standards-and-data-collections-including-extractions/publications-and-notifications/standards-and-collections/isb1523-anonymisation-standard-for-publishing-health-and-social-care-data . Cited 2023 Jul 20.
Secretary of State for Health and Social Care - UK Government. Coronavirus (COVID-19): notification to organisations to share information. 2020. Available from: https://web.archive.org/web/20200421171727/https://www.gov.uk/government/publications/coronavirus-covid-19-notification-of-data-controllers-to-shareinformation . Cited 2022 Nov 3.
Download references
Acknowledgements
We are very grateful for all the support received from the TPP Technical Operations team throughout this work and for generous assistance from the information governance and database teams at NHS England and the NHS England Transformation Directorate.
BG is guarantor.
Software and reproducibility
Data management was performed using Python 3.8, with analysis carried out using Python and R. Code for data management and analysis, as well as codelists are archived online https://github.com/opensafely/ethnicity-short-data-report/ .
Patient and public involvement
This analysis relies on the use of large volumes of patient data. Ensuring patient, professional and public trust is therefore of critical importance. Maintaining trust requires being transparent about the way OpenSAFELY works, and ensuring patient and public voices are represented in the design and use of the platform. Between February and July 2022, we ran a 6-month pilot of Patient and Public Involvement and Engagement activity designed to be aligned with the principles set out in the Consensus Statement on Public Involvement and Engagement with Data-Intensive Health Research [ 43 ]. Our engagement focused on the broader OpenSAFELY platform and comprised three sets of activities: explain and engage, involve and iterate and participate and promote. To engage and explain, we have developed a public website at opensafely.org that provides a detailed description of the OpenSAFELY platform in language suitable for a lay audience and are co-developing an accompanying explainer video. To involve and iterate, we have created the OpenSAFELY ‘Digital Critical Friends’ Group, comprised of approximately 12 members representative in terms of ethnicity, gender and educational background; this group has met every 2 weeks to engage with and review the OpenSAFELY website, governance process, principles for researchers and FAQs. To participate and promote, we are conducting a systematic review of the key enablers of public trust in data-intensive research and have participated in the stakeholder group overseeing NHS England’s ‘data stewardship public dialogue’.
The OpenSAFELY platform is principally funded by grants from:
NHS England [2023–2025];
The Wellcome Trust (222,097/Z/20/Z) [2020–2024];
MRC (MR/V015737/1) [2020–2021].
Additional contributions to OpenSAFELY have been funded by grants from:
MRC via the National Core Study programme, Longitudinal Health and Wellbeing strand (MC_PC_20030, MC_PC_20059) [2020–2022] and the Data and Connectivity strand (MC_PC_20029, MC_PC_20058) [2020–2022];
NIHR and MRC via the CONVALESCENCE programme (COV-LT-0009, MC_PC_20051) [2021–2024];
NHS England via the Primary Care Medicines Analytics Unit [2021–2024].
The views expressed are those of the authors and not necessarily those of the NIHR, NHS England, UK Health Security Agency (UKHSA), the Department of Health and Social Care or other funders. Funders had no role in the study design, collection, analysis and interpretation of data; in the writing of the report and in the decision to submit the article for publication.
Author information
Authors and affiliations.
Nuffield Department of Primary Care Health Sciences, Bennett Institute for Applied Data Science, Oxford University, Oxford, OX2 6GG, UK
Colm D. Andrews, Jon Massey, Robin Park, Helen J. Curtis, Lisa Hopcroft, Amir Mehrkar, Seb Bacon, George Hickman, Rebecca Smith, David Evans, Tom Ward, Simon Davy, Peter Inglesby, Iain Dillingham, Steven Maude, Thomas O’Dwyer, Ben F. C. Butler-Cole, Lucy Bridges, Ben Goldacre, Brian MacKenna, Alex J. Walker & William J. Hulme
London School of Hygiene and Tropical Medicine, Keppel Street, London, WC1E 7HT, UK
Rohini Mathur & Laurie A. Tomlinson
TPP, TPP House, 129 Low Lane, Horsforth, Leeds, LS18 5PX, UK
Chris Bates, John Parry, Frank Hester, Sam Harper & Jonathan Cockburn
Wolfson Institute for Population Health, University of London, London, Queen Mary, E1 2AT, UK
Rohini Mathur
You can also search for this author in PubMed Google Scholar
The OpenSAFELY Collaborative
Contributions.
