This store requires javascript to be enabled for some features to work correctly
Free, fast shipping on orders over €35
- Complete regime
- Moisturisers
- Skin consult
- The OT blog


100% natural skincare
Why choose our 100% natural organic skincare.

Results driven skincare
We prioritise high quality botanicals, probiotics and actives for pure, clean and simple skincare that delivers.

Perfect for sensitive skin
You will not find common allergens in our products such as fragrances and drying alcohols.

Vegan and cruelty free
As proud parents of 3 four legged friends, we promise that our products are totally vegan and never tested on animals.

Crafted with care
We are a proudly independent brand, where our woman led team formulate and produce everything in-house.
Check out what our community loves

Meet Aleksandra, your skincare expert
Thanks for checking out our 100% natural plant-based, eco-friendly skincare. Our collection, free from harsh chemicals and allergens, includes gentle cleansers, toners, serums, and moisturizers. We believe in beauty that's kind to both you and the planet. Choose us for a natural, skin-friendly journey.
As featured in
Discover our 100% natural organic skincare, crafted with care in Budapest. Start your 'Clean Health' journey with our cruelty-free, plant-based skin essentials. Transform your natural skincare regime today.
From the Organic blog
Understanding inci labels: expert guide by aleksandra crabtree for smart skincare choices.
We ship to Europe and the UK
Free, fast shipping on orders over €35.
Guaranteed results in 60 days
No results, then get your money back. No questions asked.
We are available 24/7
Contact us by chat, use our contact form , or send us an email.
No unexpected charges
Duties, customs and taxes are included in the final price at checkout
- American Express
© 2024 BDPST LABS Kft, trading as Organic Thesis
Coco & Gizmo
Cruelty-free.
Thesis was created in 2009 by a family of vegetarians that wanted to create a truly pure range of skincare. Their vegan and cruelty-free range is also, eco-friendly :) They use premium certified organic ingredients and no harsh chemicals and synthetics. Their products are full of pure ingredients where there has been no compromise on quality. Made by hand in small batches ensures superior quality of these luxury products. Thesis super-charged, eco-luxe skincare will leave your skin feeling nourished and getting the care it deserves...
Vegetarian Society Vegan Approved • PETA Vegan & Cruelty Free Approved • Leaping Bunny Cruelty Free Approved Brand

Thesis Facial Mask Mermaid's Cheek
Thesis Beauty Organic Skin Care Serums Review + Giveaway–Ends June 7, 2018

With so many skin care products on the market, making the right choice can be hit or miss. However, Thesis Beauty takes a healthier approach with their line of certified organic skin care judging from two products that I got to try for this review and giveaway with their Face and Eye Serums.
Thesis Beauty Takes Organic Skin Care Seriously
Though Thesis Beauty has been around since 2009, this was the first time that I tried their organic skin care. All their formulas are handcrafted and use 99% pure certified organic ingredients without any synthetics or fillers that may compromise their high standards to preserve the power of nature.
An important thing to note is many of those oils are raw, cold-pressed ones. This means they are more potent and aromatic to maximum their effectiveness for skin care.
To further protect the stability of their products, Thesis Beauty uses dark amber glass bottles for their serums to protect against light from causing deterioration.
The Face Serum that I sampled is an intense concentration of 10 oils such as rosehip, jojoba oil, sunflower, argan nut oil, grapeseed oil, macadamia, and six essential oils such as sweet orange and geranium oils to hydrate and revive dry, mature and dehydrated skin. This particular serum has a pleasant hint of orange fragrance once you pump this yellow, mildly scented serum out.
**Disclaimer: This post has been compensated through a free product (s) and/or monetary payment. Opinions are solely mine.**
One pump is enough of this rich serum to treat your whole face. This product is fast absorbing once gently patted into your skin. Though I haven’t been using this serum for all that long, I found it lightweight and gave my complexion a brighter glow and softness. Only time will tell what else this face serum will do for anti-aging.
I also got to try Thesis Beauty’s Eye Serum . This nourishing formula is full of antioxidants to help with anti-aging, dryness and relieve puffiness. With oils like jojoba, grapeseed oil, argan nut oil, rosehip oil and other essential oils such as lavender and camomile just to name a few, this is a hardworking, organic eye serum that does wonders for the eye area.

Anyone with spring allergies might have more eye puffiness than usual. When I tried this eye serum, I found it smoothed the under eye area nicely and also helped counteract some puffiness to my amazement. Being a serum also made it disappear into the skin faster than some eye creams while leaving a moisturized base for my concealer.
I don’t know how you feel about this, but I prefer keeping my skin care simple with pure ingredients instead of flooding it with countless chemicals. If you feel the same way and want organic skin care that you can trust, I recommend visiting Thesis Beauty and try their products. You just might be surprised at the difference of what healthy skin really looks like.
Thanks to Thesis Beauty , one lucky Beauty Cooks Kisses reader in the United States wins both the Dry Skin Face Serum ($24.95) and the Eye Serum ($24.95) for a ($49.90 total value)!
Giveaway details and How to Enter–
Prize: One (1) winner from the USA will win full-size bottles of Thesis Beauty’s Dry Skin Face Serum and Eye Serum ($49.90 total value)!
Dates of entry: 05/21/2018–06/07/2018 at 11:59 PM Eastern Standard Time
Eligibility: Open to the USA. The winner will need to respond to my email with name and shipping address within 48-hours or another winner will be picked.
How to Enter: Please use the Rafflecopter widget if it works. You can also enter by clicking this link or try to enter using another browser. You can return daily and log in again with your email to tweet.
You may also enjoy:
Why spiffy socks can help keep your …, why you want this inexpensive cordless vacuum …, how to get a free organic turkey …, how i improved my health and weight …, 53 comments.
Hiii I love that they are cruelty free and vegan. Turns out I was following them on Instagram. Thanks!
I learned that they do not use any synthetic or disruptive chemical.s
It’s made in the USA. Love that!
vegan and cruelty free
I learned several things about the Thesis Beauty. First of all that all products are produced in small batches and by hand, for freshness and potency. I also learned that many of the products are made with food grade ingredients.
Love how they are pure Ingredients organic, mostly raw and food-grade. Made in the USA
It’s wonderful that their products are pure and vegan…
I learned that they make products for dry skin.
I learned the products are made from Premium certified organic ingredients.
I learned that they are vegan and cruelty-free.
They are vegan and cruelty free
The products are made in the USA.
I love that their products are safe for use on babies! That’s gentle!
I learned that this company does not use synthetics or fillers.
Good to know that they’re vegan!
I learned their products have no synthetics and no fillers.
Everything in the products is pure and organic. No artificial anything.
that they are chemical free
I learned that the products are vegan, cruelty free, and there are no chemicals.
I learned that they are vegan and cruelty free love it.
I learned that it is raw focused and chemical free. Exactly the kind you want!
I learned that these products are all natural and chemical free – exactly what I have been looking for!
I learned that it is organic and vegan.
Free samples with every order! I love that!
I learned they are cruelty free
I am excited to try these products because I have dry skin, especially my under eyes, and these products are geared towards that. I’m especially excited to try the eye serum because I struggle especially with dry under eyes.
They use pure ingredients,they do not use Synthetic and Disruptive chemicals.
They use pure ingredients and they do not use Synthetic and Disruptive chemicals.
I learned that it is a family owned business that started i 2009.
I learned that they use the most potent certified organic botanicals. Thanks for the giveaway!
Learned that it is made here is the US thats great!
I learned since 2009, Thesis has been run by a family of happy vegetarians dedicated to the development of truly organic and natural skin care products at accessible price points.
I love quality skincare products and these sound nice.
I would like to try these because they are all natural and vegan. I could really use the eye serum.
i learned that your products are Vegan and cruelty-free
i am most excited to try the eye serum
I learned that your products are all Pure Ingredients organic, mostly raw and food-grade
I learned that they are a family run business and take purity in products seriously.
I like that the ingredients are pure.
I like that the ingredients are mostly raw and food grade and that they give free samples with their orders.
No synthetics. No fillers. No gimmicks.
I learned their products are cruelty free!
I learned All items are produced in small batches and by hand to keep things fresh and most potent.
Pure Ingredients organic, mostly raw and food-grade. That’s it!
You get free samples with every order, and right now there is a promotion to get a free travel bag with any 2 Face Care products.
I love the purity and the fact that they use dark glass bottles to protect the quality of the product!
That it’s as pure enough to use on a baby.
I learned that they are eco-friendly packaging with their products.
Love that the products are vegan and cruelty free
I learned their products are Vegan and Cruelty Free.
I learned they are chemical-free, that is important to me.
I learned they have an offer for a free travel bag with any 2 face products
Leave a Reply Cancel reply
This site uses Akismet to reduce spam. Learn how your comment data is processed .
- Beauty & Personal Care
- Treatments & Masks
Sorry, there was a problem.

Image Unavailable

- To view this video download Flash Player
Organic Facial Serum - Unscented - Soothing Antioxidant Moisturizer for Sensitive Skin with Sea Buckthorn
| Skin Type | mature, Combination, Sensitive, Dry, Normal |
| Scent | Unscented |
| Brand | Thesis |
| Item Form | Oil |
| Material Feature | Organic, Natural |
About this item
- 99% organic ingredients - extra high organic content plus natural vitamin E
- Luxurious organic oils fight free radicals and soothe your skin - Aloe, Sea Buckthorn, Grapeseed
- No synthetic chemicals, preservatives, fragrance, water & fillers - only potent cold-pressed, unrefined oils
- Ideal for sensitive, reactive, irritated skin (also great for all skin types, except acneic skin)
- 1 fl.oz / 30 ml dark amber apothecary bottle with pump
Product Description

Unscented Serum will rescue irritated, sensitive skin and bring soothing comfort in an instant. Does not contain any added scents for extra comfort.

Sensitive skin is very fragile and reactive to harsh chemicals and scents.This super-gentle Unscented serum is designed to rescue irritated, sensitive skin by bringing soothing comfort in an instant. It contains organic oils and does not use any added scents - for extra comfort. This moisturizing facial serum employs truly luxurious organic oils such as cold-pressed organic Grapeseed oil, cold-pressed organic Jojoba oil, cold-pressed organic Apricot oil and organic Sea Buckthorn extract - all loaded with powerful antioxidants and natural essential vitamins for the lasting comfort and glowing appearance. Cold-pressed organic Grapeseed oil, being a great moisturizer abundant in microelements, antioxidants and vitamins, such as vitamin A, B, E, promotes skin radiance and helps fight the aging effects of the elements. Due to grapeseed oil’s high linoleic acid value (greater than most other carrier oils), this oil is highly moisturizing, absorbs very quickly without clogging pores and without leaving greasy feeling. Organic Sea Buckthorn is an incredible source of carotenoids and essential vitamins and nutrients that fight free radicals.