Conceptualisation: CDA, BM, RP, RM, JM and WJH. Data curation: CDA, RP, RM and JM. Formal analysis: CDA, RP, RM, JM and WJH. Funding acquisition: BG. Methodology: CDA, BM, RP, RM, JM and WJH. Project administration: CDA, RP, RM and JM. Resources: CDA, RM, JM, RP, HJC, LH, LAT and BG. Software: CDA, RM, JM, RP, HJC, LH, AM, SB, GH, RS, DE, TW, SD, PI, ID, SM, TO’D, BFCBC, LB, CB, JP, FH, SH, JC, BG, BM, AJW and WJH. Supervision: AJW, LAT and WJH. Validation: CDA, BM, RP, RM, JM and WJH. Visualisation: CDA, RP, BM, BG, AJW and WJH. Writing—original draft: CDA. Writing—review and editing: CDA, AJW, BM, HJC and WJH.
Authors’ Twitter handles
Colm D Andrews-@colmresearcher.
Corresponding author
Correspondence to Colm D. Andrews .
Ethics declarations
Ethics approval and consent to participate.
NHS England is the data controller; TPP is the data processor; and the researchers on OpenSAFELY are acting with the approval of NHS England. This implementation of OpenSAFELY is hosted within the TPP environment which is accredited to the ISO 27001 information security standard and is NHS IG Toolkit compliant [ 44 , 45 ]; patient data has been pseudonymised for analysis and linkage using industry standard cryptographic hashing techniques; all pseudonymised datasets transmitted for linkage onto OpenSAFELY are encrypted; access to the platform is via a virtual private network (VPN) connection, restricted to a small group of researchers; the researchers hold contracts with NHS England and only access the platform to initiate database queries and statistical models; all database activity is logged; only aggregate statistical outputs leave the platform environment following best practice for anonymisation of results such as statistical disclosure control for low cell counts [ 46 ]. The OpenSAFELY research platform adheres to the obligations of the UK General Data Protection Regulation (GDPR) and the Data Protection Act 2018. In March 2020, the Secretary of State for Health and Social Care used powers under the UK Health Service (Control of Patient Information) Regulations 2002 (COPI) to require organisations to process confidential patient information for the purposes of protecting public health, providing healthcare services to the public and monitoring and managing the COVID-19 outbreak and incidents of exposure; this sets aside the requirement for patient consent [ 47 ]. Taken together, these provide the legal bases to link patient datasets on the OpenSAFELY platform. GP practices, from which the primary care data are obtained, are required to share relevant health information to support the public health response to the pandemic and have been informed of the OpenSAFELY analytics platform.
This study was approved by the Health Research Authority (REC reference 20/LO/0651) and by the LSHTM Ethics Board (reference 21863).
Consent for publication
Not applicable.
Competing interests
All authors declare the following: BG has received research funding from the Bennett Foundation, the Laura and John Arnold Foundation, the NHS National Institute for Health Research (NIHR), the NIHR School of Primary Care Research, NHS England, the NIHR Oxford Biomedical Research Centre, the Mohn-Westlake Foundation, NIHR Applied Research Collaboration Oxford and Thames Valley, the Wellcome Trust, the Good Thinking Foundation, Health Data Research UK, the Health Foundation, the World Health Organisation, UKRI MRC, Asthma UK, the British Lung Foundation, and the Longitudinal Health and Wellbeing strand of the National Core Studies programme; he is a Non-Executive Director at NHS Digital; he also receives personal income from speaking and writing for lay audiences on the misuse of science. BMK is also employed by NHS England working on medicines policy and clinical lead for primary care medicines data.
Additional information
Publisher's note.
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
12916_2024_3499_moesm1_esm.pdf.
Additional file 1: Fig. S1. Bar plot showing the proportion of 2021 Census and TPP populations (amended to 2021 grouping) per ethnicity grouped into 5 groups (excluding those without a recorded ethnicity). Annotated with percentage point difference between 2021 Census and TPP populations. Fig S2. Bar plot showing the proportion of 2021 Census and TPP populations (amended to 2021 grouping) per ethnicity grouped into 5 groups per NUTS − 1 region (excluding those without a recorded ethnicity). Annotated with percentage point difference between 2021 Census and TPP populations. Fig. S3. Recording of ethnicity over time for latest and first recorded ethnicity. Unknown dates of recording may be stored as ‘1900 − 01 − 01’. Table S1. Count of patients with a recorded ethnicity in OpenSAFELY-TPP (proportion of registered TPP population) by clinical and demographic subgroups. All counts are rounded to the nearest 5. Table S2. Count of patients with a recorded ethnicity in OpenSAFELY TPP by ethnicity group (proportion of registered TPP population) and clinical and demographic subgroups. All counts are rounded to the nearest 5. Table S3. Count of patients with a recorded ethnicity in OpenSAFELY TPP by ethnicity group (proportion of registered TPP population) and clinical and demographic subgroups. All counts are rounded to the nearest 5. Table S4. Count of patients’ most frequently recorded ethnicity (proportion of latest ethnicity). Table S6. Count of patients with a recorded ethnicity in Secondary Care by ethnicity group excluding Unknown ethnicites (proportion of Primary Care population). All counts are rounded to the nearest 5. Table S7. Count of patients with a recorded ethnicity in OpenSAFELY TPP by ethnicity group (proportion of registered TPP population) and 2021 ONS Census counts [amended to 2001 grouping] (proportion of 2021 ONS Census population). All counts are rounded to the nearest 5. Table S8. Count of patients with a recorded ethnicity in OpenSAFELY TPP [amended to the 2021 ethnicity grouping] (proportion of registered TPP population) and 2021 ONS Census counts (proportion of 2021 ONS Census population). All counts are rounded to the nearest 5. Table S9. Count of individual ethnicity code use.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/ . The Creative Commons Public Domain Dedication waiver ( http://creativecommons.org/publicdomain/zero/1.0/ ) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
Reprints and permissions
About this article
Cite this article.