Product details
- Is Discontinued By Manufacturer : No
- Package Dimensions : 5.7 x 3 x 2.3 inches; 0.96 ounces
- Item model number : fc-ser-unsc-1floz
- UPC : 856367002702
- Manufacturer : Thesis Beauty
- ASIN : B007RS3W3U
- #12,361 in Facial Serums
Important information
Ingredients.
Organic Apricot Oil; Organic Sunflower Oil; Organic Jojoba Oil; Organic Grapeseed Oil; Organic Sea Buckthorn Extract; Natural Vitamin E Complex, Soy-free D-alpha, beta, gamma, delta Tocopherols and Tocotrienols); Organic Rosemary Extract
Morning and evening, apply a very small amount (usually about 1 pump) - when your skin is still damp after cleansing. Gently tap the serum into your skin, spreading a thin layer with rolling upward motion all over your face, neck and décolleté. Avoid stretching your skin as this may lead to fine line formation. Leave for a few minutes to let sink in. Excellent under makeup.
Looking for specific info?
Customer reviews.
Customer Reviews, including Product Star Ratings help customers to learn more about the product and decide whether it is the right product for them.
To calculate the overall star rating and percentage breakdown by star, we don’t use a simple average. Instead, our system considers things like how recent a review is and if the reviewer bought the item on Amazon. It also analyzed reviews to verify trustworthiness.
Customers say
Customers like the hydration, feel and quality of the skin serum. They mention that it's excellently moisturizing, good for dry skin and that it adds a lot of moisture to their skin. They also say that it makes their skin feel dewy soft, healthy and smooth. Customers appreciate the quality and consistency of the oil. They say that the oil is luxurious and doesn't require high amounts.
AI-generated from the text of customer reviews
Customers find the skin serum excellently moisturizing, good for dry skin, and adds a lot of moisture to their skin. They say it's calming and a perfect night time moisturizer. Customers also say the oil makes their skin supple, soft, and helps remove redness.
"...This facial serum with Sea buckthorn hydrates so wonderfully making my skin feel healthy and smooth...." Read more
"...and other unnecessary ingredients, this oil will make your skin supple and glowing . It is almost like a vitamin for your skin...." Read more
"...It is all natural and made with high quality oils. Leaves your skin soft and has definitely helped remove redness from my skin." Read more
"...This serum is great for adding extra moisture and giving my skin a nice glow...." Read more
Customers like the feel of the skin serum. They say it makes their skin feel dewy, soft, and healthy. Some customers also mention that it did not irritate their skin, but did make them congested. They also say the lines just smoothed out.
"...serum with Sea buckthorn hydrates so wonderfully making my skin feel healthy and smooth ...." Read more
"I purchased this for my dry, sensitive skin. It did not irritate my skin , but it did make me congested...." Read more
"...Nope. Not puffy. The lines just smoothed out . No itch, either!..." Read more
Customers are satisfied with the quality of the skin serum. They mention that it is all natural and made with high-quality oils.
"...It is all natural and made with high quality oils . Leaves your skin soft and has definitely helped remove redness from my skin." Read more
" Nice product , didn't realize the amount a little pricey for the quantity, but feels nice on the skin." Read more
" Wonderful product , especially for winter. I use it over my serum and it keeps my skin moisturized and glowing." Read more
" Great product !" Read more
Customers are satisfied with the residue of the skin serum. They mention that it is a natural oil with a nice consistency that isn't too oily on their face. They also appreciate the luxurious oil, saying that it does not require high amounts.
"...It absorbs very quickly without any oily residue. Oils do not require high amounts ; one drop or perhaps two is sufficient enough...." Read more
"...I loved it! It is unscented, natural oil with a nice consistency that isn't too oily on your face ...." Read more
"Excellently moisturizing with no greasy residue . I put it on last night and woke up with far more luminous skin and diminished dehydration lines." Read more
- Sort reviews by Top reviews Most recent Top reviews
Top reviews from the United States
There was a problem filtering reviews right now. please try again later..
Top reviews from other countries
Disclaimer : While we work to ensure that product information is correct, on occasion manufacturers may alter their ingredient lists. Actual product packaging and materials may contain more and/or different information than that shown on our Web site. We recommend that you do not solely rely on the information presented and that you always read labels, warnings, and directions before using or consuming a product. For additional information about a product, please contact the manufacturer. Content on this site is for reference purposes and is not intended to substitute for advice given by a physician, pharmacist, or other licensed health-care professional. You should not use this information as self-diagnosis or for treating a health problem or disease. Contact your health-care provider immediately if you suspect that you have a medical problem. Information and statements regarding dietary supplements have not been evaluated by the Food and Drug Administration and are not intended to diagnose, treat, cure, or prevent any disease or health condition. Amazon.com assumes no liability for inaccuracies or misstatements about products.
- About Amazon
- Investor Relations
- Amazon Devices
- Amazon Science
- Sell products on Amazon
- Sell on Amazon Business
- Sell apps on Amazon
- Become an Affiliate
- Advertise Your Products
- Self-Publish with Us
- Host an Amazon Hub
- › See More Make Money with Us
- Amazon Business Card
- Shop with Points
- Reload Your Balance
- Amazon Currency Converter
- Amazon and COVID-19
- Your Account
- Your Orders
- Shipping Rates & Policies
- Returns & Replacements
- Manage Your Content and Devices
- Conditions of Use
- Privacy Notice
- Consumer Health Data Privacy Disclosure
- Your Ads Privacy Choices
- Research article
- Open access
- Published: 12 June 2019
The impact of skin care products on skin chemistry and microbiome dynamics
- Amina Bouslimani 1 na1 ,
- Ricardo da Silva 1 na1 ,
- Tomasz Kosciolek 2 ,
- Stefan Janssen 2 , 3 ,
- Chris Callewaert 2 , 4 ,
- Amnon Amir 2 ,
- Kathleen Dorrestein 1 ,
- Alexey V. Melnik 1 ,
- Livia S. Zaramela 2 ,
- Ji-Nu Kim 2 ,
- Gregory Humphrey 2 ,
- Tara Schwartz 2 ,
- Karenina Sanders 2 ,
- Caitriona Brennan 2 ,
- Tal Luzzatto-Knaan 1 ,
- Gail Ackermann 2 ,
- Daniel McDonald 2 ,
- Karsten Zengler 2 , 5 , 6 ,
- Rob Knight 2 , 5 , 6 , 7 &
- Pieter C. Dorrestein 1 , 2 , 5 , 8
BMC Biology volume 17 , Article number: 47 ( 2019 ) Cite this article
101k Accesses
88 Citations
122 Altmetric
Metrics details
Use of skin personal care products on a regular basis is nearly ubiquitous, but their effects on molecular and microbial diversity of the skin are unknown. We evaluated the impact of four beauty products (a facial lotion, a moisturizer, a foot powder, and a deodorant) on 11 volunteers over 9 weeks.
Mass spectrometry and 16S rRNA inventories of the skin revealed decreases in chemical as well as in bacterial and archaeal diversity on halting deodorant use. Specific compounds from beauty products used before the study remain detectable with half-lives of 0.5–1.9 weeks. The deodorant and foot powder increased molecular, bacterial, and archaeal diversity, while arm and face lotions had little effect on bacterial and archaeal but increased chemical diversity. Personal care product effects last for weeks and produce highly individualized responses, including alterations in steroid and pheromone levels and in bacterial and archaeal ecosystem structure and dynamics.
Conclusions
These findings may lead to next-generation precision beauty products and therapies for skin disorders.
The human skin is the most exposed organ to the external environment and represents the first line of defense against external chemical and microbial threats. It harbors a microbial habitat that is person-specific and varies considerably across the body surface [ 1 , 2 , 3 , 4 ]. Recent findings suggested an association between the use of antiperspirants or make-up and skin microbiota composition [ 5 , 6 , 7 ]. However, these studies were performed for a short period (7–10 days) and/or without washing out the volunteers original personal care products, leading to incomplete evaluation of microbial alterations because the process of skin turnover takes 21–28 days [ 5 , 6 , 7 , 8 , 9 ]. It is well-established that without intervention, most adult human microbiomes, skin or other microbiomes, remain stable compared to the differences between individuals [ 3 , 10 , 11 , 12 , 13 , 14 , 15 , 16 ].
Although the skin microbiome is stable for years [ 10 ], little is known about the molecules that reside on the skin surface or how skin care products influence this chemistry [ 17 , 18 ]. Mass spectrometry can be used to detect host molecules, personalized lifestyles including diet, medications, and personal care products [ 18 , 19 ]. However, although the impact of short-term dietary interventions on the gut microbiome has been assessed [ 20 , 21 ], no study has yet tested how susceptible the skin chemistry and Microbiome are to alterations in the subjects’ personal care product routine.
In our recent metabolomic/microbiome 3D cartography study [ 18 ], we observed altered microbial communities where specific skin care products were present. Therefore, we hypothesized that these products might shape specific skin microbial communities by changing their chemical environment. Some beauty product ingredients likely promote or inhibit the growth of specific bacteria: for example, lipid components of moisturizers could provide nutrients and promote the growth of lipophilic bacteria such as Staphylococcus and Propionibacterium [ 18 , 22 , 23 ]. Understanding both temporal variations of the skin microbiome and chemistry is crucial for testing whether alterations in personal habits can influence the human skin ecosystem and, perhaps, host health. To evaluate these variations, we used a multi-omics approach integrating metabolomics and microbiome data from skin samples of 11 healthy human individuals. Here, we show that many compounds from beauty products persist on the skin for weeks following their use, suggesting a long-term contribution to the chemical environment where skin microbes live. Metabolomics analysis reveals temporal trends correlated to discontinuing and resuming the use of beauty products and characteristic of variations in molecular composition of the skin. Although highly personalized, as seen with the microbiome, the chemistry, including hormones and pheromones such as androstenone and androsterone, were dramatically altered. Similarly, by experimentally manipulating the personal care regime of participants, bacterial and molecular diversity and structure are altered, particularly for the armpits and feet. Interestingly, a high person-to-person molecular and bacterial variability is maintained over time even though personal care regimes were modified in exactly the same way for all participants.
Skin care and hygiene products persist on the skin
Systematic strategies to influence both the skin chemistry and microbiome have not yet been investigated. The outermost layer of the skin turns over every 3 to 4 weeks [ 8 , 9 ]. How the microbiome and chemistry are influenced by altering personal care and how long the chemicals of personal care products persist on the skin are essentially uncharacterized. In this study, we collected samples from skin of 12 healthy individuals—six males and six females—over 9 weeks. One female volunteer had withdrawn due to skin irritations that developed, and therefore, we describe the remaining 11 volunteers. Samples were collected from each arm, armpit, foot, and face, including both the right and left sides of the body (Fig. 1 a). All participants were asked to adhere to the same daily personal care routine during the first 6 weeks of this study (Fig. 1 b). The volunteers were asked to refrain from using any personal care product for weeks 1–3 except a mild body wash (Fig. 1 b). During weeks 4–6, in addition to the body wash, participants were asked to apply selected commercial skin care products at specific body parts: a moisturizer on the arm, a sunscreen on the face, an antiperspirant on the armpits, and a soothing powder on the foot (Fig. 1 b). To monitor adherence of participants to the study protocol, molecular features found in the antiperspirant, facial lotion, moisturizer, and foot powder were directly tracked with mass spectrometry from the skin samples. For all participants, the mass spectrometry data revealed the accumulation of specific beauty product ingredients during weeks 4–6 (Additional file 1 : Figure S1A-I, Fig. 2 a orange arrows). Examples of compounds that were highly abundant during T4–T6 in skin samples are avobenzone (Additional file 1 : Figure S1A), dexpanthenol (Additional file 1 : Figure S1B), and benzalkonium chloride (Additional file 1 : Figure S1C) from the facial sunscreen; trehalose 6-phosphate (Additional file 1 : Figure S1D) and glycerol stearate (Additional file 1 : Figure S1E) from the moisturizer applied on arms; indolin (Additional file 1 : Figure S1F) and an unannotated compound ( m/z 233.9, rt 183.29 s) (Additional file 1 : Figure S1G) from the foot powder; and decapropylene glycol (Additional file 1 : Figure S1H) and nonapropylene glycol (Additional file 1 : Figure S1I) from the antiperspirant. These results suggest that there is likely a compliance of all individuals to study requirements and even if all participants confirmed using each product every day, the amount of product applied by each individual may vary. Finally, for weeks 7–9, the participants were asked to return to their normal routine by using the same personal care products they used prior to the study. In total, excluding all blanks and personal care products themselves, we analyzed 2192 skin samples for both metabolomics and microbiome analyses.
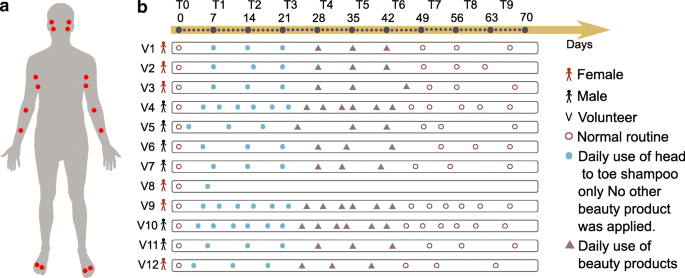
Study design and representation of changes in personal care regime over the course of 9 weeks. a Six males and six females were recruited and sampled using swabs on two locations from each body part (face, armpits, front forearms, and between toes) on the right and left side. The locations sampled were the face—upper cheek bone and lower jaw, armpit—upper and lower area, arm—front of elbow (antecubitis) and forearm (antebrachium), and feet—in between the first and second toe and third and fourth toe. Volunteers were asked to follow specific instructions for the use of skin care products. b Following the use of their personal skin care products (brown circles), all volunteers used only the same head to toe shampoo during the first 3 weeks (week 1–week 3) and no other beauty product was applied (solid blue circle). The following 3 weeks (week 4–week 6), four selected commercial beauty products were applied daily by all volunteers on the specific body part (deodorant antiperspirant for the armpits, soothing foot powder for the feet between toes, sunscreen for the face, and moisturizer for the front forearm) (triangles) and continued to use the same shampoo. During the last 3 weeks (week 7–week 9), all volunteers went back to their normal routine and used their personal beauty products (circles). Samples were collected once a week (from day 0 to day 68—10 timepoints from T0 to T9) for volunteers 1, 2, 3, 4, 5, 6, 7, 9, 10, 11, and 12, and on day 0 and day 6 for volunteer 8, who withdraw from the study after day 6. For 3 individuals (volunteers 4, 9, 10), samples were collected twice a week (19 timepoints total). Samples collected for 11 volunteers during 10 timepoints: 11 volunteers × 10 timepoints × 4 samples × 4 body sites = 1760. Samples collected from 3 selected volunteers during 9 additional timepoints: 3 volunteers × 9 timepoints × 4 samples × 4 body sites = 432. See also the “ Subject recruitment and sample collection ” section in the “ Methods ” section
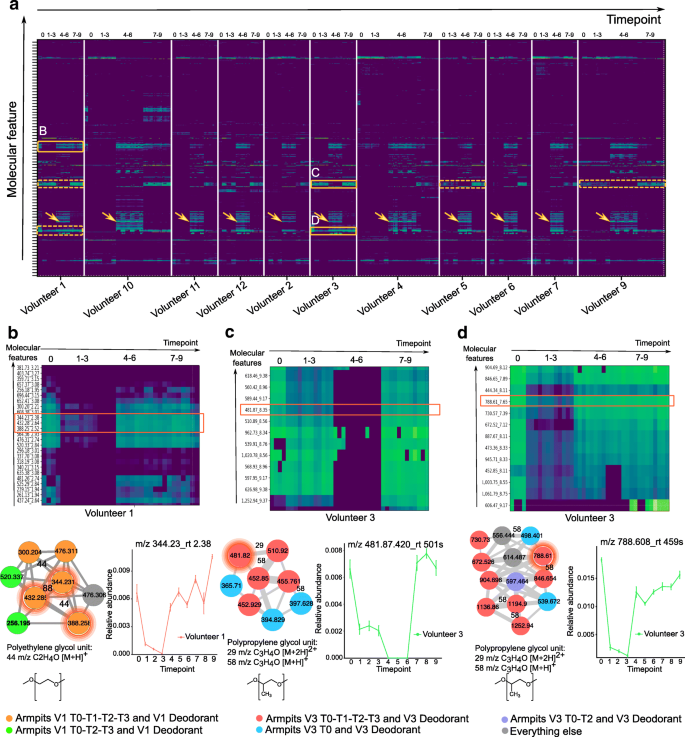
Monitoring the persistence of personal care product ingredients in the armpits over a 9-week period. a Heatmap representation of the most abundant molecular features detected in the armpits of all individuals during the four phases (0: initial, 1–3: no beauty products, 4–6: common products, and 7–9: personal products). Green color in the heatmap represents the highest molecular abundance and blue color the lowest one. Orange boxes with plain lines represent enlargement of cluster of molecules that persist on the armpits of volunteer 1 ( b ) and volunteer 3 ( c , d ). Orange clusters with dotted lines represent same clusters of molecules found on the armpits of other volunteers. Orange arrows represent the cluster of compounds characteristic of the antiperspirant used during T4–T6. b Polyethylene glycol (PEG) molecular clusters that persist on the armpits of individual 1. The molecular subnetwork, representing molecular families [ 24 ], is part of a molecular network ( http://gnps.ucsd.edu/ProteoSAFe/status.jsp?task=f5325c3b278a46b29e8860ec5791d5ad ) generated from MS/MS data collected from the armpits of volunteer 1 (T0–T3) MSV000081582 and MS/MS data collected from the deodorant used by volunteer 1 before the study started (T0) MSV000081580. c , d Polypropylene glycol (PPG) molecular families that persist on the armpits of individual 3, along with the corresponding molecular subnetwork that is part of the molecular network accessible here http://gnps.ucsd.edu/ProteoSAFe/status.jsp?task=aaa1af68099d4c1a87e9a09f398fe253 . Subnetworks were generated from MS/MS data collected from the armpits of volunteer 3 (T0–T3) MSV000081582 and MS/MS data collected from the deodorant used by volunteer 3 at T0 MSV000081580. The network nodes were annotated with colors. Nodes represent MS/MS spectra found in armpit samples of individual 1 collected during T0, T1, T2, and T3 and in personal deodorant used by individual 1 (orange nodes); armpit samples of individual 1 collected during T0, T2, and T3 and personal deodorant used by individual 1 (green nodes); armpit samples of individual 3 collected during T0, T1, T2, and T3 and in personal deodorant used by individual 3 (red nodes); armpit samples of individual 3 collected during T0 and in personal deodorant used by individual 3 (blue nodes); and armpit samples of individual 3 collected during T0 and T2 and in personal deodorant used by individual 3 (purple nodes). Gray nodes represent everything else. Error bars represent standard error of the mean calculated at each timepoint from four armpit samples collected from the right and left side of each individual separately. See also Additional file 1 : Figure S1
To understand how long beauty products persist on the skin, we monitored compounds found in deodorants used by two volunteers—female 1 and female 3—before the study (T0), over the first 3 weeks (T1–T3) (Fig. 1 b). During this phase, all participants used exclusively the same body wash during showering, making it easier to track ingredients of their personal care products. The data in the first 3 weeks (T1–T3) revealed that many ingredients of deodorants used on armpits (Fig. 2 a) persist on the skin during this time and were still detected during the first 3 weeks or at least during the first week following the last day of use. Each of the compounds detected in the armpits of individuals exhibited its own unique half-life. For example, the polyethylene glycol (PEG)-derived compounds m/z 344.227, rt 143 s (Fig. 2 b, S1J); m/z 432.279, rt 158 s (Fig. 2 b, S1K); and m/z 388.253, rt 151 s (Fig. 2 b, S1L) detected on armpits of volunteer 1 have a calculated half-life of 0.5 weeks (Additional file 1 : Figure S1J-L, all p values < 1.81e−07), while polypropylene glycol (PPG)-derived molecules m/z 481.87, rt 501 s (Fig. 2 c, S1M); m/z 560.420, rt 538 s (Fig. 2 c, S1N); m/z 788.608, rt 459 s (Fig. 2 d, S1O); m/z 846.650, rt 473 s (Fig. 2 d, S1P); and m/z 444.338, rt 486 s (Fig. 2 d, S1Q) found on armpits of volunteers 3 and 1 (Fig. 2 a) have a calculated half-life ranging from 0.7 to 1.9 weeks (Additional file 1 : Figure S1M-Q, all p values < 0.02), even though they originate from the same deodorant used by each individual. For some ingredients of deodorant used by volunteer 3 on time 0 (Additional file 1 : Figure S1M, N), a decline was observed during the first week, then little to no traces of these ingredients were detected during weeks 4–6 (T4–T6), then finally these ingredients reappear again during the last 3 weeks of personal product use (T7–T9). This suggests that these ingredients are present exclusively in the personal deodorant used by volunteer 3 before the study. Because a similar deodorant (Additional file 1 : Figure S1O-Q) and a face lotion (Additional file 1 : Figure S1R) was used by volunteer 3 and volunteer 2, respectively, prior to the study, there was no decline or absence of their ingredients during weeks 4–6 (T4–T6).
Polyethylene glycol compounds (Additional file 1 : Figure S1J-L) wash out faster from the skin than polypropylene glycol (Additional file 1 : Figure S1M-Q)(HL ~ 0.5 weeks vs ~ 1.9 weeks) and faster than fatty acids used in lotions (HL ~ 1.2 weeks) (Additional file 1 : Figure S1R), consistent with their hydrophilic (PEG) and hydrophobic properties (PPG and fatty acids) [ 25 , 26 ]. This difference in hydrophobicity is also reflected in the retention time as detected by mass spectrometry. Following the linear decrease of two PPG compounds from T0 to T1, they accumulated noticeably during weeks 2 and 3 (Additional file 1 : Figure S1M, N). This accumulation might be due to other sources of PPG such as the body wash used during this period or the clothes worn by person 3. Although PPG compounds were not listed in the ingredient list of the shampoo, we manually inspected the LC-MS data collected from this product and confirmed the absence of PPG compounds in the shampoo. The data suggest that this trend is characteristic of accumulation of PPG from additional sources. These could be clothes, beds, or sheets, in agreement with the observation of these molecules found in human habitats [ 27 ] but also in the public GNPS mass spectrometry dataset MSV000079274 that investigated the chemicals from dust collected from 1053 mattresses of children.
Temporal molecular and bacterial diversity in response to personal care use
To assess the effect of discontinuing and resuming the use of skin care products on molecular and microbiota dynamics, we first evaluated their temporal diversity. Skin sites varied markedly in their initial level (T0) of molecular and bacterial diversity, with higher molecular diversity at all sites for female participants compared to males (Fig. 3 a, b, Wilcoxon rank-sum-WR test, p values ranging from 0.01 to 0.0001, from foot to arm) and higher bacterial diversity in face (WR test, p = 0.0009) and armpits (WR test, p = 0.002) for females (Fig. 3 c, d). Temporal diversity was similar across the right and left sides of each body site of all individuals (WR test, molecular diversity: all p values > 0.05; bacterial diversity: all p values > 0.20). The data show that refraining from using beauty products (T1–T3) leads to a significant decrease in molecular diversity at all sites (Fig. 3 a, b, WR test, face: p = 8.29e−07, arm: p = 7.08e−09, armpit: p = 1.13e−05, foot: p = 0.002) and bacterial diversity mainly in armpits (WR test, p = 0.03) and feet (WR test, p = 0.04) (Fig. 3 c, d). While molecular diversity declined (Fig. 3 a, b) for arms and face, bacterial diversity (Fig. 3 c, d) was less affected in the face and arms when participants did not use skin care products (T1–T3). The molecular diversity remained stable in the arms and face of female participants during common beauty products use (T4–T6) to immediately increase as soon as the volunteers went back to their normal routines (T7–T9) (WR test, p = 0.006 for the arms and face)(Fig. 3 a, b). A higher molecular (Additional file 1 : Figure S2A) and community (Additional file 1 : Figure S2B) diversity was observed for armpits and feet of all individuals during the use of antiperspirant and foot powder (T4–T6) (WR test, molecular diversity: armpit p = 8.9e−33, foot p = 1.03e−11; bacterial diversity: armpit p = 2.14e−28, foot p = 1.26e−11), followed by a molecular and bacterial diversity decrease in the armpits when their regular personal beauty product use was resumed (T7–T9) (bacterial diversity: WR test, p = 4.780e−21, molecular diversity: WR test, p = 2.159e−21). Overall, our data show that refraining from using beauty products leads to lower molecular and bacterial diversity, while resuming the use increases their diversity. Distinct variations between male and female molecular and community richness were perceived at distinct body parts (Fig. 3 a–d). Although the chemical diversity of personal beauty products does not explain these variations (Additional file 1 : Figure S2C), differences observed between males and females may be attributed to many environmental and lifestyle factors including different original skin care and different frequency of use of beauty products (Additional file 2 : Table S1), washing routines, and diet.
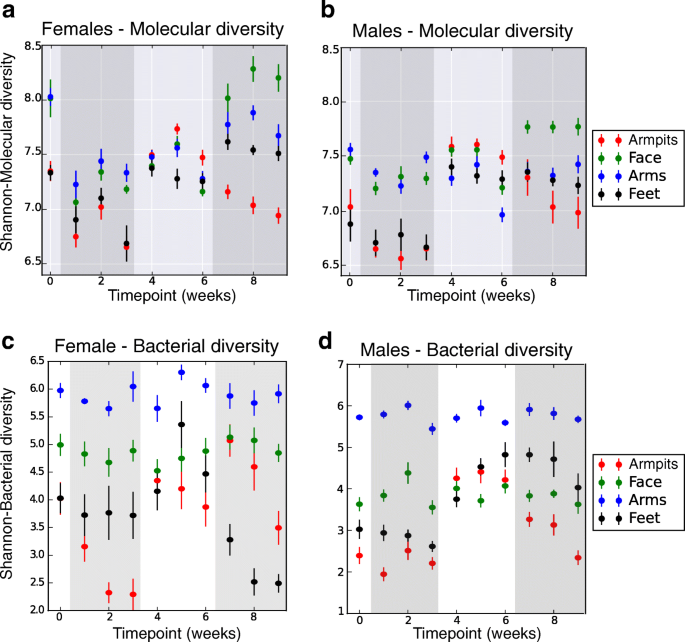
Molecular and bacterial diversity over a 9-week period, comparing samples based on their molecular (UPLC-Q-TOF-MS) or bacterial (16S rRNA amplicon) profiles. Molecular and bacterial diversity using the Shannon index was calculated from samples collected from each body part at each timepoint, separately for female ( n = 5) and male ( n = 6) individuals. Error bars represent standard error of the mean calculated at each timepoint, from up to four samples collected from the right and left side of each body part, of females ( n = 5) and males ( n = 6) separately. a , b Molecular alpha diversity measured using the Shannon index from five females (left panel) and six males (right panel), over 9 weeks, from four distinct body parts (armpits, face, arms, feet). c , d Bacterial alpha diversity measured using the Shannon index, from skin samples collected from five female (left panel) and six male individuals (right panel), over 9 weeks, from four distinct body parts (armpits, face, arms, feet). See also Additional file 1 : Figure S2
Longitudinal variation of skin metabolomics signatures
To gain insights into temporal metabolomics variation associated with beauty product use, chemical inventories collected over 9 weeks were subjected to multivariate analysis using the widely used Bray–Curtis dissimilarity metric (Fig. 4 a–c, S3A). Throughout the 9-week period, distinct molecular signatures were associated to each specific body site: arm, armpit, face, and foot (Additional file 1 : Figure S3A, Adonis test, p < 0.001, R 2 0.12391). Mass spectrometric signatures displayed distinct individual trends at each specific body site (arm, armpit, face, and foot) over time, supported by their distinct locations in PCoA (principal coordinate analysis) space (Fig. 4 a, b) and based on the Bray–Curtis distances between molecular profiles (Additional file 1 : Figure S3B, WR test, all p values < 0.0001 from T0 through T9). This suggests a high molecular inter-individual variability over time despite similar changes in personal care routines. Significant differences in molecular patterns associated to ceasing (T1–T3) (Fig. 4 b, Additional file 1 : Figure S3C, WR test, T0 vs T1–T3 p < 0.001) and resuming the use of common beauty products (T4–T6) (Additional file 1 : Figure S3C) were observed in the arm, face, and foot (Fig. 4 b), although the armpit exhibited the most pronounced changes (Fig. 4 b, Additional file 1 : Figure S3D, E, random forest highlighting that 100% of samples from each phase were correctly predicted). Therefore, we focused our analysis on this region. Molecular changes were noticeable starting the first week (T1) of discontinuing beauty product use. As shown for armpits in Fig. 4 c, these changes at the chemical level are specific to each individual, possibly due to the extremely personalized lifestyles before the study and match their original use of deodorant. Based on the initial use of underarm products (T0) (Additional file 2 : Table S1), two groups of participants can be distinguished: a group of five volunteers who used stick deodorant as evidenced by the mass spectrometry data and another group of volunteers where we found few or no traces suggesting they never or infrequently used stick deodorants (Additional file 2 : Table S1). Based on this criterion, the chemical trends shown in Fig. 4 c highlight that individuals who used stick deodorant before the beginning of the study (volunteers 1, 2, 3, 9, and 12) displayed a more pronounced shift in their armpits’ chemistries as soon as they stopped using deodorant (T1–T3), compared to individuals who had low detectable levels of stick deodorant use (volunteers 4, 6, 7, and 10), or “rarely-to-never” (volunteers 5 and 11) use stick deodorants as confirmed by the volunteers (Additional file 1 : Figure S3F, WR test, T0 vs T1–T3 all p values < 0.0001, with greater distance for the group of volunteers 1, 2, 3, 9, and 12, compared to volunteers 4, 5, 6, 7, 10, and 11). The most drastic shift in chemical profiles was observed during the transition period, when all participants applied the common antiperspirant on a daily basis (T4–T6) (Additional file 1 : Figure S3D, E). Finally, the molecular profiles became gradually more similar to those collected before the experiment (T0) as soon as the participants resumed using their personal beauty products (T7–T9) (Additional file 1 : Figure S3C), although traces of skin care products did last through the entire T7–T9 period in people who do not routinely apply these products (Fig. 4 c).
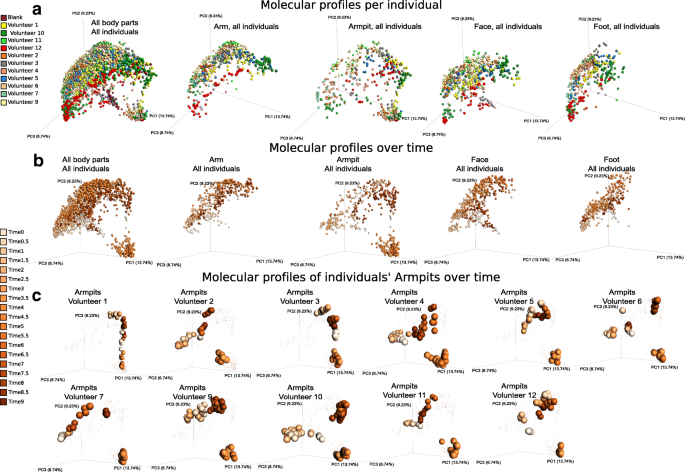
Individualized influence of beauty product application on skin metabolomics profiles over time. a Multivariate statistical analysis (principal coordinate analysis (PCoA)) comparing mass spectrometry data collected over 9 weeks from the skin of 11 individuals, all body parts, combined (first plot from the left) and then displayed separately (arm, armpits, face, feet). Color scale represents volunteer ID. The PCoA was calculated on all samples together, and subsets of the data are shown in this shared space and the other panels. b The molecular profiles collected over 9 weeks from all body parts, combined then separately (arm, armpits, face, feet). c Representative molecular profiles collected over 9 weeks from armpits of 11 individuals (volunteers 1, 2, 3, 4, 5, 6, 7, 9, 10, 11, 12). Color gradient in b and c represents timepoints (time 0 to time 9), ranging from the lightest orange color to the darkest one that represent the earliest (time 0) to the latest (time 9) timepoint, respectively. 0.5 timepoints represent additional timepoints where three selected volunteers were samples (volunteers 4, 9, and 10). PCoA plots were generated using the Bray–Curtis dissimilarity matrix and visualized in Emperor [ 28 ]. See also Additional file 1 : Figure S3
Comparing chemistries detected in armpits at the end timepoints—when no products were used (T3) and during product use (T6)—revealed distinct molecular signatures characteristic of each phase (random forest highlighting that 100% of samples from each group were correctly predicted, see Additional file 1 : Figure S3D, E). Because volunteers used the same antiperspirant during T4–T6, molecular profiles converged during that time despite individual patterns at T3 (Fig. 4 b, c, Additional file 1 : Figure S3D). These distinct chemical patterns reflect the significant impact of beauty products on skin molecular composition. Although these differences may in part be driven by beauty product ingredients detected on the skin (Additional file 1 : Figure S1), we anticipated that additional host- and microbe-derived molecules may also be involved in these molecular changes.
To characterize the chemistries that vary over time, we used molecular networking, a MS visualization approach that evaluates the relationship between MS/MS spectra and compares them to reference MS/MS spectral libraries of known compounds [ 29 , 30 ]. We recently showed that molecular networking can successfully organize large-scale mass spectrometry data collected from the human skin surface [ 18 , 19 ]. Briefly, molecular networking uses the MScluster algorithm [ 31 ] to merge all identical spectra and then compares and aligns all unique pairs of MS/MS spectra based on their similarities where 1.0 indicates a perfect match. Similarities between MS/MS spectra are calculated using a similarity score, and are interpreted as molecular families [ 19 , 24 , 32 , 33 , 34 ]. Here, we used this method to compare and characterize chemistries found in armpits, arms, face, and foot of 11 participants. Based on MS/MS spectral similarities, chemistries highlighted through molecular networking (Additional file 1 : Figure S4A) were associated with each body region with 8% of spectra found exclusively in the arms, 12% in the face, 14% in the armpits, and 2% in the foot, while 18% of the nodes were shared between all four body parts and the rest of spectra were shared between two body sites or more (Additional file 1 : Figure S4B). Greater spectral similarities were highlighted between armpits, face, and arm (12%) followed by the arm and face (9%) (Additional file 1 : Figure S4B).
Molecules were annotated with Global Natural Products Social Molecular Networking (GNPS) libraries [ 29 ], using accurate parent mass and MS/MS fragmentation patterns, according to level 2 or 3 of annotation defined by the 2007 metabolomics standards initiative [ 35 ]. Through annotations, molecular networking revealed that many compounds derived from steroids (Fig. 5 a–d), bile acids (Additional file 1 : Figure S5A-D), and acylcarnitines (Additional file 1 : Figure S5E-F) were exclusively detected in the armpits. Using authentic standards, the identity of some pheromones and bile acids were validated to a level 1 identification with matched retention times (Additional file 1 : Figure S6B, S7A, C, D). Other steroids and bile acids were either annotated using standards with identical MS/MS spectra but slightly different retention times (Additional file 1 : Figure S6A) or annotated with MS/MS spectra match with reference MS/MS library spectra (Additional file 1 : Figure S6C, D, S7B, S6E-G). These compounds were therefore classified as level 3 [ 35 ]. Acylcarnitines were annotated to a family of possible acylcarnitines (we therefore classify as level 3), as the positions of double bonds or cis vs trans configurations are unknown (Additional file 1 : Figure S8A, B).
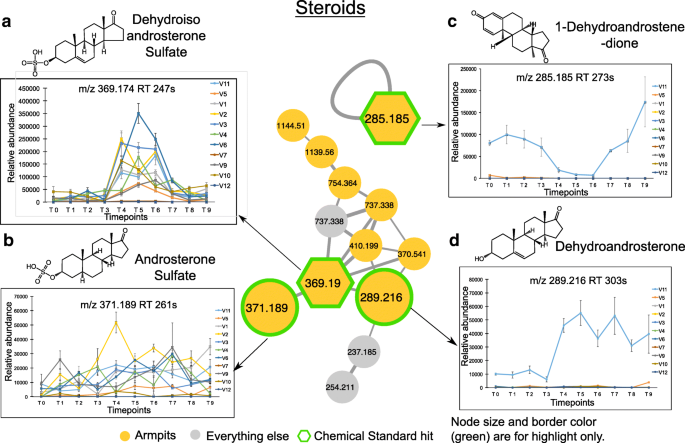
Underarm steroids and their longitudinal abundance. a – d Steroid molecular families in the armpits and their relative abundance over a 9-week period. Molecular networking was applied to characterize chemistries from the skin of 11 healthy individuals. The full network is shown in Additional file 1 : Figure S4A, and networking parameters can be found here http://gnps.ucsd.edu/ProteoSAFe/status.jsp?task=284fc383e4c44c4db48912f01905f9c5 for MS/MS datasets MSV000081582. Each node represents a consensus of a minimum of 3 identical MS/MS spectra. Yellow nodes represent MS/MS spectra detected in armpits samples. Hexagonal shape represents MS/MS spectra match between skin samples and chemical standards. Plots are representative of the relative abundance of each compound over time, calculated separately from LC-MS1 data collected from the armpits of each individual. Steroids detected in armpits are a , dehydroisoandrosterone sulfate ( m/z 369.190, rt 247 s), b androsterone sulfate ( m/z 371.189, rt 261 s), c 1-dehydroandrostenedione ( m/z 285.185, rt 273 s), and d dehydroandrosterone ( m/z 289.216, rt 303 s). Relative abundance over time of each steroid compound is represented. Error bars represent the standard error of the mean calculated at each timepoint from four armpit samples from the right and left side of each individual separately. See also Additional file 1 : Figures S4-S8
Among the steroid compounds, several molecular families were characterized: androsterone (Fig. 5 a, b, d), androstadienedione (Fig. 5 c), androstanedione (Additional file 1 : Figure S6E), androstanolone (Additional file 1 : Figure S6F), and androstenedione (Additional file 1 : Figure S6G). While some steroids were detected in the armpits of several individuals, such as dehydroisoandrosterone sulfate ( m/z 369.19, rt 247 s) (9 individuals) (Fig. 5 a, Additional file 1 : Figure S6A), androsterone sulfate ( m/z 371.189, rt 261 s) (9 individuals) (Fig. 5 b, Additional file 1 : Figure S6C), and 5-alpha-androstane-3,17-dione ( m/z 271.205, rt 249 s) (9 individuals) (Additional file 1 : Figure S6E), other steroids including 1-dehydroandrostenedione ( m/z 285.185, rt 273 s) (Fig. 5 c, Additional file 1 : Figure S6B), dehydroandrosterone ( m/z 289.216, rt 303 s) (Fig. 5 d, Additional file 1 : Figure S6D), and 5-alpha-androstan-17.beta-ol-3-one ( m/z 291.231, rt 318 s) (Additional file 1 : Figure S6F) were only found in the armpits of volunteer 11 and 4-androstene-3,17-dione ( m/z 287.200, rt 293 s) in the armpits of volunteer 11 and volunteer 5, both are male that never applied stick deodorants (Additional file 1 : Figure S6G). Each molecular species exhibited a unique pattern over the 9-week period. The abundance of dehydroisoandrosterone sulfate (Fig. 5 a, WR test, p < 0.01 for 7 individuals) and dehydroandrosterone (Fig. 5 a, WR test, p = 0.00025) significantly increased during the use of antiperspirant (T4–T6), while androsterone sulfate (Fig. 5 b) and 5-alpha-androstane-3,17-dione (Additional file 1 : Figure S6E) display little variation over time. Unlike dehydroisoandrosterone sulfate (Fig. 5 a) and dehydroandrosterone (Fig. 5 d), steroids including 1-dehydroandrostenedione (Fig. 5 c, WR test, p = 0.00024) and 4-androstene-3,17-dione (Additional file 1 : Figure S6G, WR test, p = 0.00012) decreased in abundance during the 3 weeks of antiperspirant application (T4–T6) in armpits of male 11, and their abundance increased again when resuming the use of his normal skin care routines (T7–T9). Interestingly, even within the same individual 11, steroids were differently impacted by antiperspirant use as seen for 1-dehydroandrostenedione that decreased in abundance during T4–T6 (Fig. 5 c, WR test, p = 0.00024), while dehydroandrosterone increased in abundance (Fig. 5 d, WR test, p = 0.00025), and this increase was maintained during the last 3 weeks of the study (T7–T9).
In addition to steroids, many bile acids (Additional file 1 : Figure S5A-D) and acylcarnitines (Additional file 1 : Figure S5E-F) were detected on the skin of several individuals through the 9-week period. Unlike taurocholic acid found only on the face (Additional file 1 : Figures S5A, S7A) and tauroursodeoxycholic acid detected in both armpits and arm samples (Additional file 1 : Figures S5B, S7B), other primary bile acids such as glycocholic (Additional file 1 : Figures S5C, S7C) and chenodeoxyglycocholic acid (Additional file 1 : Figures S5D, S7D) were exclusively detected in the armpits. Similarly, acylcarnitines were also found either exclusively in the armpits (hexadecanoyl carnitines) (Additional file 1 : Figures S5E, S8A) or in the armpits and face (tetradecenoyl carnitine) (Additional file 1 : Figures S5F, S8B) and, just like the bile acids, they were also stably detected during the whole 9-week period.
Bacterial communities and their variation over time
Having demonstrated the impact of beauty products on the chemical makeup of the skin, we next tested the extent to which skin microbes are affected by personal care products. We assessed temporal variation of bacterial communities detected on the skin of healthy individuals by evaluating dissimilarities of bacterial collections over time using unweighted UniFrac distance [ 36 ] and community variation at each body site in association to beauty product use [ 3 , 15 , 37 ]. Unweighted metrics are used for beta diversity calculations because we are primarily concerned with changes in community membership rather than relative abundance. The reason for this is that skin microbiomes can fluctuate dramatically in relative abundance on shorter timescales than that assessed here. Longitudinal variations were revealed for the armpits (Fig. 6 a) and feet microbiome by their overall trend in the PCoA plots (Fig. 6 b), while the arm (Fig. 6 c) and face (Fig. 6 d) displayed relatively stable bacterial profiles over time. As shown in Fig. 6 a–d, although the microbiome was site-specific, it varied more between individuals and this inter-individual variability was maintained over time despite same changes in personal care routine (WR test, all p values at all timepoints < 0.05, T5 p = 0.07), in agreement with previous findings that individual differences in the microbiome are large and stable over time [ 3 , 4 , 10 , 37 ]. However, we show that shifts in the microbiome can be induced by changing hygiene routine and therefore skin chemistry. Changes associated with using beauty products (T4–T6) were more pronounced for the armpits (Fig. 6 a, WR test, p = 1.61e−52) and feet (Fig. 6 b, WR test, p = 6.15e−09), while little variations were observed for the face (Fig. 6 d, WR test, p = 1.402.e−83) and none for the arms (Fig. 6 c, WR test, p = 0.296).
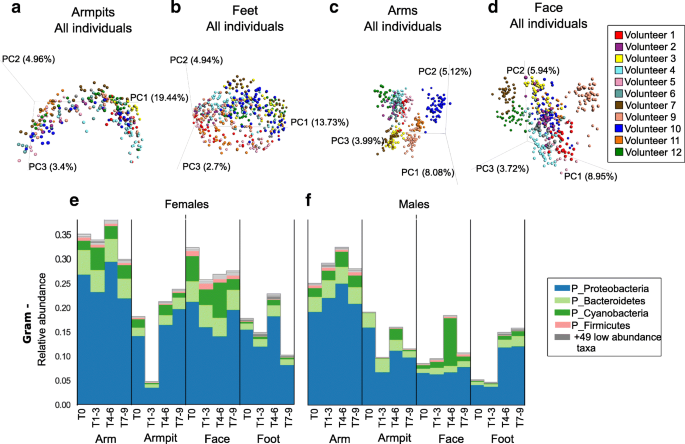
Longitudinal variation of skin bacterial communities in association with beauty product use. a - d Bacterial profiles collected from skin samples of 11 individuals, over 9 weeks, from four distinct body parts a) armpits, b) feet, c) arms and d) face, using multivariate statistical analysis (Principal Coordinates Analysis PCoA) and unweighted Unifrac metric. Each color represents bacterial samples collected from an individual. PCoA were calculated separately for each body part. e , f Representative Gram-negative (Gram -) bacteria collected from arms, armpits, face and feet of e) female and f) male participants. See also Additional file 1 : Figure S9A, B showing Gram-negative bacterial communities represented at the genus level
A significant increase in abundance of Gram-negative bacteria including the phyla Proteobacteria and Bacteroidetes was noticeable for the armpits and feet of both females (Fig. 6 e; Mann–Whitney U , p = 8.458e−07) and males (Fig. 6 f; Mann–Whitney U , p = 0.0004) during the use of antiperspirant (T4–T6), while their abundance remained stable for the arms and face during that time (Fig. 6 e, f; female arm p = 0.231; female face p value = 0.475; male arm p = 0.523;male face p = 6.848751e−07). These Gram-negative bacteria include Acinetobacter and Paracoccus genera that increased in abundance in both armpits and feet of females (Additional file 1 : Figure S9A), while a decrease in abundance of Enhydrobacter was observed in the armpits of males (Additional file 1 : Figure S9B). Cyanobacteria, potentially originating from plant material (Additional file 1 : Figure S9C) also increased during beauty product use (T4–T6) especially in males, in the armpits and face of females (Fig. 6 e) and males (Fig. 6 f). Interestingly, although chloroplast sequences (which group phylogenetically within the cyanobacteria [ 38 ]) were only found in the facial cream (Additional file 1 : Figure S9D), they were detected in other locations as well (Fig. 6 e, f. S9E, F), highlighting that the application of a product in one region will likely affect other regions of the body. For example, when showering, a face lotion will drip down along the body and may be detected on the feet. Indeed, not only did the plant material from the cream reveal this but also the shampoo used for the study for which molecular signatures were readily detected on the feet as well (Additional file 1 : Figure S10A). Minimal average changes were observed for Gram-positive organisms (Additional file 1 : Figure S10B, C), although in some individuals the variation was greater than others (Additional file 1 : Figure S10D, E) as discussed for specific Gram-positive taxa below.
At T0, the armpit’s microflora was dominated by Staphylococcus (26.24%, 25.11% of sequencing reads for females and 27.36% for males) and Corynebacterium genera (26.06%, 17.89% for females and 34.22% for males) (Fig. 7 a—first plot from left and Additional file 1 : Figure S10D, E). They are generally known as the dominant armpit microbiota and make up to 80% of the armpit microbiome [ 39 , 40 ]. When no deodorants were used (T1–T3), an overall increase in relative abundance of Staphylococcus (37.71%, 46.78% for females and 30.47% for males) and Corynebacterium (31.88%, 16.50% for females and 44.15% for males) genera was noticeable (WR test, p < 3.071e−05) (Fig. 7 a—first plot from left), while the genera Anaerococcus and Peptoniphilus decreased in relative abundance (WR test, p < 0.03644) (Fig. 7 a—first plot from left and Additional file 1 : Figure S10D, E). When volunteers started using antiperspirants (T4–T6), the relative abundance of Staphylococcus (37.71%, 46.78% females and 30.47% males, to 21.71%, 25.02% females and 19.25% males) and Corynebacterium (31.88%, 16.50% females and 44.15% males, to 15.83%, 10.76% females and 19.60% males) decreased (WR test, p < 3.071e−05) (Fig. 7 a, Additional file 1 : Figure S10D, E) and at the same time, the overall alpha diversity increased significantly (WR test, p = 3.47e−11) (Fig. 3 c, d). The microbiota Anaerococcus (WR test, p = 0.0006018) , Peptoniphilus (WR test, p = 0.008639), and Micrococcus (WR test, p = 0.0377) increased significantly in relative abundance, together with a lot of additional low-abundant species that lead to an increase in Shannon alpha diversity (Fig. 3 c, d). When participants went back to normal personal care products (T7–T9), the underarm microbiome resembled the original underarm community of T0 (WR test, p = 0.7274) (Fig. 7 a). Because armpit bacterial communities are person-specific (inter-individual variability: WR test, all p values at all timepoints < 0.05, besides T5 p n.s), variation in bacterial abundance upon antiperspirant use (T4–T6) differ between individuals and during the whole 9-week period (Fig. 7a —taxonomic plots per individual). For example, the underarm microbiome of male 5 exhibited a unique pattern, where Corynebacterium abundance decreased drastically during the use of antiperspirant (82.74 to 11.71%, WR test, p = 3.518e−05) while in the armpits of female 9 a huge decrease in Staphylococcus abundance was observed (Fig. 7 a) (65.19 to 14.85%, WR test, p = 0.000113). Unlike other participants, during T0–T3, the armpits of individual 11 were uniquely characterized by the dominance of a sequence that matched most closely to the Enhydrobacter genera . The transition to antiperspirant use (T4–T6) induces the absence of Enhydrobacter (30.77 to 0.48%, WR test, p = 0.01528) along with an increase of Corynebacterium abundance (26.87 to 49.74%, WR test, p = 0.1123) (Fig. 7 a—male 11).
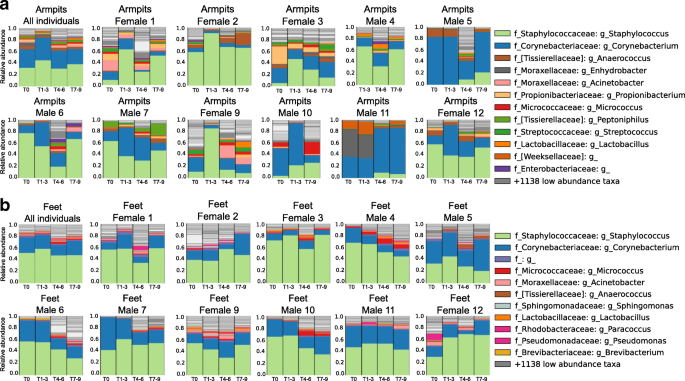
Person-to-person bacterial variabilities over time in the armpits and feet. a Armpit microbiome changes when stopping personal care product use, then resuming. Armpit bacterial composition of the 11 volunteers combined, then separately, (female 1, female 2, female 3, male 4, male 5, male 6, male 7, female 9, male 10, male 11, female 12) according to the four periods within the experiment. b Feet bacterial variation over time of the 12 volunteers combined, then separately (female 1, female 2, female 3, male 4, male 5, male 6, male 7, female 9, male 10, male 11, female 12) according to the four periods within the experiment. See also Additional file 1 : Figure S9-S13
In addition to the armpits, a decline in abundance of Staphylococcus and Corynebacterium was perceived during the use of the foot powder (46.93% and 17.36%, respectively) compared to when no beauty product was used (58.35% and 22.99%, respectively) (WR test, p = 9.653e−06 and p = 0.02032, respectively), while the abundance of low-abundant foot bacteria significantly increased such as Micrococcus (WR test, p = 1.552e−08), Anaerococcus (WR test, p = 3.522e−13), Streptococcus (WR test, p = 1.463e−06), Brevibacterium (WR test, p = 6.561e−05), Moraxellaceae (WR test, p = 0.0006719), and Acinetobacter (WR test, p = 0.001487), leading to a greater bacterial diversity compared to other phases of the study (Fig. 7 b first plot from left, Additional file 1 : Figure S10D, E, Fig. 3 c, d).
We further evaluated the relationship between the two omics datasets by superimposing the principal coordinates calculated from metabolome and microbiome data (Procrustes analysis) (Additional file 1 : Figure S11) [ 34 , 41 , 42 ]. Metabolomics data were more correlated with patterns observed in microbiome data in individual 3 (Additional file 1 : Figure S11C, Mantel test, r = 0.23, p < 0.001), individual 5 (Additional file 1 : Figure S11E, r = 0.42, p < 0.001), individual 9 (Additional file 1 : Figure S11H, r = 0.24, p < 0.001), individual 10 (Additional file 1 : Figure S11I, r = 0.38, p < 0.001), and individual 11 (Additional file 1 : Figure S11J, r = 0.35, p < 0.001) when compared to other individuals 1, 2, 4, 6, 7, and 12 (Additional file 1 : Figure S11A, B, D, F, G, K, respectively) (Mantel test, all r < 0.2, all p values < 0.002, for volunteer 2 p n.s). Furthermore, these correlations were individually affected by ceasing (T1–T3) or resuming the use of beauty products (T4–T6 and T7–T9) (Additional file 1 : Figure S11A-K).
Overall, metabolomics–microbiome correlations were consistent over time for the arms, face, and feet although alterations were observed in the arms of volunteers 7 (Additional file 1 : Figure S11G) and 10 (Additional file 1 : Figure S11I) and the face of volunteer 7 (Additional file 1 : Figure S11G) during product use (T4–T6). Molecular–bacterial correlations were mostly affected in the armpits during antiperspirant use (T4–T6), as seen for volunteers male 7 (Additional file 1 : Figure S11G) and 11 (Additional file 1 : Figure S11J) and females 2 (Additional file 1 : Figure S11B), 9 (Additional file 1 : Figure S11H), and 12 (Additional file 1 : Figure S11K). This perturbation either persisted during the last 3 weeks (Additional file 1 : Figure S11D, E, H, I, K) when individuals went back to their normal routine (T7–T9) or resembled the initial molecular–microbial correlation observed in T0 (Additional file 1 : Figure S11C, G, J). These alterations in molecular–bacterial correlation are driven by metabolomics changes during antiperspirant use as revealed by metabolomics shifts on the PCoA space (Additional file 1 : Figure S11), partially due to the deodorant’s chemicals (Additional file 1 : Figure S1J, K) but also to changes observed in steroid levels in the armpits (Fig. 5A, C, D , Additional file 1 : Figure S6G), suggesting metabolome-dependant changes of the skin microbiome. In agreement with previous findings that showed efficient biotransformation of steroids by Corynebacterium [ 43 , 44 ], our correlation analysis associates specific steroids that were affected by antiperspirant use in the armpits of volunteer 11 (Fig. 5 c, d, Additional file 1 : Figure S6G) with microbes that may produce or process them: 1-dehydroandrostenedione, androstenedione, and dehydrosterone with Corynebacterium ( r = − 0.674, p = 6e−05; r = 0.671, p = 7e−05; r = 0.834, p < 1e−05, respectively) (Additional file 1 : Figure S12A, B, C, respectively) and Enhydrobacter ( r = 0.683, p = 4e−05; r = 0.581, p = 0.00095; r = 0.755, p < 1e−05 respectively) (Additional file 1 : Figure S12D, E, F, respectively).
Despite the widespread use of skin care and hygiene products, their impact on the molecular and microbial composition of the skin is poorly studied. We established a workflow that examines individuals to systematically study the impact of such lifestyle characteristics on the skin by taking a broad look at temporal molecular and bacterial inventories and linking them to personal skin care product use. Our study reveals that when the hygiene routine is modified, the skin metabolome and microbiome can be altered, but that this alteration depends on product use and location on the body. We also show that like gut microbiome responses to dietary changes [ 20 , 21 ], the responses are individual-specific.
We recently reported that traces of our lifestyle molecules can be detected on the skin days and months after the original application [ 18 , 19 ]. Here, we show that many of the molecules associated with our personal skin and hygiene products had a half-life of 0.5 to 1.9 weeks even though the volunteers regularly showered, swam, or spent time in the ocean. Thus, a single application of some of these products has the potential to alter the microbiome and skin chemistry for extensive periods of time. Our data suggests that although host genetics and diet may play a role, a significant part of the resilience of the microbiome that has been reported [ 10 , 45 ] is due to the resilience of the skin chemistry associated with personal skin and hygiene routines, or perhaps even continuous re-exposure to chemicals from our personal care routines that are found on mattresses, furniture, and other personal objects [ 19 , 27 , 46 ] that are in constant contact. Consistent with this observation is that individuals in tribal regions and remote villages that are infrequently exposed to the types of products used in this study have very different skin microbial communities [ 47 , 48 ] and that the individuals in this study who rarely apply personal care products had a different starting metabolome. We observed that both the microbiome and skin chemistry of these individuals were most significantly affected by these products. This effect by the use of products at T4–T6 on the volunteers that infrequently used them lasted to the end phase of the study even though they went back to infrequent use of personal care products. What was notable and opposite to what the authors originally hypothesized is that the use of the foot powder and antiperspirant increased the diversity of microbes and that some of this diversity continued in the T7–T9 phase when people went back to their normal skin and hygiene routines. It is likely that this is due to the alteration in the nutrient availability such as fatty acids and moisture requirements, or alteration of microbes that control the colonization via secreted small molecules, including antibiotics made by microbes commonly found on the skin [ 49 , 50 ].
We detected specific molecules on the skin that originated from personal care products or from the host. One ingredient that lasts on the skin is propylene glycol, which is commonly used in deodorants and antiperspirants and added in relatively large amounts as a humectant to create a soft and sleek consistency [ 51 ]. As shown, daily use of personal care products is leading to high levels of exposure to these polymers. Such polymers cause contact dermatitis in a subset of the population [ 51 , 52 ]. Our data reveal a lasting accumulation of these compounds on the skin, suggesting that it may be possible to reduce their dose in deodorants or frequency of application and consequently decrease the degree of exposure to such compounds. Formulation design of personal care products may be influenced by performing detailed outcome studies. In addition, longer term impact studies are needed, perhaps in multiple year follow-up studies, to assess if the changes we observed are permanent or if they will recover to the original state.
Some of the host- and microbiome-modified molecules were also detected consistently, such as acylcarnitines, bile acids, and certain steroids. This means that a portion of the molecular composition of a person’s skin is not influenced by the beauty products applied to the skin, perhaps reflecting the level of exercise for acylcarnitines [ 53 , 54 ] or the liver (dominant location where they are made) or gallbladder (where they are stored) function for bile acids. The bile acid levels are not related to sex and do not change in amount during the course of this study. While bile acids are typically associated with the human gut microbiome [ 34 , 55 , 56 , 57 , 58 ], it is unclear what their role is on the skin and how they get there. One hypothesis is that they are present in the sweat that is excreted through the skin, as this is the case for several food-derived molecules such as caffeine or drugs and medications that have been previously reported on the human skin [ 19 ] or that microbes synthesize them de novo [ 55 ]. The only reports we could find on bile acids being associated with the skin describe cholestasis and pruritus diseases. Cholestasis and pruritus in hepatobiliary disease have symptoms of skin bile acid accumulation that are thought to be responsible for severe skin itching [ 59 , 60 ]. However, since bile acids were found in over 50% of the healthy volunteers, their detection on the skin is likely a common phenotype among the general population and not only reflective of disease, consistent with recent reports challenging these molecules as biomarkers of disease [ 59 ]. Other molecules that were detected consistently came from personal care products.
Aside from molecules that are person-specific and those that do not vary, there are others that can be modified via personal care routines. Most striking is how the personal care routines influenced changes in hormones and pheromones in a personalized manner. This suggests that there may be personalized recipes that make it possible to make someone more or less attractive to others via adjustments of hormonal and pheromonal levels through alterations in skin care.
Here, we describe the utilization of an approach that combines metabolomics and microbiome analysis to assess the effect of modifying personal care regime on skin chemistry and microbes. The key findings are as follows: (1) Compounds from beauty products last on the skin for weeks after their first use despite daily showering. (2) Beauty products alter molecular and bacterial diversity as well as the dynamic and structure of molecules and bacteria on the skin. (3) Molecular and bacterial temporal variability is product-, site-, and person-specific, and changes are observed starting the first week of beauty product use. This study provides a framework for future investigations to understand how lifestyle characteristics such as diet, outdoor activities, exercise, and medications shape the molecular and microbial composition of the skin. These factors have been studied far more in their impact on the gut microbiome and chemistry than in the skin. Revealing how such factors can affect skin microbes and their associated metabolites may be essential to define long-term skin health by restoring the appropriate microbes particularly in the context of skin aging [ 61 ] and skin diseases [ 49 ] as has shown to be necessary for amphibian health [ 62 , 63 ], or perhaps even create a precision skin care approach that utilizes the proper care ingredients based on the microbial and chemical signatures that could act as key players in host defense [ 49 , 64 , 65 ].
Subject recruitment and sample collection
Twelve individuals between 25 and 40 years old were recruited to participate in this study, six females and six males. Female volunteer 8 dropped out of the study as she developed a skin irritation during the T1–T3 phase. All volunteers signed a written informed consent in accordance with the sampling procedure approved by the UCSD Institutional Review Board (Approval Number 161730). Volunteers were required to follow specific instructions during 9 weeks. They were asked to bring in samples of their personal care products they used prior to T0 so they could be sampled as well. Following the initial timepoint time 0 and during the first 3 weeks (week 1–week 3), volunteers were asked not to use any beauty products (Fig. 1 b). During the next 3 weeks (week 4–week 6), four selected commercial beauty products provided to all volunteers were applied once a day at specific body part (deodorant for the armpits, soothing foot powder between the toes, sunscreen for the face, and moisturizer for front forearms) (Fig. 1 b, Additional file 3 : Table S2 Ingredient list of beauty products). During the first 6 weeks, volunteers were asked to shower with a head to toe shampoo. During the last 3 weeks (week 7–week 9), all volunteers went back to their normal routine and used the personal care products used before the beginning of the study (Fig. 1 b). Volunteers were asked not to shower the day before sampling. Samples were collected by the same three researchers to ensure consistency in sampling and the area sampled. Researchers examined every subject together and collected metabolomics and microbiome samples from each location together. Samples were collected once a week (from day 0 to day 68—10 timepoints total) for volunteers 1, 2, 3, 4, 5, 6, 7, 9, 10, 11, and 12, and on day 0 and day 6 for volunteer 8. For individuals 4, 9, and 10, samples were collected twice a week. Samples collected for 11 volunteers during 10 timepoints: 11 volunteers × 10 timepoints × 4 samples × 4 body sites = 1760. Samples collected from 3 selected volunteers during 9 additional timepoints: 3 volunteers × 9 timepoints × 4 samples × 4 body sites = 432. All samples were collected following the same protocol described in [ 18 ]. Briefly, samples were collected over an area of 2 × 2 cm, using pre-moistened swabs in 50:50 ethanol/water solution for metabolomics analysis or in Tris-EDTA buffer for 16S rRNA sequencing. Four samples were collected from each body part right and left side. The locations sampled were the face—upper cheek bone and lower jaw, armpit—upper and lower area, arm—front of the elbow (antecubitis) and forearm (antebrachium), and feet—in between the first and second toe and third and fourth toe. Including personal care product references, a total of 2275 samples were collected over 9 weeks and were submitted to both metabolomics and microbial inventories.