The OpenSAFELY Collaborative., Andrews, C.D., Mathur, R. et al. Consistency, completeness and external validity of ethnicity recording in NHS primary care records: a cohort study in 25 million patients’ records at source using OpenSAFELY. BMC Med 22 , 288 (2024). https://doi.org/10.1186/s12916-024-03499-5
Download citation
Received : 31 January 2024
Accepted : 24 June 2024
Published : 10 July 2024
DOI : https://doi.org/10.1186/s12916-024-03499-5
Share this article
Anyone you share the following link with will be able to read this content:
Sorry, a shareable link is not currently available for this article.
Provided by the Springer Nature SharedIt content-sharing initiative
- Primary care health sciences
- Electronic health records
- Data curation
BMC Medicine
ISSN: 1741-7015
- Submission enquiries: [email protected]
- General enquiries: [email protected]

Infosys partners with a Government department for implementation of Electronic Health records
The Electronic Health Records (EHR) project is a massive one and so a demonstrator was to be built for a particular county. This would function as a test bed for proving the concept and the operations would yield new ideas for actual fulltime implementation. Technically, the EHR would enable the General Practitioner (GP) to view a patient's full details and also help doctors giving emergency treatment to quickly determine the allergies and previous history of diseases but until a demonstrator was built and deployed, the actual evidence would not emerge. Infosys had to build a demonstrator to show what was actually possible and also suggest improvements.
Key Challenges
Complexities were overwhelming and the ownership of the patient was confusing
Patient care services were impacted by rising costs
An increasing proportion of the ageing population had long-term diseases
Data provided would come from disparate legacy systems
To enable doctors to keep abreast of the latest developments, Infosys developed a Knowledge Management (KM) portal subsystem
Infosys used a Web Services model to ensure that EHR could utilize the information to deliver accurate results.
The Solution
Integrate disparate legacy systems
Looking for a breakthrough solution?

Infosys relied on the Global Delivery Model to conclusively depict the benefits of EHR. As a first step, Infosys analyzed the existing high-level user requirement documents and assisted in preparing requirements for defining the solution of the EHR demonstrator.

Digital Healthcare ecosystem
Ready for Disruption?
Next Case Study
Developing a PHR solution for a health insurer


COMMENTS
We describe surveillance case studies and future directions for enhancing opportunities to use EHR data for public health surveillance. ... Electronic health records (EHRs) contain extensive longitudinal health information about patients and populations (1). Over the last decade, prompted by federal meaningful use guidelines and incentives ...
Abstract. Implementing an electronic health record (EHR) can be a difficult task to take on and planning the process is of utmost importance to minimize errors. Evaluating the selection criteria and implementation plan of an EHR system, intending interoperability, confidentiality, availability, and integrity of the patient health information ...
Despite the great advances in the field of electronic health records (EHRs) over the past 25 years, implementation and adoption challenges persist, and the benefits realized remain below expectations. This scoping review aimed to present current knowledge ...
Read EHR case studies and hear stories of doctors, practices and organizations that have implemented EHRs.
In this chapter, we present each of our four case study sites in turn. We summarise key facts about the four sites in Table 9. Our case study ambulance services used a range of terminology for describing their electronic records systems, including electronic PRFs, electronic PCRs and electronic patient records. For consistency, we use the expression 'electronic health records' ('EHRs ...
A qualitative study design was used. We collected the opinions from different groups of clinicians (physicians, hospitalists, nurse practitioners, nurses, and patient safety officers) using semi-structured interviews. Organizations represented were trauma hospitals, academic medical centers, medical clinics, home health centers, and small hospitals.
With the increasing availability of rich, longitudinal, real-world clinical data recorded in electronic health records (EHRs) for millions of patients, there is a growing interest in leveraging ...