Metabolite extraction and UPLC-Q-TOF mass spectrometry analysis
Skin swabs were extracted and analyzed using a previously validated workflow described in [ 18 , 19 ]. All samples were extracted in 200 μl of 50:50 ethanol/water solution for 2 h on ice then overnight at − 20 °C. Swab sample extractions were dried down in a centrifugal evaporator then resuspended by vortexing and sonication in a 100 μl 50:50 ethanol/water solution containing two internal standards (fluconazole 1 μM and amitriptyline 1 μM). The ethanol/water extracts were then analyzed using a previously validated UPLC-MS/MS method [ 18 , 19 ]. We used a ThermoScientific UltiMate 3000 UPLC system for liquid chromatography and a Maxis Q-TOF (Quadrupole-Time-of-Flight) mass spectrometer (Bruker Daltonics), controlled by the Otof Control and Hystar software packages (Bruker Daltonics) and equipped with ESI source. UPLC conditions of analysis are 1.7 μm C18 (50 × 2.1 mm) UHPLC Column (Phenomenex), column temperature 40 °C, flow rate 0.5 ml/min, mobile phase A 98% water/2% acetonitrile/0.1% formic acid ( v / v ), mobile phase B 98% acetonitrile/2% water/0.1% formic acid ( v / v ). A linear gradient was used for the chromatographic separation: 0–2 min 0–20% B, 2–8 min 20–99% B, 8–9 min 99–99% B, 9–10 min 0% B. Full-scan MS spectra ( m/z 80–2000) were acquired in a data-dependant positive ion mode. Instrument parameters were set as follows: nebulizer gas (nitrogen) pressure 2 Bar, capillary voltage 4500 V, ion source temperature 180 °C, dry gas flow 9 l/min, and spectra rate acquisition 10 spectra/s. MS/MS fragmentation of 10 most intense selected ions per spectrum was performed using ramped collision induced dissociation energy, ranged from 10 to 50 eV to get diverse fragmentation patterns. MS/MS active exclusion was set after 4 spectra and released after 30 s.
Mass spectrometry data collected from the skin of 12 individuals can be found here MSV000081582.
LC-MS data processing
LC-MS raw data files were converted to mzXML format using Compass Data analysis software (Bruker Daltonics). MS1 features were selected for all LC-MS datasets collected from the skin of 12 individuals and blank samples (total 2275) using the open-source software MZmine [ 66 ]—see Additional file 4 : Table S3 for parameters. Subsequent blank filtering, total ion current, and internal standard normalization were performed (Additional file 5 : Table S4) for representation of relative abundance of molecular features (Fig. 2 , Additional file 1 : Figure S1), principal coordinate analysis (PCoA) (Fig. 4 ). For steroid compounds in Fig. 5 a–d, bile acids (Additional file 1 : Figure S5A-D), and acylcarnitines (Additional file 1 : Figure S5E, F) compounds, crop filtering feature available in MZmine [ 66 ] was used to identify each feature separately in all LC-MS data collected from the skin of 12 individuals (see Additional file 4 : Table S3 for crop filtering parameters and feature finding in Additional file 6 : Table S5).
Heatmap in Fig. 2 was constructed from the bucket table generated from LC-MS1 features (Additional file 7 : Table S6) and associated metadata (Additional file 8 : Table S7) using the Calour command line available here: https://github.com/biocore/calour . Calour parameters were as follows: normalized read per sample 5000 and cluster feature minimum reads 50. Procrustes and Pearson correlation analyses in Additional file 1 : Figures S10 and S11 were performed using the feature table in Additional file 9 : Table S8, normalized using the probabilistic quotient normalization method [ 67 ].
16S rRNA amplicon sequencing
16S rRNA sequencing was performed following the Earth Microbiome Project protocols [ 68 , 69 ], as described before [ 18 ]. Briefly, DNA was extracted using MoBio PowerMag Soil DNA Isolation Kit and the V4 region of the 16S rRNA gene was amplified using barcoded primers [ 70 ]. PCR was performed in triplicate for each sample, and V4 paired-end sequencing [ 70 ] was performed using Illumina HiSeq (La Jolla, CA). Raw sequence reads were demultiplexed and quality controlled using the defaults, as provided by QIIME 1.9.1 [ 71 ]. The primary OTU table was generated using Qiita ( https://qiita.ucsd.edu/ ), using UCLUST ( https://academic.oup.com/bioinformatics/article/26/19/2460/230188 ) closed-reference OTU picking method against GreenGenes 13.5 database [ 72 ]. Sequences can be found in EBI under accession number EBI: ERP104625 or in Qiita ( qiita.ucsd.edu ) under Study ID 10370. Resulting OTU tables were then rarefied to 10,000 sequences/sample for downstream analyses (Additional file 10 Table S9). See Additional file 11 : Table S10 for read count per sample and Additional file 1 : Figure S13 representing the samples that fall out with rarefaction at 10,000 threshold. The dataset includes 35 blank swab controls and 699 empty controls. The blank samples can be accessed through Qiita ( qiita.ucsd.edu ) as study ID 10370 and in EBI with accession number EBI: ERP104625. Blank samples can be found under the metadata category “sample_type” with the name “empty control” and “Swabblank.” These samples fell below the rarefaction threshold at 10,000 (Additional file 11 : Table S10).
To rule out the possibility that personal care products themselves contained the microbes that induced the changes in the armpit and foot microbiomes that were observed in this study (Fig. 7 ), we subjected the common personal care products that were used in this study during T4–T6 also to 16S rRNA sequencing. The data revealed that within the limit of detectability of the current experiment, few 16S signatures were detected. One notable exception was the most dominant plant-originated bacteria chloroplast detected in the sunscreen lotion applied on the face (Additional file 1 : Figure S9D), that was also detected on the face of individuals and at a lower level on their arms, sites where stable microbial communities were observed over time (Additional file 1 : Figure S9E, F). This finding is in agreement with our previous data from the 3D cartographical skin maps that revealed the presence of co-localized chloroplast and lotion molecules [ 18 ]. Other low-abundant microbial signatures found in the sunscreen lotion include additional plant-associated bacteria: mitochondria [ 73 ], Bacillaceae [ 74 , 75 ], Planococcaceae [ 76 ], and Ruminococcaceae family [ 77 ], but all these bacteria are not responsible for microbial changes associated to beauty product use, as they were poorly detected in the armpits and feet (Fig. 7 ).
To assess the origin of Cyanobacteria detected in skin samples, each Greengenes [ 72 ] 13_8 97% OTU table (per lane; obtained from Qiita [ 78 ] study 10,370) was filtered to only features with a p__Cyanobacteria phylum. The OTU maps for these tables—which relate each raw sequence to an OTU ID—were then filtered to only those observed p__Cyanobacteria OTU IDs. The filtered OTU map was used to extract the raw sequences into a single file. Separately, the unaligned Greengenes 13_8 99% representative sequences were filtered into two sets, first the set of representatives associated with c__Chloroplast (our interest database), and second the set of sequences associated with p__Cyanobacteria without the c__Chloroplast sequences (our background database). Platypus Conquistador [ 79 ] was then used to determine what reads were observed exclusively in the interest database and not in the background database. Of the 4,926,465 raw sequences associated with a p__Cyanobacteria classification (out of 318,686,615 total sequences), at the 95% sequence identity level with 100% alignment, 4,860,258 sequences exclusively recruit to full-length chloroplast 16S by BLAST [ 80 ] with the bulk recruiting to streptophytes (with Chlorophyta and Stramenopiles to a lesser extent). These sequences do not recruit non-chloroplast Cyanobacteria full length 16S.
Half-life calculation for metabolomics data
In order to estimate the biological half-life of molecules detected in the skin, the first four timepoints of the study (T0, T1, T2, T3) were considered for the calculation to allow the monitoring of personal beauty products used at T0. The IUPAC’s definition of biological half-life as the time required to a substance in a biological system to be reduced to half of its value, assuming an approximately exponential removal [ 81 ] was used. The exponential removal can be described as C ( t ) = C 0 e − tλ where t represents the time in weeks, C 0 represents the initial concentration of the molecule, C ( t ) represents the concentration of the molecule at time t , and λ is the rate of removal [ http://onlinelibrary.wiley.com/doi/10.1002/9780470140451.ch2/summary ]. The parameter λ was estimated by a mixed linear effects model in order to account for the paired sample structure. The regression model tests the null hypothesis that λ is equal to zero and only the significant ( p value < 0.05) parameters were considered.
Principal coordinate analysis
We performed principal coordinate analysis (PCoA) on both metabolomics and microbiome data. For metabolomics, we used MS1 features (Additional file 5 : Table S4) and calculated Bray–Curtis dissimilarity metric using ClusterApp ( https://github.com/mwang87/q2_metabolomics ).
For microbiome data, we used rarefied OTU table (Additional file 10 : Table S9) and used unweighted UniFrac metric [ 36 ] to calculate beta diversity distance matrix using QIIME2 (https://qiime2.org). Results from both data sources were visualized using Emperor ( https://biocore.github.io/emperor/ ) [ 28 ].
Molecular networking
Molecular networking was generated from LC-MS/MS data collected from skin samples of 11 individuals MSV000081582, using the Global Natural Products Social Molecular Networking platform (GNPS) [ 29 ]. Molecular network parameters for MS/MS data collected from all body parts of 11 individuals during T0–T9 MSV000081582 are accessible here http://gnps.ucsd.edu/ProteoSAFe/status.jsp?task=284fc383e4c44c4db48912f01905f9c5 . Molecular network parameters for MS/MS data collected from armpits T0–T3 MSV000081582 and deodorant used by individual 1 and 3 MSV000081580 can be found here http://gnps.ucsd.edu/ProteoSAFe/status.jsp?task=f5325c3b278a46b29e8860ec57915ad and here http://gnps.ucsd.edu/ProteoSAFe/status.jsp?task=aaa1af68099d4c1a87e9a09f398fe253 , respectively. Molecular networks were exported and visualized in Cytoscape 3.4.0. [ 82 ]. Molecular networking parameters were set as follows: parent mass tolerance 1 Da, MS/MS fragment ion tolerance 0.5 Da, and cosine threshold 0.65 or greater, and only MS/MS spectral pairs with at least 4 matched fragment ions were included. Each MS/MS spectrum was only allowed to connect to its top 10 scoring matches, resulting in a maximum of 10 connections per node. The maximum size of connected components allowed in the network was 600, and the minimum number of spectra required in a cluster was 3. Venn diagrams were generated from Cytoscape data http://gnps.ucsd.edu/ProteoSAFe/status.jsp?task=284fc383e4c44c4db48912f01905f9c5 using Cytoscape [ 82 ] Venn diagram app available here http://apps.cytoscape.org/apps/all .
Shannon molecular and bacterial diversity
The diversity analysis was performed separately for 16S rRNA data and LC-MS data. For each sample in each feature table (LC-MS data and microbiome data), we calculated the value of the Shannon diversity index. For LC-MS data, we used the full MZmine feature table (Additional file 5 : Table S4). For microbiome data, we used the closed-reference BIOM table rarefied to 10,000 sequences/sample. For diversity changes between timepoints, we aggregated Shannon diversity values across groups of individuals (all, females, males) and calculated mean values and standard errors. All successfully processed samples (detected features in LC-MS or successful sequencing with 10,000 or more sequences/sample) were considered.
Beauty products and chemical standards
Samples (10 mg) from personal care products used during T0 and T7–T9 MSV000081580 (Additional file 2 : Table S1) and common beauty products used during T4–T6 MSV000081581 (Additional file 3 : Table S2) were extracted in 1 ml 50:50 ethanol/water. Sample extractions were subjected to the same UPLC-Q-TOF MS method used to analyze skin samples and described above in the section “ Metabolite extraction and UPLC-Q-TOF mass spectrometry analysis .” Authentic chemical standards MSV000081583 including 1-dehydroandrostenedion (5 μM), chenodeoxyglycocholic acid (5 μM), dehydroisoandrosterone sulfate (100 μM), glycocholic acid (5 μM), and taurocholic acid (5 μM) were analyzed using the same mass spectrometry workflow used to run skin and beauty product samples.
Monitoring beauty product ingredients in skin samples
In order to monitor beauty product ingredients used during T4–T6, we selected only molecular features present in each beauty product sample (antiperspirant, facial lotion, body moisturizer, soothing powder) and then filtered the aligned MZmine feature table (Additional file 5 : Table S4) for the specific feature in specific body part samples. After feature filtering, we selected all features that had a higher average intensity on beauty product phase (T4–T6) compared to non-beauty product phase (T1–T3). The selected features were annotated using GNPS dereplication output http://gnps.ucsd.edu/ProteoSAFe/status.jsp?task=69319caf219642a5a6748a3aba8914df , plotted using R package ggplot2 ( https://cran.r-project.org/web/packages/ggplot2/index.html ) and visually inspected for meaningful patterns.
Random forest analysis
Random forest analysis was performed in MetaboAnalyst 3.0 online platform http://www.metaboanalyst.ca/faces/home.xhtml . Using LC-MS1 features found in armpit samples collected on T3 and T6. Random forest parameters were set as follows: top 1000 most abundant features, number of predictors to try for each node 7, estimate of error rate (0.0%).
BugBase analysis
To determine the functional potential of microbial communities within our samples, we used BugBase [ 83 ]. Because we do not have direct access to all of the gene information due to the use of 16S rRNA marker gene sequencing, we can only rely on phylogenetic information inferred from OTUs. BugBase takes advantage of this information to predict microbial phenotypes by associating OTUs with gene content using PICRUSt [ 84 ]. Thus, using BugBase, we can predict such phenotypes as Gram staining, or oxidative stress tolerance at each timepoint or each phase. All statistical analyses in BugBase are performed using non-parametric differentiation tests (Mann–Whitney U ).
Taxonomic plots
Rarefied OTU counts were collapsed according to the OTU’s assigned family and genus name per sample, with a single exception for the class of chloroplasts. Relative abundances of each family-genus group are obtained by dividing by overall reads per sample, i.e., 10,000. Samples are grouped by volunteer, body site, and time/phase. Abundances are aggregated by taking the mean overall samples, and resulting abundances are again normalized to add up to 1. Low-abundant taxa are not listed in the legend and plotted in grayscale. Open-source code is available at https://github.com/sjanssen2/ggmap/blob/master/ggmap/snippets.py
Dissimilarity-based analysis
Pairwise dissimilarity matrices were generated for metabolomics and 16S metagenomics quantification tables, described above, using Bray–Curtis dissimilarity through QIIME 1.9.1 [ 71 ]. Those distance matrices were used to perform Procrustes analysis (QIIME 1.9.1), and Mantel test (scikit-bio version 0.5.1) to measure the correlation between the metabolome and microbiome over time. The metabolomics dissimilarities were used to perform the PERMANOVA test to assess the significance of body part grouping. The PCoA and Procrustes plots were visualized in EMPeror. The dissimilarity matrices were also used to perform distance tests, comparing the distances within and between individuals and distances from time 0 to times 1, 2, and 3 using Wilcoxon rank-sum tests (SciPy version 0.19.1) [ 19 ].
Statistical analysis for molecular and microbial data
Statistical analyses were performed in R and Python (R Core Team 2018). Monotonic relationships between two variables were tested using non-parametric Spearman correlation tests. The p values for correlation significance were subsequently corrected using Benjamini and Hochberg false discovery rate control method. The relationship between two groups was tested using non-parametric Wilcoxon rank-sum tests. The relationship between multiple groups was tested using non-parametric Kruskal–Wallis test. The significance level was set to 5%, unless otherwise mentioned, and all tests were performed as two-sided tests.
Oh J, Byrd AL, Deming C, Conlan S, Kong HH, Segre JA. Biogeography and individuality shape function in the human skin metagenome. Nature. 2014;514(7520):59–64.
Article CAS PubMed PubMed Central Google Scholar
Grice EA, Segre JA. The skin microbiome. Nat Rev Microbiol. 2011;9(4):244–53.
Costello EK, Lauber CL, Hamady M, Fierer N, Gordon JI, Knight R. Bacterial community variation in human body habitats across space and time. Science. 2009;326(5960):1694–7.
Grice EA, Kong HH, Conlan S, Deming CB, Davis J, Young AC, et al. Topographical and temporal diversity of the human skin microbiome. Science. 2009;324(5931):1190–2.
Urban J, Fergus DJ, Savage AM, Ehlers M, Menninger HL, Dunn RR, et al. The effect of habitual and experimental antiperspirant and deodorant product use on the armpit microbiome. PeerJ. 2016;4:e1605.
Article PubMed PubMed Central Google Scholar
Callewaert C, Hutapea P, Van de Wiele T, Boon N. Deodorants and antiperspirants affect the axillary bacterial community. Arch Dermatol Res. 2014;306(8):701–10.
Article CAS PubMed Google Scholar
Staudinger T, Pipal A, Redl B. Molecular analysis of the prevalent microbiota of human male and female forehead skin compared to forearm skin and the influence of make-up. J Appl Microbiol. 2011;110(6):1381–9.
Houben E, De Paepe K, Rogiers V. A keratinocyte’s course of life. Skin Pharmacol Physiol. 2007;20(3):122–32.
Hoath SB, Leahy DG. The organization of human epidermis: functional epidermal units and phi proportionality. J Invest Dermatol. 2003;121(6):1440–6.
Oh J, Byrd AL, Park M, Kong HH, Segre JA. Temporal stability of the human skin microbiome. Cell. 2016;165(4):854–66.
Schloissnig S, Arumugam M, Sunagawa S, Mitreva M, Tap J, Zhu A, et al. Genomic variation landscape of the human gut microbiome. Nature. 2013;493(7430):45–50.
Article PubMed Google Scholar
Faith JJ, Guruge JL, Charbonneau M, Subramanian S, Seedorf H, Goodman AL, et al. The long-term stability of the human gut microbiota. Science. 2013;341(6141):1237439.
Hall MW, Singh N, Ng KF, Lam DK, Goldberg MB, Tenenbaum HC, et al. Inter-personal diversity and temporal dynamics of dental, tongue, and salivary microbiota in the healthy oral cavity. NPJ Biofilms Microbiomes. 2017;3:2.
Utter DR, Mark Welch JL, Borisy GG. Individuality, stability, and variability of the plaque microbiome. Front Microbiol. 2016;7:564.
Flores GE, Caporaso JG, Henley JB, Rideout JR, Domogala D, Chase J, et al. Temporal variability is a personalized feature of the human microbiome. Genome Biol. 2014;15(12):531.
The Human Microbiome Project C. Structure, function and diversity of the healthy human microbiome. Nature [Article]. 2012;486:207.
Article Google Scholar
Dorrestein PC, Gallo RL, Knight R. Microbial skin inhabitants: friends forever. Cell. 2016;165(4):771–2.
Bouslimani A, Porto C, Rath CM, Wang M, Guo Y, Gonzalez A, et al. Molecular cartography of the human skin surface in 3D. Proc Natl Acad Sci U S A. 2015;112(17):E2120–9.
Bouslimani A, Melnik AV, Xu Z, Amir A, da Silva RR, Wang M, et al. Lifestyle chemistries from phones for individual profiling. Proc Natl Acad Sci U S A. 2016;113(48):E7645–E54.
David LA, Maurice CF, Carmody RN, Gootenberg DB, Button JE, Wolfe BE, et al. Diet rapidly and reproducibly alters the human gut microbiome. Nature. 2014;505(7484):559–63.
Wu GD, Chen J, Hoffmann C, Bittinger K, Chen YY, Keilbaugh SA, et al. Linking long-term dietary patterns with gut microbial enterotypes. Science. 2011;334(6052):105–8.
Unno M, Cho O, Sugita T. Inhibition of Propionibacterium acnes lipase activity by the antifungal agent ketoconazole. Microbiol Immunol. 2017;61(1):42–4.
Holland C, Mak TN, Zimny-Arndt U, Schmid M, Meyer TF, Jungblut PR, et al. Proteomic identification of secreted proteins of Propionibacterium acnes. BMC Microbiol. 2010;10:230.
Nguyen DD, Wu CH, Moree WJ, Lamsa A, Medema MH, Zhao X, et al. MS/MS networking guided analysis of molecule and gene cluster families. Proc Natl Acad Sci U S A. 2013;110(28):E2611–20.
Soltanpour S, Jouyban A. Solubility of acetaminophen and ibuprofen in polyethylene glycol 600, propylene glycol and water mixtures at 25°C. J Mol Liq. 2010;155(2):80–4.
Article CAS Google Scholar
Haglund BO. Solubility studies of polyethylene glycols in ethanol and water. Thermochimica Acta. 1987;114(1):97–102.
Petras D, Nothias LF, Quinn RA, Alexandrov T, Bandeira N, Bouslimani A, et al. Mass spectrometry-based visualization of molecules associated with human habitats. Anal Chem. 2016;88(22):10775–84.
Vazquez-Baeza Y, Pirrung M, Gonzalez A, Knight R. EMPeror: a tool for visualizing high-throughput microbial community data. Gigascience. 2013;2(1):16.
Wang M, Carver JJ, Phelan VV, Sanchez LM, Garg N, Peng Y, et al. Sharing and community curation of mass spectrometry data with Global Natural Products Social Molecular Networking. Nat Biotechnol. 2016;34(8):828–37.
Watrous J, Roach P, Alexandrov T, Heath BS, Yang JY, Kersten RD, et al. Mass spectral molecular networking of living microbial colonies. Proc Natl Acad Sci U S A. 2012;109(26):E1743–52.
Frank AM, Monroe ME, Shah AR, Carver JJ, Bandeira N, Moore RJ, et al. Spectral archives: extending spectral libraries to analyze both identified and unidentified spectra. Nat Methods. 2011;8(7):587–91.
Quinn RA, Nothias LF, Vining O, Meehan M, Esquenazi E, Dorrestein PC. Molecular networking as a drug discovery, drug metabolism, and precision medicine strategy. Trends Pharmacol Sci. 2017;38(2):143–54.
Luzzatto-Knaan T, Garg N, Wang M, Glukhov E, Peng Y, Ackermann G, et al. Digitizing mass spectrometry data to explore the chemical diversity and distribution of marine cyanobacteria and algae. Elife. 2017;6:e24214.
Melnik AV, da Silva RR, Hyde ER, Aksenov AA, Vargas F, Bouslimani A, et al. Coupling targeted and untargeted mass spectrometry for metabolome-microbiome-wide association studies of human fecal samples. Anal Chem. 2017;89(14):7549–59.
Sumner LW, Amberg A, Barrett D, Beale MH, Beger R, Daykin CA, et al. Proposed minimum reporting standards for chemical analysis Chemical Analysis Working Group (CAWG) Metabolomics Standards Initiative (MSI). Metabolomics. 2007;3(3):211–21.
Lozupone C, Knight R. UniFrac: a new phylogenetic method for comparing microbial communities. Appl Environ Microbiol. 2005;71(12):8228–35.
Caporaso JG, Lauber CL, Costello EK, Berg-Lyons D, Gonzalez A, Stombaugh J, et al. Moving pictures of the human microbiome. Genome Biol. 2011;12(5):R50.
Green BR. Chloroplast genomes of photosynthetic eukaryotes. Plant J. 2011;66(1):34–44.
Callewaert C, Kerckhof FM, Granitsiotis MS, Van Gele M, Van de Wiele T, Boon N. Characterization of Staphylococcus and Corynebacterium clusters in the human axillary region. PLoS One. 2013;8(8):e70538.
Callewaert C, Lambert J, Van de Wiele T. Towards a bacterial treatment for armpit malodour. Exp Dermatol. 2017;26(5):388–91.
Tripathi A, Melnik AV, Xue J, Poulsen O, Meehan MJ, Humphrey G, et al. Intermittent hypoxia and hypercapnia, a hallmark of obstructive sleep apnea, alters the gut microbiome and metabolome. mSystems. 2018;3(3):e00020-18.
Gower JC. Generalized procrustes analysis. Psychometrika [journal article]. 1975;40(1):33–51.
Decreau RA, Marson CM, Smith KE, Behan JM. Production of malodorous steroids from androsta-5,16-dienes and androsta-4,16-dienes by Corynebacteria and other human axillary bacteria. J Steroid Biochem Mol Biol. 2003;87(4–5):327–36.
Austin C, Ellis J. Microbial pathways leading to steroidal malodour in the axilla. J Steroid Biochem Mol Biol. 2003;87(1):105–10.
Lloyd-Price J, Mahurkar A, Rahnavard G, Crabtree J, Orvis J, Hall AB, et al. Strains, functions and dynamics in the expanded Human Microbiome Project. Nature. 2017;550(7674):61-6.
Kapono CA, Morton JT, Bouslimani A, Melnik AV, Orlinsky K, Knaan TL, et al. Creating a 3D microbial and chemical snapshot of a human habitat. Sci Rep. 2018;8(1):3669.
Clemente JC, Pehrsson EC, Blaser MJ, Sandhu K, Gao Z, Wang B, et al. The microbiome of uncontacted Amerindians. Sci Adv. 2015;1(3):e1500183.
Blaser MJ, Dominguez-Bello MG, Contreras M, Magris M, Hidalgo G, Estrada I, et al. Distinct cutaneous bacterial assemblages in a sampling of South American Amerindians and US residents. ISME J. 2013;7(1):85–95.
Nakatsuji T, Chen TH, Narala S, Chun KA, Two AM, Yun T, et al. Antimicrobials from human skin commensal bacteria protect against Staphylococcus aureus and are deficient in atopic dermatitis. Sci Transl Med 2017;9(378)eaah4680.
Hollands A, Gonzalez D, Leire E, Donald C, Gallo RL, Sanderson-Smith M, et al. A bacterial pathogen co-opts host plasmin to resist killing by cathelicidin antimicrobial peptides. J Biol Chem. 2012;287(49):40891–7.
Zirwas MJ, Moennich J. Antiperspirant and deodorant allergy: diagnosis and management. J Clin Aesthet Dermatol. 2008;1(3):38–43.
PubMed PubMed Central Google Scholar
Funk JO, Maibach HI. Propylene glycol dermatitis: re-evaluation of an old problem. Contact Dermatitis. 1994;31(4):236–41.
Lehmann R, Zhao X, Weigert C, Simon P, Fehrenbach E, Fritsche J, et al. Medium chain acylcarnitines dominate the metabolite pattern in humans under moderate intensity exercise and support lipid oxidation. PLoS One. 2010;5(7):e11519.
Hiatt WR, Regensteiner JG, Wolfel EE, Ruff L, Brass EP. Carnitine and acylcarnitine metabolism during exercise in humans. Dependence on skeletal muscle metabolic state. J Clin Invest. 1989;84(4):1167–73.
Fischbach MA, Segre JA. Signaling in host-associated microbial communities. Cell. 2016;164(6):1288–300.
Devlin AS, Fischbach MA. A biosynthetic pathway for a prominent class of microbiota-derived bile acids. Nat Chem Biol [Article]. 2015;11(9):685–90.
Ridlon JM, Kang DJ, Hylemon PB, Bajaj JS. Bile acids and the gut microbiome. Curr Opin Gastroenterol. 2014;30(3):332–8.
Humbert L, Maubert MA, Wolf C, Duboc H, Mahe M, Farabos D, et al. Bile acid profiling in human biological samples: comparison of extraction procedures and application to normal and cholestatic patients. J Chromatogr B Analyt Technol Biomed Life Sci. 2012;899:135–45.
Ghent CN, Bloomer JR. Itch in liver disease: facts and speculations. Yale J Biol Med. 1979;52(1):77–82.
CAS PubMed PubMed Central Google Scholar
Herndon JH Jr. Pathophysiology of pruritus associated with elevated bile acid levels in serum. Arch Intern Med. 1972;130(4):632–7.
Zapata HJ, Quagliarello VJ. The microbiota and microbiome in aging: potential implications in health and age-related diseases. J Am Geriatr Soc. 2015;63(4):776–81.
Kueneman JG, Woodhams DC, Harris R, Archer HM, Knight R, McKenzie VJ. Probiotic treatment restores protection against lethal fungal infection lost during amphibian captivity. Proc Biol Sci. 2016;283(1839):e20161553.
Woodhams DC, Brandt H, Baumgartner S, Kielgast J, Kupfer E, Tobler U, et al. Interacting symbionts and immunity in the amphibian skin mucosome predict disease risk and probiotic effectiveness. PLoS One. 2014;9(4):e96375.
Belkaid Y, Tamoutounour S. The influence of skin microorganisms on cutaneous immunity. Nat Rev Immunol. 2016;16(6):353–66.
Belkaid Y, Segre JA. Dialogue between skin microbiota and immunity. Science. 2014;346(6212):954–9.
Pluskal T, Castillo S, Villar-Briones A, Oresic M. MZmine 2: modular framework for processing, visualizing, and analyzing mass spectrometry-based molecular profile data. BMC Bioinformatics. 2010;11:395.
Dieterle F, Ross A, Schlotterbeck G, Senn H. Probabilistic quotient normalization as robust method to account for dilution of complex biological mixtures. Application in 1H NMR metabonomics. Anal Chem. 2006;78(13):4281–90.
Gilbert JA, Jansson JK, Knight R. The Earth Microbiome project: successes and aspirations. BMC Biol. 2014;12:69.
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Huntley J, Fierer N, et al. Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J. 2012;6(8):1621–4.
Walters W, Hyde ER, Berg-Lyons D, Ackermann G, Humphrey G, Parada A, et al. Improved bacterial 16S rRNA gene (V4 and V4-5) and fungal internal transcribed spacer marker gene primers for microbial community surveys. mSystems. 2016;1(1):e00009-15.
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, et al. QIIME allows analysis of high-throughput community sequencing data. Nat Methods. 2010;7(5):335–6.
McDonald D, Price MN, Goodrich J, Nawrocki EP, DeSantis TZ, Probst A, et al. An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea. ISME J. 2012;6(3):610–8.
Haferkamp I. The diverse members of the mitochondrial carrier family in plants. FEBS Lett. 2007;581(12):2375–9.
Burgess SA, Flint SH, Lindsay D, Cox MP, Biggs PJ. Insights into the Geobacillus stearothermophilus species based on phylogenomic principles. BMC Microbiol. 2017;17(1):140.
Goh KM, Gan HM, Chan KG, Chan GF, Shahar S, Chong CS, et al. Analysis of Anoxybacillus genomes from the aspects of lifestyle adaptations, prophage diversity, and carbohydrate metabolism. PLoS One. 2014;9(6):e90549.
Carvalhais LC, Dennis PG, Badri DV, Tyson GW, Vivanco JM, Schenk PM. Activation of the jasmonic acid plant defence pathway alters the composition of rhizosphere bacterial communities. PLoS One. 2013;8(2):e56457.
Barelli C, Albanese D, Donati C, Pindo M, Dallago C, Rovero F, et al. Habitat fragmentation is associated to gut microbiota diversity of an endangered primate: implications for conservation. Sci Rep. 2015;5:14862.
Gonzalez A, Navas-Molina JA, Kosciolek T, McDonald D, Vazquez-Baeza Y, Ackermann G, et al. Qiita: rapid, web-enabled microbiome meta-analysis. Nat Methods. 2018;15(10):796–8.
Gonzalez A, Vazquez-Baeza Y, Pettengill JB, Ottesen A, McDonald D, Knight R. Avoiding pandemic fears in the subway and conquering the platypus. mSystems. 2016;1(3):e00050-16.
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215(3):403–10.
Wilkinson ADMaA. IUPAC. Compendium of chemical terminology, 2nd ed. (the "Gold Book": Blackwell Scientific Publications, Oxford 1997.
Smoot ME, Ono K, Ruscheinski J, Wang PL, Ideker T. Cytoscape 2.8: new features for data integration and network visualization. Bioinformatics. 2011;27(3):431–2.
Ward T, Larson J, Meulemans J, Hillmann B, Lynch J, Sidiropoulos D, et al. BugBase Predicts Organism Level Microbiome Phenotypes. bioRxiv. 2017;133462. https://doi.org/10.1101/133462 .
Langille MGI, Zaneveld J, Caporaso JG, McDonald D, Knights D, Reyes JA, et al. Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat Biotech [Computational Biology]. 2013;31(9):814–21.
Download references

Acknowledgements
We thank all volunteers who were recruited in this study for their participation and Carla Porto for discussions regarding beauty products selected in this study. We further acknowledge Bruker for the support of the shared instrumentation infrastructure that enabled this work.
This work was partially supported by US National Institutes of Health (NIH) Grant. P.C.D. acknowledges funding from the European Union’s Horizon 2020 Programme (Grant 634402). A.B was supported by the National Institute of Justice Award 2015-DN-BX-K047. C.C. was supported by a fellowship of the Belgian American Educational Foundation and the Research Foundation Flanders. L.Z., J.K, and K.Z. acknowledge funding from the US National Institutes of Health under Grant No. AR071731. TLK was supported by Vaadia-BARD Postdoctoral Fellowship Award No. FI-494-13.
Availability of data and materials
The mass spectrometry data have been deposited in the MassIVE database (MSV000081582, MSV000081580 and MSV000081581). Molecular network parameters for MS/MS data collected from all body parts of 11 individuals during T0-T9 MSV000081582 are accessible here http://gnps.ucsd.edu/ProteoSAFe/status.jsp?task=284fc383e4c44c4db48912f01905f9c5 . Molecular network parameters for MS/MS data collected from armpits T0–T3 MSV000081582 and deodorant used by individual 1 and 3 MSV000081580 can be found here http://gnps.ucsd.edu/ProteoSAFe/status.jsp?task=f5325c3b278a46b29e8860ec5791d5ad and here http://gnps.ucsd.edu/ProteoSAFe/status.jsp?task=aaa1af68099d4c1a87e9a09f398fe253 , respectively. OTU tables can be found in Qiita ( qiita.ucsd.edu ) as study ID 10370, and sequences can be found in EBI under accession number EBI: ERP104625.
Author information
Amina Bouslimani and Ricardo da Silva contributed equally to this work.
Authors and Affiliations
Collaborative Mass Spectrometry Innovation Center, Skaggs School of Pharmacy and Pharmaceutical Sciences, San Diego, USA
Amina Bouslimani, Ricardo da Silva, Kathleen Dorrestein, Alexey V. Melnik, Tal Luzzatto-Knaan & Pieter C. Dorrestein
Department of Pediatrics, University of California, San Diego, La Jolla, CA, 92037, USA
Tomasz Kosciolek, Stefan Janssen, Chris Callewaert, Amnon Amir, Livia S. Zaramela, Ji-Nu Kim, Gregory Humphrey, Tara Schwartz, Karenina Sanders, Caitriona Brennan, Gail Ackermann, Daniel McDonald, Karsten Zengler, Rob Knight & Pieter C. Dorrestein
Department for Pediatric Oncology, Hematology and Clinical Immunology, University Children’s Hospital, Medical Faculty, Heinrich-Heine-University Düsseldorf, Düsseldorf, Germany
Stefan Janssen
Center for Microbial Ecology and Technology, Ghent University, 9000, Ghent, Belgium
Chris Callewaert
Center for Microbiome Innovation, University of California, San Diego, La Jolla, CA, 92307, USA
Karsten Zengler, Rob Knight & Pieter C. Dorrestein
Department of Bioengineering, University of California, San Diego, La Jolla, CA, 92093, USA
Karsten Zengler & Rob Knight
Department of Computer Science and Engineering, University of California, San Diego, La Jolla, CA, 92093, USA
Department of Pharmacology, University of California, San Diego, La Jolla, CA, 92037, USA
Pieter C. Dorrestein
You can also search for this author in PubMed Google Scholar
Contributions
AB and PCD contributed to the study and experimental design. AB, KD, and TLK contributed to the metabolite and microbial sample collection. AB contributed to the mass spectrometry data collection. AB, RS, and AVM contributed to the mass spectrometry data analysis. RS contributed to the metabolomics statistical analysis and microbial–molecular correlations. GH, TS, KS, and CB contributed to the 16S rRNA sequencing. AB and GA contributed to the metadata organization. TK, SJ, CC, AA, and DMD contributed to the microbial data analysis and statistics. LZ, JK, and KZ contributed to the additional data analysis. AB, PCD, and RK wrote the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Correspondence to Rob Knight or Pieter C. Dorrestein .
Ethics declarations
Ethics approval and consent to participate.
All participants signed a written informed consent in accordance with the sampling procedure approved by the UCSD Institutional Review Board (Approval Number 161730).
Competing interests
Dorrestein is on the advisory board for SIRENAS, a company that aims to find therapeutics from ocean environments. There is no overlap between this research and the company. The other authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Additional files
Additional file 1:.
Figure S1. Beauty products ingredients persist on skin of participants. Figure S2. Beauty product application impacts the molecular and bacterial diversity on skin of 11 individuals while the chemical diversity from personal beauty products used by males and females on T0 is similar. Figure S3. Longitudinal impact of ceasing and resuming the use of beauty products on the molecular composition of the skin over time. Figure S4. Molecular networking to highlight MS/MS spectra found in each body part. Figure S5. Longitudinal abundance of bile acids and acylcarnitines in skin samples. Figure S6. Characterization of steroids in armpits samples. Figure S7. Characterization of bile acids in armpit samples. Figure S8. Characterization of Acylcarnitine family members in skin samples. Figure S9. Beauty products applied at one body part might affect other areas of the body, while specific products determine stability versus variability of microflora at each body site. Figure S10. Representation of Gram-positive bacteria over time and the molecular features from the shampoo detected on feet. Figure S11. Procrustes analysis to correlate the skin microbiome and metabolome over time. Figure S12. Correlation between specific molecules and bacteria that change over time in armpits of individual 11. Figure S13. Representation of the number of samples that were removed (gray) and those retained (blue) after rarefaction at 10,000 threshold. (DOCX 1140 kb)
Additional file 2:
Table S1. List of personal (T0 and T7–9) beauty products and their frequency of use. (XLSX 30 kb)
Additional file 3:
Table S2. List of ingredients of common beauty products used during T4–T6. (PDF 207 kb)
Additional file 4:
Table S3. Mzmine feature finding and crop filtering parameters. (XLSX 4 kb)
Additional file 5:
Table S4. Feature table for statistical analysis with blank filtering and total ion current normalization. (CSV 150242 kb)
Additional file 6:
Table S5. Feature table for individual feature abundance in armpits. (XLSX 379 kb)
Additional file 7:
Table S6. Feature table for Calour analysis. (CSV 91651 kb)
Additional file 8:
Table S7. Metadata for Calour analysis. (TXT 129 kb)
Additional file 9:
Table S8. feature table with Probabilistic quotient normalization for molecular–microbial analysis. (ZIP 29557 kb)
Additional file 10:
Table S9. OTU table rarefied to 10,000 sequences per sample. (BIOM 9493 kb)
Additional file 11:
Table S10. 16S rRNA sequencing read counts per sample. (TSV 2949 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License ( http://creativecommons.org/licenses/by/4.0/ ), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The Creative Commons Public Domain Dedication waiver ( http://creativecommons.org/publicdomain/zero/1.0/ ) applies to the data made available in this article, unless otherwise stated.
Reprints and permissions
About this article
Cite this article.
Bouslimani, A., da Silva, R., Kosciolek, T. et al. The impact of skin care products on skin chemistry and microbiome dynamics. BMC Biol 17 , 47 (2019). https://doi.org/10.1186/s12915-019-0660-6
Download citation
Received : 20 February 2019
Accepted : 30 April 2019
Published : 12 June 2019
DOI : https://doi.org/10.1186/s12915-019-0660-6
Share this article
Anyone you share the following link with will be able to read this content:
Sorry, a shareable link is not currently available for this article.
Provided by the Springer Nature SharedIt content-sharing initiative
- Skin care products
- Mass spectrometry
- Metabolomics
- 16S rRNA sequencing
BMC Biology
ISSN: 1741-7007
- Submission enquiries: [email protected]
- General enquiries: [email protected]