The Shenandoah Community Health Center (SCHC) in Martinsburg, WV, is a federally qualified health center with about 30,000 patients that averages 129,000 visits per year, and has been working on implementing electronic health records and achieving meaningful use. It is also a Level 1 NCQA Certified Medical Home and Joint Commission Accredited.
Abstract. There is a national focus on the adoption and use of electronic health records (EHRs) with electronic prescribing (e-Rx) for the goal of providing safe and quality care. Although there is a large body of literature on the benefits of adoption, there is also increasing evidence of the unintentional consequences resulting from use.
This retrospective cohort study used electronic health records from TriNetX US Collaborative Network, covering >100 million patients in the USA.
Well before the Covid-19 pandemic struck, electronic health records were the bane of physicians' existences. In all too many cases, EHRs seemed to create a huge amount of extra work and generate ...
This mixed-methods study was conducted among CMC, their family caregivers, and physicians in SC. Electronic health records data from a primary care clinic within a large health system (7/1/2022-6/30/2023) was analyzed. Logistic regression examined factors associated with hospitalizations among CMC.
This cross-sectional study examines whether electronic health record (EHR) system safety performance is associated with EHR frontline user experience in a national sample of hospitals.
Rural Health Clinic Exchanges Information with Hospitals and Physicians for Improved Coordination of Care
One of the structures that centralizes many of the tools for achieving comprehensive and convenient sleep health lifelong is the electronic health record (EHR). EHRs that are widely distributed and provide patient access and communication can be designed to foster best practices, coordinate more eficient care long-term, and provide proof of efectiveness in achieving outcomes. After examining ...
The surge in electronic health records (EHRs), big data analytics, and telemedicine raises significant questions about privacy, consent, data ownership, and equity. Integrating these technologies into our healthcare systems is crucial to navigating these ethical dilemmas thoughtfully.
The second part describes two implementations of electronic medical record systems and compares the theory against the findings of these two case studies. The paper provides implementers with research-informed guidance about effective implementation, contributes to developing implementation theory and notes policy implications for current ...
The purpose of this paper is to do a case study on the implementation of an EHR system in a local healthcare institution, and then to analyze this case study to give directions so as to avoid some arising issues. Download to read the full chapter text.
Objective: We aimed to present a descriptive case study of the implementation of an open source electronic health record system in public health care facilities in Kenya.
This cohort study assesses the ability of a natural language processing model to classify patient-initiated electronic health record (EHR) messages and triage COVID-19 cases to reduce clinician response time and improve access to antiviral treatment.
Background In contrast to the acute hospital sector, there have been relatively few implementations of integrated electronic health record (EHR) systems into specialist mental health settings. The National Programme for Information Technology (NPfIT) in England was the most expensive IT-based transformation of public services ever undertaken, which aimed amongst other things, to implement ...
High-quality data are fundamental to healthcare research, future applications of artificial intelligence and advancing healthcare delivery and outcomes through a learning health system. Although routinely collected administrative health and electronic medical record data are rich sources of information, they have significant limitations.
Background: Electronic health records (EHRs) are a promising approach to document and map (complex) health information gathered in health care worldwide. However, possible unintended consequences during use, which can occur owing to low usability or the lack of adaption to existing workflows (e.g., high cognitive load), may pose a challenge. To prevent this, the involvement of users in the ...
The purpose of this study was to develop and validate an algorithm for identifying Veterans with a history of traumatic brain injury (TBI) in the Veterans Affairs (VA) electronic health record usin...
Mobile health (mHealth) technologies, such as wearable devices and sensors that can be placed in the home, allow for the capture of physiologic, behavioral, and environmental data from patients between clinic visits. The inclusion of these data in the medical record may benefit patients and providers. Most health systems now have electronic health records (EHRs), and the ability to pull and ...
Methods: AESI phenotype algorithms were developed to apply to electronic health record data at health provider organizations across the country by querying for standard and interoperable codes. The codes queried in the rules represent symptoms, diagnoses, or treatments of the AESI sourced from published case definitions and input from clinicians.
At the same time, improved data collection efficiency reduces clinical trial costs. Objective: Explore how to extract clinical trial-related data from hospital electronic health record system (EHR), transform the data into an electronic data capture system (EDC) required format, and transfer it into sponsor's environment.
PDF | The main objective of this study was to explore the impact of EHRs on healthcare quality at the Asamankese Government Hospital. The research used... | Find, read and cite all the research ...
The key strengths of this study are the use of large Electronic Health Record (EHR) datasets representing roughly 40% of the population of England registered with a GP, which enabled us to assess the quality of ethnicity data against a variety of important clinical characteristics.
The Electronic Health Records (EHR) project is a massive one and so a demonstrator was to be built for a particular county. This would function as a test bed for proving the concept and the operations would yield new ideas for actual fulltime implementation. Technically, the EHR would enable the General Practitioner (GP) to view a patient's ...