At Skin Thesis®, we value and appreciate every client. Please note our policies and procedures:
8969 Santa Monica Blvd., West Hollywood, CA 90069

PHILOSOPHY TO ACHIEVE THE MOST NATURAL LOOK
Using latest innovations in aesthetic medicine..

Information
- Author Services
Initiatives
You are accessing a machine-readable page. In order to be human-readable, please install an RSS reader.
All articles published by MDPI are made immediately available worldwide under an open access license. No special permission is required to reuse all or part of the article published by MDPI, including figures and tables. For articles published under an open access Creative Common CC BY license, any part of the article may be reused without permission provided that the original article is clearly cited. For more information, please refer to https://www.mdpi.com/openaccess .
Feature papers represent the most advanced research with significant potential for high impact in the field. A Feature Paper should be a substantial original Article that involves several techniques or approaches, provides an outlook for future research directions and describes possible research applications.
Feature papers are submitted upon individual invitation or recommendation by the scientific editors and must receive positive feedback from the reviewers.
Editor’s Choice articles are based on recommendations by the scientific editors of MDPI journals from around the world. Editors select a small number of articles recently published in the journal that they believe will be particularly interesting to readers, or important in the respective research area. The aim is to provide a snapshot of some of the most exciting work published in the various research areas of the journal.
Original Submission Date Received: .
- Active Journals
- Find a Journal
- Proceedings Series
- For Authors
- For Reviewers
- For Editors
- For Librarians
- For Publishers
- For Societies
- For Conference Organizers
- Open Access Policy
- Institutional Open Access Program
- Special Issues Guidelines
- Editorial Process
- Research and Publication Ethics
- Article Processing Charges
- Testimonials
- Preprints.org
- SciProfiles
- Encyclopedia

Article Menu

- Subscribe SciFeed
- Recommended Articles
- Google Scholar
- on Google Scholar
- Table of Contents
Find support for a specific problem in the support section of our website.
Please let us know what you think of our products and services.
Visit our dedicated information section to learn more about MDPI.
JSmol Viewer
Regenerative cosmetics: skin tissue engineering for anti-aging, repair, and hair restoration.

1. Introduction
1.1. importance of healthy skin and hair in society, 1.2. limitations of traditional cosmetic approaches, 1.3. the promise of te-based dermocosmetics.
- Addressing the root cause: TE-based dermocosmetics aim to address the underlying biological processes responsible for various skin and hair concerns. This can involve stimulating collagen production for anti-aging effects, promoting wound healing through the delivery of growth factors, or even facilitating hair follicle (HF) regeneration through the use of bioengineered scaffolds [ 18 ].
- Enhanced efficacy: By targeting specifically the dermal compartment, dermocosmetics derived from TE, including new delivery methods, improve the efficacy of the bioactive compounds or key proteins such as collagen. This can significantly improve areas like scar regeneration and wound healing [ 19 , 20 ].
- Long-lasting results: Some TE techniques, like the application of stem cells or their exosomes, show promise for promoting long-lasting results by stimulating cell proliferation and collagen production. This can significantly reduce the need for frequent product application and improve patient compliance [ 21 , 22 ].
- Better testers: Ex vivo skin models, such as 3D “skin-on-a-chip” (SoC) systems combined with microfluidics, offer a promising alternative to traditional testing methods. These models provide a more realistic recreation of human skin architecture and function, enabling more accurate dermocosmetic product testing [ 23 , 24 ].
2. Mechanisms Involved in Regenerative Cosmetics
2.1. skin aging.
- Skin aging is characterized by a decline in collagen production and a reduction of cell proliferation, together with a decrease in stemness from each tissue, among other factors [ 25 ]. Regenerative cosmetics offer solutions to combat these age-related changes, here is a summary of the key approaches in regenerative medicine applied to address age-related skin changes ( Table 2 ).
2.2. Oxidative Stress
2.3. repair vs. regeneration.
- Promoting wound healing: Engineered skin substitutes like biocompatible scaffolds provide a structure for cell migration and tissue regeneration, accelerating wound healing and minimizing scar formation [ 53 ]. Current skin substitutes have been tested for addressing regeneration in burn patients, chronic ulcers (diabetes), and rare genodermatoses (Epidermolysis bullosa) [ 54 ]. Novel technologies, including injectable cell suspensions and 3D scaffolds, are promising for improving wound healing and skin regeneration [ 55 ].
- Scar reduction: Microneedling and fractional laser therapy, combined with regenerative ingredients like growth factors, can stimulate collagen production and improve the appearance of existing scars [ 56 , 57 ]. Additionally, platelet-rich plasma (PRP) therapy is gaining traction as a potential scar reduction technique. Studies suggest that PRP injections may improve scar quality and reduce scar tissue formation [ 58 ], which is particularly interesting in relation to acne scars.
2.4. Fibrosis and Connective Tissue in Skin Rejuvenation
- Modulate the fibrotic process: by understanding the molecular mechanisms underlying fibrosis, researchers can develop strategies to control collagen deposition and promote scarless wound healing, and also highlight the role of macrophages in the inflammatory phase [ 59 ].
- Enhance the functionality of the connective tissue: supporting the health and organization of the connective tissue, which provides structural support and elasticity to the skin, is crucial for maintaining a youthful appearance and function, and this is particularly interesting when the role of MSCs is studied in UV-associated skin aging [ 61 ].
2.5. Hair Follicle Regeneration
3. regenerative cosmetics: a transformative alternative, 3.1. omic approaches: a key tool for regenerative cosmetics, 3.1.1. proteomics, 3.1.2. metabolomics, 3.1.3. multi-omics integration, 3.2. skin modeling accelerates drug development, 3d skin-on-chip models and microfluidics for dermocosmetics.
- Materials: The 3D SoC models require balancing cost and biomimicry through material selection. Synthetic polymers (PDMS, PCL, PLA) offer affordability and biocompatibility but lack the intricate structure of natural tissues. Hydrogels (alginate, collagen) mimic tissues but struggle with maintaining precise mechanical properties. Animal-derived materials (decellularized ECM, silk) provide the most biomimetic environment, closely resembling natural skin, but are expensive and raise ethical concerns [ 95 , 98 , 99 , 100 , 101 , 102 ]. Recent innovations like decellularized ECM and silk offer promising solutions, aiming to bridge the gap between affordability and biomimicry [ 15 , 103 ].
- Challenges : SoC models face challenges in controlling chemical gradients, technical sampling, and analysis. Integrating vasculature and microbiomes is crucial for physiological accuracy. Despite these, models like EpiDerm from MatTek Corporation or SkinEthic from L’Oréal show promise for dermocosmetics, offering safety and efficacy benefits over traditional methods [ 93 , 104 , 105 ].
3.3. Potential Solutions for Hair Loss and Promotion of Thicker, Healthier Hair Growth
- Low-level laser therapy (LLLT) stimulates hair growth with minimal side effects by exposing tissues to low-level light energy, showing a synergistic effect on promoting hair regrowth [ 107 ].
- Mesotherapy , involving the intradermal infusion of a mixture of factors, has demonstrated improvements in hair growth [ 93 , 108 , 109 ].
- Carboxytherapy , which entails the intradermic or subcutaneous insufflation of medical-grade sterile CO 2 , enhances blood flow and nutrient delivery to the HF, potentially aiding in cases of alopecia [ 93 , 110 , 111 ].
- Microneedling promotes hair growth by inducing percutaneous wounds with sterile microneedles, stimulating the activation of HFSCs and the release of growth factors that promote wound healing and angiogenesis [ 112 , 113 ].
- Autologous PRP treatment, derived from the patient’s blood, stimulates hair growth through the release of growth factors, cytokines, and chemokines, promoting cell proliferation, differentiation, and angiogenesis [ 114 ].
- Nanoparticles have been studied for drug delivery directly into the HF, minimizing the systemic adverse effects [ 107 ].
4. Revolutionizing Beauty: The Convergence of Regenerative Medicine and Cosmetic Science
5. challenges, future opportunities, and the role of ai in this field, 6. conclusions, author contributions, institutional review board statement, informed consent statement, data availability statement, acknowledgments, conflicts of interest.
- Bouwstra, J.A.; Nadaban, A.; Bras, W.; McCabe, C.; Bunge, A.; Gooris, G.S. The skin barrier: An extraordinary interface with an exceptional lipid organization. Prog. Lipid Res. 2023 , 92 , 101252. [ Google Scholar ] [ CrossRef ]
- Mansbridge, J. Skin tissue engineering. J. Biomater. Sci. Polym. Ed. 2008 , 19 , 955–968. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Lasisi, T.; Smallcombe, J.W.; Kenney, W.L.; Shriver, M.D.; Zydney, B.; Jablonski, N.G.; Havenith, G. Human scalp hair as a thermoregulatory adaptation. Proc. Natl. Acad. Sci. USA 2023 , 120 , e2301760120. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Silverberg, J.I. Comorbidities and the impact of atopic dermatitis. Ann. Allergy Asthma Immunol. 2019 , 123 , 144–151. [ Google Scholar ] [ CrossRef ]
- Hwang, H.W.; Ryou, S.; Jeong, J.H.; Lee, J.W.; Lee, K.J.; Lee, S.B.; Shin, H.T.; Byun, J.W.; Shin, J.; Choi, G.S. The Quality of Life and Psychosocial Impact on Female Pattern Hair Loss. Ann. Dermatol. 2024 , 36 , 44–52. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Ahluwalia, J.; Fabi, S.G. The psychological and aesthetic impact of age-related hair changes in females. J. Cosmet. Dermatol. 2019 , 18 , 1161–1169. [ Google Scholar ] [ CrossRef ]
- Schielein, M.C.; Tizek, L.; Ziehfreund, S.; Sommer, R.; Biedermann, T.; Zink, A. Stigmatization caused by hair loss—A systematic literature review. JDDG J. Der Dtsch. Dermatol. Ges. 2020 , 18 , 1357–1368. [ Google Scholar ] [ CrossRef ]
- Nandy, P.; Shrivastava, T. Exploring the Multifaceted Impact of Acne on Quality of Life and Well-Being. Cureus 2024 , 16 , e52727. [ Google Scholar ] [ CrossRef ]
- Bertoli, M.J.; Sadoughifar, R.; Schwartz, R.A.; Lotti, T.M.; Janniger, C.K. Female pattern hair loss: A comprehensive review. Dermatol. Ther. 2020 , 33 , e14055. [ Google Scholar ] [ CrossRef ]
- York, K.; Meah, N.; Bhoyrul, B.; Sinclair, R. A review of the treatment of male pattern hair loss. Expert Opin. Pharmacother. 2020 , 21 , 603–612. [ Google Scholar ] [ CrossRef ]
- Dhapte-Pawar, V.; Kadam, S.; Saptarsi, S.; Kenjale, P.P. Nanocosmeceuticals: Facets and aspects. Future Sci. OA 2020 , 6 , FSO613. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Vyas, K.S.; Kaufman, J.; Munavalli, G.S.; Robertson, K.; Behfar, A.; Wyles, S.P. Exosomes: The latest in regenerative aesthetics. Regen. Med. 2023 , 18 , 181–194. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Mathes, S.H.; Ruffner, H.; Graf-Hausner, U. The use of skin models in drug development. Adv. Drug Deliv. Rev. 2014 , 69–70 , 81–102. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Sarkiri, M.; Fox, S.C.; Fratila-Apachitei, L.E.; Zadpoor, A.A. Bioengineered Skin Intended for Skin Disease Modeling. Int. J. Mol. Sci. 2019 , 20 , 1407. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Gao, G.; Yonezawa, T.; Hubbell, K.; Dai, G.; Cui, X. Inkjet-bioprinted acrylated peptides and PEG hydrogel with human mesenchymal stem cells promote robust bone and cartilage formation with minimal printhead clogging. Biotechnol. J. 2015 , 10 , 1568–1577. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Yin, J.; Yan, M.; Wang, Y.; Fu, J.; Suo, H. 3D Bioprinting of Low-Concentration Cell-Laden Gelatin Methacrylate (GelMA) Bioinks with a Two-Step Cross-linking Strategy. ACS Appl. Mater. Interfaces 2018 , 10 , 6849–6857. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Jin, Z.; Li, Y.; Yu, K.; Liu, L.; Fu, J.; Yao, X.; Zhang, A.; He, Y. 3D Printing of Physical Organ Models: Recent Developments and Challenges. Adv. Sci. 2021 , 8 , e2101394. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Khalili, M.H.; Zhang, R.; Wilson, S.; Goel, S.; Impey, S.A.; Aria, A.I. Additive Manufacturing and Physicomechanical Characteristics of PEGDA Hydrogels: Recent Advances and Perspective for Tissue Engineering. Polymers 2023 , 15 , 2341. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Matama, T.; Costa, C.; Fernandes, B.; Araujo, R.; Cruz, C.F.; Tortosa, F.; Sheeba, C.J.; Becker, J.D.; Gomes, A.; Cavaco-Paulo, A. Changing human hair fibre colour and shape from the follicle. J. Adv. Res. 2023 . [ Google Scholar ] [ CrossRef ]
- Monavarian, M.; Kader, S.; Moeinzadeh, S.; Jabbari, E. Regenerative Scar-Free Skin Wound Healing. Tissue Eng. Part B Rev. 2019 , 25 , 294–311. [ Google Scholar ] [ CrossRef ]
- Dehkordi, A.N.; Babaheydari, F.M.; Chehelgerdi, M.; Dehkordi, S.R. Skin tissue engineering: Wound healing based on stem-cell-based therapeutic strategies. Stem Cell Res. Ther. 2019 , 10 , 111. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Jo, H.; Brito, S.; Kwak, B.M.; Park, S.; Lee, M.G.; Bin, B.H. Applications of Mesenchymal Stem Cells in Skin Regeneration and Rejuvenation. Int. J. Mol. Sci. 2021 , 22 , 2410. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Zhou, C.; Zhang, B.; Yang, Y.; Jiang, Q.; Li, T.; Gong, J.; Tang, H.; Zhang, Q. Stem cell-derived exosomes: Emerging therapeutic opportunities for wound healing. Stem Cell Res. Ther. 2023 , 14 , 107. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Eberlin, S.; Silva, M.S.D.; Facchini, G.; Silva, G.H.D.; Pinheiro, A.; Eberlin, S.; Pinheiro, A.D.S. The Ex Vivo Skin Model as an Alternative Tool for the Efficacy and Safety Evaluation of Topical Products. Altern. Lab. Anim. 2020 , 48 , 10–22. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Bataillon, M.; Lelievre, D.; Chapuis, A.; Thillou, F.; Autourde, J.B.; Durand, S.; Boyera, N.; Rigaudeau, A.S.; Besne, I.; Pellevoisin, C. Characterization of a New Reconstructed Full Thickness Skin Model, T-Skin, and its Application for Investigations of Anti-Aging Compounds. Int. J. Mol. Sci. 2019 , 20 , 2240. [ Google Scholar ] [ CrossRef ]
- Shin, S.H.; Lee, Y.H.; Rho, N.K.; Park, K.Y. Skin aging from mechanisms to interventions: Focusing on dermal aging. Front. Physiol. 2023 , 14 , 1195272. [ Google Scholar ] [ CrossRef ]
- Gref, R.; Delomenie, C.; Maksimenko, A.; Gouadon, E.; Percoco, G.; Lati, E.; Desmaele, D.; Zouhiri, F.; Couvreur, P. Vitamin C-squalene bioconjugate promotes epidermal thickening and collagen production in human skin. Sci. Rep. 2020 , 10 , 16883. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Skibska, A.; Perlikowska, R. Signal Peptides—Promising Ingredients in Cosmetics. Curr. Protein Pept. Sci. 2021 , 22 , 716–728. [ Google Scholar ] [ CrossRef ]
- Bukhari, S.N.A.; Roswandi, N.L.; Waqas, M.; Habib, H.; Hussain, F.; Khan, S.; Sohail, M.; Ramli, N.A.; Thu, H.E.; Hussain, Z. Hyaluronic acid, a promising skin rejuvenating biomedicine: A review of recent updates and pre-clinical and clinical investigations on cosmetic and nutricosmetic effects. Int. J. Biol. Macromol. 2018 , 120 , 1682–1695. [ Google Scholar ] [ CrossRef ]
- Spataro, E.A.; Dierks, K.; Carniol, P.J. Microneedling-Associated Procedures to Enhance Facial Rejuvenation. Facial Plast. Surg. Clin. N. Am. 2022 , 30 , 389–397. [ Google Scholar ] [ CrossRef ]
- Kim, Y.J.; Yoo, S.M.; Park, H.H.; Lim, H.J.; Kim, Y.L.; Lee, S.; Seo, K.W.; Kang, K.S. Exosomes derived from human umbilical cord blood mesenchymal stem cells stimulates rejuvenation of human skin. Biochem. Biophys. Res. Commun. 2017 , 493 , 1102–1108. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Sierra-Sanchez, A.; Kim, K.H.; Blasco-Morente, G.; Arias-Santiago, S. Cellular human tissue-engineered skin substitutes investigated for deep and difficult to heal injuries. NPJ Regen. Med. 2021 , 6 , 35. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Baek, J.; Lee, M.G. Oxidative stress and antioxidant strategies in dermatology. Redox Rep. 2016 , 21 , 164–169. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Trouba, K.J.; Hamadeh, H.K.; Amin, R.P.; Germolec, D.R. Oxidative stress and its role in skin disease. Antioxid. Redox Signal. 2002 , 4 , 665–673. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Popa, G.L.; Mitran, C.I.; Mitran, M.I.; Tampa, M.; Matei, C.; Popa, M.I.; Georgescu, S.R. Markers of Oxidative Stress in Patients with Acne: A Literature Review. Life 2023 , 13 , 1433. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Jones, D.A. Rosacea, reactive oxygen species, and azelaic Acid. J. Clin. Aesthet. Dermatol. 2009 , 2 , 26–30. [ Google Scholar ] [ PubMed ]
- Rahimi, H.; Mirnezami, M.; Yazdabadi, A.; Hajihashemi, A. Evaluation of systemic oxidative stress in patients with melasma. J. Cosmet. Dermatol. 2024 , 23 , 284–288. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Prie, B.E.; Voiculescu, V.M.; Ionescu-Bozdog, O.B.; Petrutescu, B.; Iosif, L.; Gaman, L.E.; Clatici, V.G.; Stoian, I.; Giurcaneanu, C. Oxidative stress and alopecia areata. J. Med. Life 2015 , 8 , 43–46. [ Google Scholar ]
- Zapatero-Solana, E.; Garcia-Gimenez, J.L.; Guerrero-Aspizua, S.; Garcia, M.; Toll, A.; Baselga, E.; Duran-Moreno, M.; Markovic, J.; Garcia-Verdugo, J.M.; Conti, C.J.; et al. Oxidative stress and mitochondrial dysfunction in Kindler syndrome. Orphanet J. Rare Dis. 2014 , 9 , 211. [ Google Scholar ] [ CrossRef ]
- Padayatty, S.J.; Katz, A.; Wang, Y.; Eck, P.; Kwon, O.; Lee, J.H.; Chen, S.; Corpe, C.; Dutta, A.; Dutta, S.K.; et al. Vitamin C as an antioxidant: Evaluation of its role in disease prevention. J. Am. Coll. Nutr. 2003 , 22 , 18–35. [ Google Scholar ] [ CrossRef ]
- Niki, E. Interaction of ascorbate and alpha-tocopherol. Ann. N. Y. Acad. Sci. 1987 , 498 , 186–199. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Farris, P.K. Topical vitamin C: A useful agent for treating photoaging and other dermatologic conditions. Dermatol. Surg. 2005 , 31 , 814–817; discussion 818. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Turcov, D.; Zbranca-Toporas, A.; Suteu, D. Bioactive Compounds for Combating Oxidative Stress in Dermatology. Int. J. Mol. Sci. 2023 , 24 , 17517. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Shaban, S.; El-Husseny, M.W.A.; Abushouk, A.I.; Salem, A.M.A.; Mamdouh, M.; Abdel-Daim, M.M. Effects of Antioxidant Supplements on the Survival and Differentiation of Stem Cells. Oxidative Med. Cell. Longev. 2017 , 2017 , 5032102. [ Google Scholar ] [ CrossRef ]
- Luo, L.; Kawakatsu, M.; Guo, C.W.; Urata, Y.; Huang, W.J.; Ali, H.; Doi, H.; Kitajima, Y.; Tanaka, T.; Goto, S.; et al. Effects of antioxidants on the quality and genomic stability of induced pluripotent stem cells. Sci. Rep. 2014 , 4 , 3779. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Choi, K.M.; Seo, Y.K.; Yoon, H.H.; Song, K.Y.; Kwon, S.Y.; Lee, H.S.; Park, J.K. Effect of ascorbic acid on bone marrow-derived mesenchymal stem cell proliferation and differentiation. J. Biosci. Bioeng. 2008 , 105 , 586–594. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Sies, H.; Jones, D.P. Reactive oxygen species (ROS) as pleiotropic physiological signalling agents. Nat. Rev. Mol. Cell Biol. 2020 , 21 , 363–383. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Loo, A.E.; Wong, Y.T.; Ho, R.; Wasser, M.; Du, T.; Ng, W.T.; Halliwell, B. Effects of hydrogen peroxide on wound healing in mice in relation to oxidative damage. PLoS ONE 2012 , 7 , e49215. [ Google Scholar ] [ CrossRef ]
- Lyublinskaya, O.G.; Borisov, Y.G.; Pugovkina, N.A.; Smirnova, I.S.; Obidina, J.V.; Ivanova, J.S.; Zenin, V.V.; Shatrova, A.N.; Borodkina, A.V.; Aksenov, N.D.; et al. Reactive Oxygen Species Are Required for Human Mesenchymal Stem Cells to Initiate Proliferation after the Quiescence Exit. Oxidative Med. Cell. Longev. 2015 , 2015 , 502105. [ Google Scholar ] [ CrossRef ]
- De Deken, X.; Corvilain, B.; Dumont, J.E.; Miot, F. Roles of DUOX-mediated hydrogen peroxide in metabolism, host defense, and signaling. Antioxid. Redox Signal. 2014 , 20 , 2776–2793. [ Google Scholar ] [ CrossRef ]
- Guo, L.; Du, J.; Yuan, D.F.; Zhang, Y.; Zhang, S.; Zhang, H.C.; Mi, J.W.; Ning, Y.L.; Chen, M.J.; Wen, D.L.; et al. Optimal H 2 O 2 preconditioning to improve bone marrow mesenchymal stem cells’ engraftment in wound healing. Stem Cell Res. Ther. 2020 , 11 , 434. [ Google Scholar ] [ CrossRef ]
- Buron, M.; Palomares, T.; Garrido-Pascual, P.; Herrero de la Parte, B.; Garcia-Alonso, I.; Alonso-Varona, A. Conditioned Medium from H 2 O 2 —Preconditioned Human Adipose-Derived Stem Cells Ameliorates UVB-Induced Damage to Human Dermal Fibroblasts. Antioxidants 2022 , 11 , 2011. [ Google Scholar ] [ CrossRef ]
- Lee, Y.; Son, J.Y.; Kang, J.I.; Park, K.M.; Park, K.D. Hydrogen Peroxide-Releasing Hydrogels for Enhanced Endothelial Cell Activities and Neovascularization. ACS Appl. Mater. Interfaces 2018 , 10 , 18372–18379. [ Google Scholar ] [ CrossRef ]
- Pleguezuelos-Beltran, P.; Galvez-Martin, P.; Nieto-Garcia, D.; Marchal, J.A.; Lopez-Ruiz, E. Advances in spray products for skin regeneration. Bioact. Mater. 2022 , 16 , 187–203. [ Google Scholar ] [ CrossRef ]
- Martinez-Santamaria, L.; Guerrero-Aspizua, S.; Del Rio, M. Skin bioengineering: Preclinical and clinical applications. Actas Dermosifiliogr. 2012 , 103 , 5–11. [ Google Scholar ] [ CrossRef ]
- Chocarro-Wrona, C.; Lopez-Ruiz, E.; Peran, M.; Galvez-Martin, P.; Marchal, J.A. Therapeutic strategies for skin regeneration based on biomedical substitutes. J. Eur. Acad. Dermatol. Venereol. 2019 , 33 , 484–496. [ Google Scholar ] [ CrossRef ]
- Disphanurat, W.; Sivapornpan, N.; Srisantithum, B.; Leelawattanachai, J. Efficacy of a triamcinolone acetonide-loaded dissolving microneedle patch for the treatment of hypertrophic scars and keloids: A randomized, double-blinded, placebo-controlled split-scar study. Arch. Dermatol. Res. 2023 , 315 , 989–997. [ Google Scholar ] [ CrossRef ]
- Waghmare, K.B.; Sequeira, J.; Rao, B.H.S. An objective assessment of microneedling therapy in atrophic facial acne scars. Natl. J. Maxillofac. Surg. 2022 , 13 , S103–S107. [ Google Scholar ] [ CrossRef ]
- Long, T.; Gupta, A.; Ma, S.; Hsu, S. Platelet-rich plasma in noninvasive procedures for atrophic acne scars: A systematic review and meta-analysis. J. Cosmet. Dermatol. 2020 , 19 , 836–844. [ Google Scholar ] [ CrossRef ]
- Mony, M.P.; Harmon, K.A.; Hess, R.; Dorafshar, A.H.; Shafikhani, S.H. An Updated Review of Hypertrophic Scarring. Cells 2023 , 12 , 678. [ Google Scholar ] [ CrossRef ]
- Tartaglia, G.; Cao, Q.; Padron, Z.M.; South, A.P. Impaired Wound Healing, Fibrosis, and Cancer: The Paradigm of Recessive Dystrophic Epidermolysis Bullosa. Int. J. Mol. Sci. 2021 , 22 , 5104. [ Google Scholar ] [ CrossRef ]
- Cao, Z.; Jin, S.; Wang, P.; He, Q.; Yang, Y.; Gao, Z.; Wang, X. Microneedle based adipose derived stem cells-derived extracellular vesicles therapy ameliorates UV-induced photoaging in SKH-1 mice. J. Biomed. Mater. Res. A 2021 , 109 , 1849–1857. [ Google Scholar ] [ CrossRef ]
- Sorg, H.; Tilkorn, D.J.; Hager, S.; Hauser, J.; Mirastschijski, U. Skin Wound Healing: An Update on the Current Knowledge and Concepts. Eur. Surg. Res. 2017 , 58 , 81–94. [ Google Scholar ] [ CrossRef ]
- Lin, X.; Zhu, L.; He, J. Morphogenesis, Growth Cycle and Molecular Regulation of Hair Follicles. Front. Cell Dev. Biol. 2022 , 10 , 899095. [ Google Scholar ] [ CrossRef ]
- Vandishi, A.K.; Esmaeili, A.; Taghipour, N. The promising prospect of human hair follicle regeneration in the shadow of new tissue engineering strategies. Tissue Cell 2024 , 87 , 102338. [ Google Scholar ] [ CrossRef ]
- Llamas-Molina, J.M.; Carrero-Castano, A.; Ruiz-Villaverde, R.; Campos, A. Tissue Engineering and Regeneration of the Human Hair Follicle in Androgenetic Alopecia: Literature Review. Life 2022 , 12 , 117. [ Google Scholar ] [ CrossRef ]
- Castro, A.R.; Logarinho, E. Tissue engineering strategies for human hair follicle regeneration: How far from a hairy goal? Stem Cells Transl. Med. 2020 , 9 , 342–350. [ Google Scholar ] [ CrossRef ]
- Xu, K.; Yu, E.; Wu, M.; Wei, P.; Yin, J. Cells, growth factors and biomaterials used in tissue engineering for hair follicles regeneration. Regen. Ther. 2022 , 21 , 596–610. [ Google Scholar ] [ CrossRef ]
- Kageyama, T.; Yan, L.; Shimizu, A.; Maruo, S.; Fukuda, J. Preparation of hair beads and hair follicle germs for regenerative medicine. Biomaterials 2019 , 212 , 55–63. [ Google Scholar ] [ CrossRef ]
- Wang, J.; Miao, Y.; Huang, Y.; Lin, B.; Liu, X.; Xiao, S.; Du, L.; Hu, Z.; Xing, M. Bottom-up Nanoencapsulation from Single Cells to Tunable and Scalable Cellular Spheroids for Hair Follicle Regeneration. Adv. Healthc. Mater. 2018 , 7 , 1700447. [ Google Scholar ] [ CrossRef ]
- Zhang, Y.; Yin, P.; Huang, J.; Yang, L.; Liu, Z.; Fu, D.; Hu, Z.; Huang, W.; Miao, Y. Scalable and high-throughput production of an injectable platelet-rich plasma (PRP)/cell-laden microcarrier/hydrogel composite system for hair follicle tissue engineering. J. Nanobiotechnol. 2022 , 20 , 465. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Kageyama, T.; Nanmo, A.; Yan, L.; Nittami, T.; Fukuda, J. Effects of platelet-rich plasma on in vitro hair follicle germ preparation for hair regenerative medicine. J. Biosci. Bioeng. 2020 , 130 , 666–671. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Zhang, K.; Bai, X.; Yuan, Z.; Cao, X.; Jiao, X.; Qin, Y.; Wen, Y.; Zhang, X. Cellular Nanofiber Structure with Secretory Activity-Promoting Characteristics for Multicellular Spheroid Formation and Hair Follicle Regeneration. ACS Appl. Mater. Interfaces 2020 , 12 , 7931–7941. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Fernandez-Martos, S.; Calvo-Sanchez, M.; Garcia-Alonso, K.; Castro, B.; Hashtroody, B.; Espada, J. Sustained Human Hair Follicle Growth Ex Vivo in a Glycosaminoglycan Hydrogel Matrix. Int. J. Mol. Sci. 2019 , 20 , 1741. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Xie, S.; Chen, L.; Zhang, M.; Zhang, C.; Li, H. Self-assembled complete hair follicle organoids by coculture of neonatal mouse epidermal cells and dermal cells in Matrigel. Ann. Transl. Med. 2022 , 10 , 767. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Zhang, X.; Xiao, S.; Liu, B.; Miao, Y.; Hu, Z. Use of extracellular matrix hydrogel from human placenta to restore hair-inductive potential of dermal papilla cells. Regen. Med. 2019 , 14 , 741–751. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Barat, T.; Abdollahimajd, F.; Dadkhahfar, S.; Moravvej, H. Evaluation of the efficacy and safety of cow placenta extract lotion versus minoxidil 2% in the treatment of female pattern androgenetic alopecia. Int. J. Women’s Dermatol. 2020 , 6 , 318–321. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Motter Catarino, C.; Cigaran Schuck, D.; Dechiario, L.; Karande, P. Incorporation of hair follicles in 3D bioprinted models of human skin. Sci. Adv. 2023 , 9 , eadg0297. [ Google Scholar ] [ CrossRef ]
- Kang, D.; Liu, Z.; Qian, C.; Huang, J.; Zhou, Y.; Mao, X.; Qu, Q.; Liu, B.; Wang, J.; Hu, Z.; et al. 3D bioprinting of a gelatin-alginate hydrogel for tissue-engineered hair follicle regeneration. Acta Biomater. 2023 , 165 , 19–30. [ Google Scholar ] [ CrossRef ]
- Kang, M.S.; Kwon, M.; Lee, S.H.; Kim, W.H.; Lee, G.W.; Jo, H.J.; Kim, B.; Yang, S.Y.; Kim, K.S.; Han, D.W. 3D Printing of Skin Equivalents with Hair Follicle Structures and Epidermal-Papillary-Dermal Layers Using Gelatin/Hyaluronic Acid Hydrogels. Chem. Asian J. 2022 , 17 , e202200620. [ Google Scholar ] [ CrossRef ]
- Zheng, S.Y.; Hu, X.M.; Huang, K.; Li, Z.H.; Chen, Q.N.; Yang, R.H.; Xiong, K. Proteomics as a tool to improve novel insights into skin diseases: What we know and where we should be going. Front. Surg. 2022 , 9 , 1025557. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Benoit, I.; Burty-Valin, E.; Radman, M. A Proteome-Centric View of Ageing, including that of the Skin and Age-Related Diseases: Considerations of a Common Cause and Common Preventative and Curative Interventions. Clin. Cosmet. Investig. Dermatol. 2023 , 16 , 79–85. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Pien, N.; Bray, F.; Gheysens, T.; Tytgat, L.; Rolando, C.; Mantovani, D.; Dubruel, P.; Vlierberghe, S.V. Proteomics as a tool to gain next level insights into photo-crosslinkable biopolymer modifications. Bioact. Mater. 2022 , 17 , 204–220. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Tan, K.X.; Chang, T.; Lin, X. Secretomes as an emerging class of bioactive ingredients for enhanced cosmeceutical applications. Exp. Dermatol. 2022 , 31 , 674–688. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Masutin, V.; Kersch, C.; Schmitz-Spanke, S. A systematic review: Metabolomics-based identification of altered metabolites and pathways in the skin caused by internal and external factors. Exp. Dermatol. 2022 , 31 , 700–714. [ Google Scholar ] [ CrossRef ]
- Knox, S.; O’Boyle, N.M. Skin lipids in health and disease: A review. Chem. Phys. Lipids 2021 , 236 , 105055. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Chen, T.; Zhu, Z.; Du, Q.; Wang, Z.; Wu, W.; Xue, Y.; Wang, Y.; Wu, Y.; Zeng, Q.; Jiang, C.; et al. A Skin Lipidomics Study Reveals the Therapeutic Effects of Tanshinones in a Rat Model of Acne. Front. Pharmacol. 2021 , 12 , 675659. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Chen, H.; Zhao, Q.; Zhong, Q.; Duan, C.; Krutmann, J.; Wang, J.; Xia, J. Skin Microbiome, Metabolome and Skin Phenome, from the Perspectives of Skin as an Ecosystem. Phenomics 2022 , 2 , 363–382. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Gueniche, A.; Perin, O.; Bouslimani, A.; Landemaine, L.; Misra, N.; Cupferman, S.; Aguilar, L.; Clavaud, C.; Chopra, T.; Khodr, A. Advances in Microbiome-Derived Solutions and Methodologies Are Founding a New Era in Skin Health and Care. Pathogens 2022 , 11 , 121. [ Google Scholar ] [ CrossRef ]
- Tong, T.; Geng, R.; Kang, S.G.; Li, X.; Huang, K. Revitalizing Photoaging Skin through Eugenol in UVB-Exposed Hairless Mice: Mechanistic Insights from Integrated Multi-Omics. Antioxidants 2024 , 13 , 168. [ Google Scholar ] [ CrossRef ]
- Bonafont, J.; Mencia, A.; Garcia, M.; Torres, R.; Rodriguez, S.; Carretero, M.; Chacon-Solano, E.; Modamio-Hoybjor, S.; Marinas, L.; Leon, C.; et al. Clinically Relevant Correction of Recessive Dystrophic Epidermolysis Bullosa by Dual sgRNA CRISPR/Cas9-Mediated Gene Editing. Mol. Ther. 2019 , 27 , 986–998. [ Google Scholar ] [ CrossRef ]
- Carretero, M.; Guerrero-Aspizua, S.; Del Rio, M. Bioengineered skin humanized model of psoriasis. Methods Mol. Biol. 2013 , 961 , 305–323. [ Google Scholar ] [ CrossRef ]
- Guerrero-Aspizua, S.; Carretero, M.; Conti, C.J.; Del Rio, M. The importance of immunity in the development of reliable animal models for psoriasis and atopic dermatitis. Immunol. Cell Biol. 2020 , 98 , 626–638. [ Google Scholar ] [ CrossRef ]
- Fernandez-Carro, E.; Angenent, M.; Gracia-Cazana, T.; Gilaberte, Y.; Alcaine, C.; Ciriza, J. Modeling an Optimal 3D Skin-on-Chip within Microfluidic Devices for Pharmacological Studies. Pharmaceutics 2022 , 14 , 1417. [ Google Scholar ] [ CrossRef ]
- Rodrigues, R.O.; Sousa, P.C.; Gaspar, J.; Banobre-Lopez, M.; Lima, R.; Minas, G. Organ-on-a-Chip: A Preclinical Microfluidic Platform for the Progress of Nanomedicine. Small 2020 , 16 , e2003517. [ Google Scholar ] [ CrossRef ]
- Ponmozhi, J.; Dhinakaran, S.; Varga-Medveczky, Z.; Fonagy, K.; Bors, L.A.; Ivan, K.; Erdo, F. Development of Skin-On-A-Chip Platforms for Different Utilizations: Factors to Be Considered. Micromachines 2021 , 12 , 294. [ Google Scholar ] [ CrossRef ]
- Mohamadali, M.; Ghiaseddin, A.; Irani, S.; Amirkhani, M.A.; Dahmardehei, M. Design and evaluation of a skin-on-a-chip pumpless microfluidic device. Sci. Rep. 2023 , 13 , 8861. [ Google Scholar ] [ CrossRef ]
- Wufuer, M.; Lee, G.; Hur, W.; Jeon, B.; Kim, B.J.; Choi, T.H.; Lee, S. Skin-on-a-chip model simulating inflammation, edema and drug-based treatment. Sci. Rep. 2016 , 6 , 37471. [ Google Scholar ] [ CrossRef ]
- Randall, M.J.; Jungel, A.; Rimann, M.; Wuertz-Kozak, K. Advances in the Biofabrication of 3D Skin in vitro: Healthy and Pathological Models. Front. Bioeng. Biotechnol. 2018 , 6 , 154. [ Google Scholar ] [ CrossRef ]
- Bellas, E.; Seiberg, M.; Garlick, J.; Kaplan, D.L. In vitro 3D full-thickness skin-equivalent tissue model using silk and collagen biomaterials. Macromol. Biosci. 2012 , 12 , 1627–1636. [ Google Scholar ] [ CrossRef ]
- Parenteau-Bareil, R.; Gauvin, R.; Cliche, S.; Gariepy, C.; Germain, L.; Berthod, F. Comparative study of bovine, porcine and avian collagens for the production of a tissue engineered dermis. Acta Biomater. 2011 , 7 , 3757–3765. [ Google Scholar ] [ CrossRef ]
- Rosa, E.; Diaferia, C.; Gianolio, E.; Sibillano, T.; Gallo, E.; Smaldone, G.; Stornaiuolo, M.; Giannini, C.; Morelli, G.; Accardo, A. Multicomponent Hydrogel Matrices of Fmoc-FF and Cationic Peptides for Application in Tissue Engineering. Macromol. Biosci. 2022 , 22 , e2200128. [ Google Scholar ] [ CrossRef ]
- Arab, W.T.; Susapto, H.H.; Alhattab, D.; Hauser, C.A.E. Peptide nanogels as a scaffold for fabricating dermal grafts and 3D vascularized skin models. J. Tissue Eng. 2022 , 13 , 20417314221111868. [ Google Scholar ] [ CrossRef ]
- Bal-Ozturk, A.; Miccoli, B.; Avci-Adali, M.; Mogtader, F.; Sharifi, F.; Cecen, B.; Yasayan, G.; Braeken, D.; Alarcin, E. Current Strategies and Future Perspectives of Skin-on-a-Chip Platforms: Innovations, Technical Challenges and Commercial Outlook. Curr. Pharm. Des. 2018 , 24 , 5437–5457. [ Google Scholar ] [ CrossRef ]
- Vurat, M.T.; Ergun, C.; Elcin, A.E.; Elcin, Y.M. 3D Bioprinting of Tissue Models with Customized Bioinks. Adv. Exp. Med. Biol. 2020 , 1249 , 67–84. [ Google Scholar ] [ CrossRef ]
- Jimenez, F.; Alam, M.; Vogel, J.E.; Avram, M. Hair transplantation: Basic overview. J. Am. Acad. Dermatol. 2021 , 85 , 803–814. [ Google Scholar ] [ CrossRef ]
- Katzer, T.; Leite Junior, A.; Beck, R.; da Silva, C. Physiopathology and current treatments of androgenetic alopecia: Going beyond androgens and anti-androgens. Dermatol. Ther. 2019 , 32 , e13059. [ Google Scholar ] [ CrossRef ]
- Gupta, A.K.; Polla Ravi, S.; Wang, T.; Talukder, M.; Starace, M.; Piraccini, B.M. Systematic review of mesotherapy: A novel avenue for the treatment of hair loss. J. Dermatol. Treat. 2023 , 34 , 2245084. [ Google Scholar ] [ CrossRef ]
- Tang, Z.; Hu, Y.; Wang, J.; Fan, Z.; Qu, Q.; Miao, Y. Current application of mesotherapy in pattern hair loss: A systematic review. J. Cosmet. Dermatol. 2022 , 21 , 4184–4193. [ Google Scholar ] [ CrossRef ]
- Bagherani, N.; Smoller, B.R.; Tavoosidana, G.; Ghanadan, A.; Wollina, U.; Lotti, T. An overview of the role of carboxytherapy in dermatology. J. Cosmet. Dermatol. 2023 , 22 , 2399–2407. [ Google Scholar ] [ CrossRef ]
- Kroumpouzos, G.; Arora, G.; Kassir, M.; Galadari, H.; Wollina, U.; Lotti, T.; Grabbe, S.; Goldust, M. Carboxytherapy in dermatology. Clin. Dermatol. 2022 , 40 , 305–309. [ Google Scholar ] [ CrossRef ]
- English, R.S., Jr.; Ruiz, S.; DoAmaral, P. Microneedling and Its Use in Hair Loss Disorders: A Systematic Review. Dermatol. Ther (Heidelb) 2022 , 12 , 41–60. [ Google Scholar ] [ CrossRef ]
- Ocampo-Garza, S.S.; Fabbrocini, G.; Ocampo-Candiani, J.; Cinelli, E.; Villani, A. Micro needling: A novel therapeutic approach for androgenetic alopecia, A Review of Literature. Dermatol. Ther. 2020 , 33 , e14267. [ Google Scholar ] [ CrossRef ]
- Paichitrojjana, A.; Paichitrojjana, A. Platelet Rich Plasma and Its Use in Hair Regrowth: A Review. Drug Des. Dev. Ther. 2022 , 16 , 635–645. [ Google Scholar ] [ CrossRef ]
- Yuan, A.R.; Bian, Q.; Gao, J.Q. Current advances in stem cell-based therapies for hair regeneration. Eur. J. Pharmacol. 2020 , 881 , 173197. [ Google Scholar ] [ CrossRef ]
- Rose, P.T. Advances in Hair Restoration. Dermatol. Clin. 2018 , 36 , 57–62. [ Google Scholar ] [ CrossRef ]
- Kwack, M.H.; Seo, C.H.; Gangadaran, P.; Ahn, B.C.; Kim, M.K.; Kim, J.C.; Sung, Y.K. Exosomes derived from human dermal papilla cells promote hair growth in cultured human hair follicles and augment the hair-inductive capacity of cultured dermal papilla spheres. Exp. Dermatol. 2019 , 28 , 854–857. [ Google Scholar ] [ CrossRef ]
- Mahmoudian-Sani, M.R.; Jamshidi, M.; Asgharzade, S. Combined Growth Factor and Gene Therapy: An Approach for Hair Cell Regeneration and Hearing Recovery. ORL J. Otorhinolaryngol. Relat. Spec. 2018 , 80 , 326–337. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Elder, A.; Cappelli, M.O.; Ring, C.; Saedi, N. Artificial intelligence in cosmetic dermatology: An update on current trends. Clin. Dermatol. 2024 , 42 , 216–220. [ Google Scholar ] [ CrossRef ]
- Waddell, S.J.; de Andres, M.C.; Tsimbouri, P.M.; Alakpa, E.V.; Cusack, M.; Dalby, M.J.; Oreffo, R.O. Biomimetic oyster shell-replicated topography alters the behaviour of human skeletal stem cells. J. Tissue Eng. 2018 , 9 , 2041731418794007. [ Google Scholar ] [ CrossRef ]
Click here to enlarge figure
| Component | Description | References |
|---|---|---|
| : : | [ , ] | |
| : : : | [ , ] | |
| : | [ , ] |
| Approach | Components | Function | References |
|---|---|---|---|
| (e.g., TGF-β, PDGF, IGF-1) | Promote collagen synthesis by skin cells | [ , ] | |
| (e.g., vitamin C, vitamin A) | Enhance collagen production and skin cell proliferation | [ ] | |
| (e.g., Palmitoyl pentapeptide-4) | Stimulate collagen and elastin synthesis | [ ] | |
| Plump up the skin and reduce the appearance of wrinkles | [ ] | ||
| Create controlled micro-injuries to stimulate collagen production and reduce wrinkle depth | [ ] | ||
| Promote collagen synthesis and improve skin elasticity through paracrine effects | [ ] | ||
| Enhance barrier function, hydration, immune response, and wound healing | [ ] |
| The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
Share and Cite
Pleguezuelos-Beltrán, P.; Herráiz-Gil, S.; Martínez-Moreno, D.; Medraño-Fernandez, I.; León, C.; Guerrero-Aspizua, S. Regenerative Cosmetics: Skin Tissue Engineering for Anti-Aging, Repair, and Hair Restoration. Cosmetics 2024 , 11 , 121. https://doi.org/10.3390/cosmetics11040121
Pleguezuelos-Beltrán P, Herráiz-Gil S, Martínez-Moreno D, Medraño-Fernandez I, León C, Guerrero-Aspizua S. Regenerative Cosmetics: Skin Tissue Engineering for Anti-Aging, Repair, and Hair Restoration. Cosmetics . 2024; 11(4):121. https://doi.org/10.3390/cosmetics11040121
Pleguezuelos-Beltrán, Paula, Sara Herráiz-Gil, Daniel Martínez-Moreno, Iria Medraño-Fernandez, Carlos León, and Sara Guerrero-Aspizua. 2024. "Regenerative Cosmetics: Skin Tissue Engineering for Anti-Aging, Repair, and Hair Restoration" Cosmetics 11, no. 4: 121. https://doi.org/10.3390/cosmetics11040121
Article Metrics
Article access statistics, further information, mdpi initiatives, follow mdpi.

Subscribe to receive issue release notifications and newsletters from MDPI journals

IMAGES
COMMENTS
Skincare rooted in innovative formulation, that focuses on supporting healthy skin function. Facials An opportunity to learn more about your skin, and receive beneficial treatments. ... an approach to makeup that perfectly fits the thesis client. easy, multi-use, high pigment delivery, gorgeously crafted. manasi7 in the know
skin skin; cleanser mist + tonic serum oil spf mask tools all manasi7 body grocery gift card wear thesis shop all treatments about contact faqs blog blog; Skin Health Product Use Lauren's Health / A Series
thesis is a small skincare studio in the heart of missoula, montana. LAUREN HANSEN. Born and raised two blocks away from Thesis and having spent much of my childhood in the 425 building, this space truly is home. My family has been in this neighborhood for over 100 years. Opening this business has been a dream extending beyond becoming an ...
Meet Aleksandra, your skincare expert. Thanks for checking out our 100% natural plant-based, eco-friendly skincare. Our collection, free from harsh chemicals and allergens, includes gentle cleansers, toners, serums, and moisturizers. We believe in beauty that's kind to both you and the planet. Choose us for a natural, skin-friendly journey.
Thesis beauty is a luxury natural and organic range of skincare and bodycare. 100% Vegan & Cruelty-free! Thesis brand is available to buy at Coco & Gizmo. We sell Thesis Eye Serum, strawberry face mask, chocolate face mask, face serum, gentle cleanser, facial oils for dry skin, eye serum, facial serums for dry skin.
Thesis, Somerville, Massachusetts. 4,349 likes · 2 talking about this. Providing the purest Organic, Vegan, Eco-friendly products -- beauty as nature intended!
Thesis Beauty Takes Organic Skin Care Seriously . Though Thesis Beauty has been around since 2009, this was the first time that I tried their organic skin care. All their formulas are handcrafted and use 99% pure certified organic ingredients without any synthetics or fillers that may compromise their high standards to preserve the power of nature.
The facial serum from Thesis Beauty is a must beauty product for anyone looking for a non-toxic, highly effective and inexpensive skincare product. I have very dry sensitive skin. ... Even my make up looks better now that I have incorporated the serum into my skin care routine. I have used many oils in the past; some were ok, some did not meet ...
I have sensitive skin and allergic to many fragranced and chemical laden products on the market. This facial serum with Sea buckthorn hydrates so wonderfully making my skin feel healthy and smooth. My order was received in a timely fashion, with the items arriving safely. The Thesis Beauty surprised me with a small gift of another product to try.
Use of skin personal care products on a regular basis is nearly ubiquitous, but their effects on molecular and microbial diversity of the skin are unknown. We evaluated the impact of four beauty products (a facial lotion, a moisturizer, a foot powder, and a deodorant) on 11 volunteers over 9 weeks. Mass spectrometry and 16S rRNA inventories of the skin revealed decreases in chemical as well as ...
Each treatment at Thesis is customized to what your skin is experiencing at that time. We utilize a full backbar of Marie Veronique products and techniques that work to support your skin. Our treatment options reflect the time on the bed, please budget 20-30 additional minutes for your visit. First time clients are recommended to book for a 60 ...
Skin Thesis uses advanced skincare technology including lasers, lights, and injectables to achieve natural, youthful skin. Our Philosophy is to achieve the most natural look. Using latest innovations in aesthetic medicine. top of page. POLICIES. At Skin Thesis®, we value and appreciate every client. Please note our policies and procedures:
skin care products purchasing obtain the effects of the interaction between their personal demographics and attitude, the emergence of metrosexual and marketing mix (especially ... this thesis such their kind encouragements, generosity, and patience in providing valuable comments and recommendations on every earlier draft. Without them, this ...
3.1.2 Descriptive research - The purpose of the descriptive research was to provide a clearer picture of consumers' attitudes toward organic skin and personal care including key value benefits that affect purchase intentions. Figure 3.2 displays the picture of all variables in this study.
The quest for youthful, healthy skin and full, vibrant hair has long been a driving force in the dermocosmetics field. However, traditional approaches often struggle to address the underlying causes of aging, damage, and hair loss. Regenerative cosmetics powered by skin tissue engineering offer a transformative alternative. This review explores the emerging field of using engineered skin ...
skin skin; cleanser mist + tonic serum oil spf mask tools all manasi7 body grocery gift card wear thesis shop all treatments about book now contact faqs blog blog; Skin Health Product Use Lauren's Health / A Series Search Search Search. Register / Login Currency Currency ...
Top Dzerzhinsky Landmarks: See reviews and photos of sights to see in Dzerzhinsky, Russia on Tripadvisor.
您可以向此项目. Zhukovsky coat of arms. Date of adoption: April 25, 2002. Russian Heraldic Register no. 959. Textual description: "On a sky-blue (azure) field there are three wide arrow-heads in a triangle (two and one in a form of a plane). Above them there're two wings. All figures in gold". 2009年2月2日. File:Zhukovsky coat of arms ...
skin skin; cleanser mist + tonic serum oil spf mask tools all manasi7 body grocery gift card wear thesis shop all treatments about contact faqs blog blog; Skin Health Product Use Lauren's Health / A Series
Media in category "Elektrostal" The following 200 files are in this category, out of 222 total. (previous page) ()
State Housing Inspectorate of the Moscow Region Elektrostal postal code 144009. See Google profile, Hours, Phone, Website and more for this business. 2.0 Cybo Score. Review on Cybo.
Thesis 425 South Orange Street Missoula, Montana 406-546-5491 [email protected]